-
Články
- Časopisy
- Kurzy
- Témy
- Kongresy
- Videa
- Podcasty
Entry and Fusion of Emerging Paramyxoviruses
article has not abstract
Published in the journal: Entry and Fusion of Emerging Paramyxoviruses. PLoS Pathog 6(6): e32767. doi:10.1371/journal.ppat.1000881
Category: Pearls
doi: https://doi.org/10.1371/journal.ppat.1000881Summary
article has not abstract
Paramyxoviruses are a family of non-segmented RNA viruses that includes major human pathogens such as measles virus and respiratory syncytial virus (RSV) and significant animal viruses like rinderpest [1]. In recent years, several new paramyxoviruses have been identified, further increasing the breadth and importance of this viral family. While many elements of the fusion and entry mechanisms of these recently identified pathogens are conserved, there are interesting differences, including variations in receptor binding, cell tropism, fusion (F) protein proteolytic activation, and triggering of membrane fusion. Thus, study of their entry mechanisms has highlighted the diversity of these critical events in the family.
Paramyxoviruses: An Expanding Group of Important Viral Pathogens
Hendra virus and Nipah virus are the only identified zoonotic members of the paramyxovirus family, and both are highly pathogenic in humans [2]. Hendra virus infection has resulted in multiple horse and four human fatalities since its emergence in Australia in 1994, with outbreaks in 2008 and 2009 leading to rising concern in the Australian horse breeding industry. Nipah virus emerged in Malaysia in 1999, causing an outbreak of viral encephalitis that led to 105 human fatalities out of 265 reported cases. Containment of the 1999 Nipah virus epidemic required the sacrifice of more than 1 million swine. Continued Nipah outbreaks have occurred in Southeast Asia, with mortality rates of up to 70% and suspected human-to-human transmission. Numerous molecular features have led to the placement of Hendra and Nipah viruses within a new genus in the paramyxovirus family, the henipaviruses (Figure 1). The principal reservoir species for both viruses is thought to be Pteropus fruit bats, but a number of other species have been shown to be susceptible to infection [3].
Fig. 1. A phylogenetic tree of the paramyxovirus family, built using fusion protein sequence comparison. 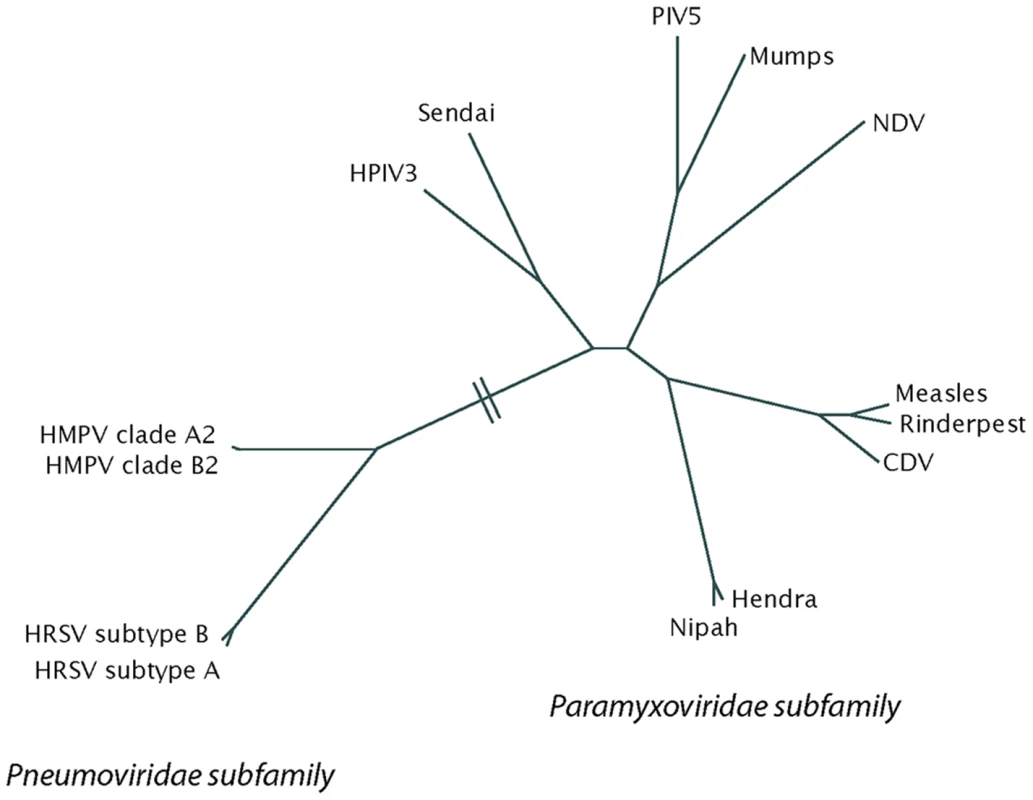
The tree was generated from a PROMALS multiple sequence alignment using PROTDIST and FITCH from the PHYLIP 3.65 software package and displayed using HYPERTREE 1.0.0. CDV, canine distemper virus; HMPV, human metapneumovirus; HPIV3, human parainfluenza virus 3; HRSV, human respiratory syncytial virus; NDV, Newcastle disease virus; PIV5, parainfluenza virus 5. Human metapneumovirus (HMPV) was first identified in 2001, but unlike Hendra and Nipah, HMPV is not a new human virus resulting from zoonotic transmission. Instead, HMPV is a long-term human pathogen that was only identified by careful analysis of samples from children with respiratory tract disease for which an etiological agent had not been identified [4]. Subsequent studies indicate that HMPV is a major causative agent of respiratory tract infections worldwide, causing between 5% and 15% of lower respiratory tract infections in young children [5]. HMPV has been circulating in the human population since at least 1958 [4]. Sequence analysis places HMPV in the Pneumovirinae subfamily, along with RSV.
Fusion Mechanisms: Conserved Features in Newly Identified Paramyxoviruses
To enter host cells, paramyxoviruses must go through the key steps of viral attachment to the target cell, followed by fusion of the viral membrane to a host cell membrane [6]. Two major viral glycoproteins promote these events: the attachment protein facilitates primary receptor binding of the virus to the target cell, while the F protein promotes the subsequent membrane fusion events. Both events are hypothesized to occur at the cell surface in a neutral pH environment. Interactions between the F protein and the homotypic attachment protein are hypothesized to control initiation of the fusion process for most paramyxoviruses, though the mechanistic details of triggering control remain elusive. Once begun, fusion is promoted by a series of conformational changes in the F protein that first lead to insertion of a hydrophobic region (termed the fusion peptide) into the target membrane, forming a protein bridge between the two membranes. Additional conformational changes lead to formation of a helical bundle, formed by interactions between two heptad repeat regions that do not interact in the prefusion form of the protein [1], and subsequent membrane fusion.
A number of factors point to an overall conserved mechanism of fusion promotion among the paramyxovirus F proteins. While there is considerable heterogeneity at the amino acid level, F proteins from both established and newly identified paramyxoviruses display conserved positioning of cysteine, glycine, and proline residues, suggesting an overall conservation of structure. F proteins also contain similarly placed fusion peptide and heptad repeat regions. Peptides corresponding to the F protein heptad repeat regions have been shown to block fusion and entry for previously studied paramyxoviruses, and similar peptides inhibit Hendra, Nipah, and HMPV fusion and entry, indicating that the requirement for formation of the final helical bundle is a conserved feature [2], [6]. Like previously identified members of the family, fusion activity of the Hendra and Nipah F proteins requires the presence of a viral attachment protein, though either the Hendra or Nipah attachment protein can be used interchangeably [6]. As was seen with measles virus, recent evidence suggests that fusion activity for the Hendra and Nipah F proteins is inversely proportional to the strength of the F attachment protein interactions, in contrast to results from other paramyxovirus systems such as Newcastle disease viruses [7], suggesting slightly different mechanisms of control of fusion initiation.
Identification of Ephrin B2 as the Receptor for Hendra and Nipah Viruses: Implications for Tissue and Species Range
Initial characterization of the activity of the Hendra and Nipah attachment proteins indicated interaction with a protein receptor, as is the case for measles virus attachment protein, rather than the sialic acid binding observed for most paramyxovirus attachment proteins [3]. Further study identified EphrinB2 as the receptor for the Hendra and Nipah viruses [8], [9], with EphrinB3 later shown to serve as an additional receptor for both viruses. Structural analysis of Nipah G alone or in complex with Ephrin B3 interestingly showed little conformational change upon receptor binding, suggesting that only subtle alterations in the attachment protein lead to F protein activation [10]. EphrinB2 and B3 serve as ligands for the Eph tyrosine receptor family, and their cellular expression in neurons, arterial endothelial cells, and smooth muscle is consistent with the tissue distribution observed during Hendra and Nipah infection [3]. EphrinB2 and B3 are also highly conserved between species, fitting with the large number of species shown to be infected by these pathogens. EphrinB2 and B3 from multiple infectable species, including human, horse, pig, cat, dog, and bat, have been shown to serve as functional receptors for Hendra and Nipah [11], suggesting that the conserved expression of this receptor plays an important role in the unusually broad host range of these pathogens. Interestingly, murine EphrinB2 can serve as a functional receptor for these viruses, but mice are resistant to henipavirus infection, indicating that additional factors modulate overall host range.
Cathepsin L Processing of the Hendra and Nipah F Proteins: A New Paradigm for Fusion Protein Proteolytic Activation
Like other paramyxovirus F proteins, the Hendra and Nipah virus F proteins are initially synthesized as a precursor (F0) that must be proteolytically processed to two subunits (F1 and F2) to be fusogenically active (Figure 2A). For the majority of F proteins, this critical proteolytic processing event is promoted by furin, a cellular protease present primarily in the trans-Golgi network. Interestingly, the mechanism for proteolytic activation of the henipavirus F proteins is completely novel. Furin is clearly not involved, as there is no furin consensus at the cleavage site, furin inhibitors have no effect on henipavirus F processing, and processing occurs efficiently in furin-negative cell lines [2]. Instead, inhibitors or shRNA knock-downs of the cellular endosomal protease cathepsin L were shown to inhibit cleavage of the Hendra and Nipah F proteins, and in vitro studies confirmed proteolytic cleavage of the henipavirus F proteins at a single specific site by purified cathepsin L [6], [12]. To facilitate this key interaction with cathepsin L, endocytosis of the Hendra F protein [13] and the Nipah F protein [14] must occur, followed by a retrafficking event to the cell surface after proteolytic processing (Figure 2B). As cleaved F protein is present within the packaged virion [3], this complex trafficking of the henipavirus F proteins through the endosomal pathway occurs prior to viral assembly. Interestingly, the Hendra G attachment protein does not follow this complicated trafficking pathway, indicating that the critical attachment protein: fusion protein interactions needed for fusion occur only after F protein endocytic trafficking and proteolytic cleavage [15]. The reason for this novel activation pathway is unclear, though it is intriguing to note that Ebola virus and SARS coronavirus also have a role for cathepsin L at some point during the viral life cycle, and like Hendra and Nipah virus, the reservoir species for these important pathogens is thought to be bats. Future studies on protease profiles in bat cells may shed light on the reason for the unusual role of cathepsins in the life cycles of these pathogens.
Fig. 2. Proteolytic processing of the henipavirus fusion proteins. 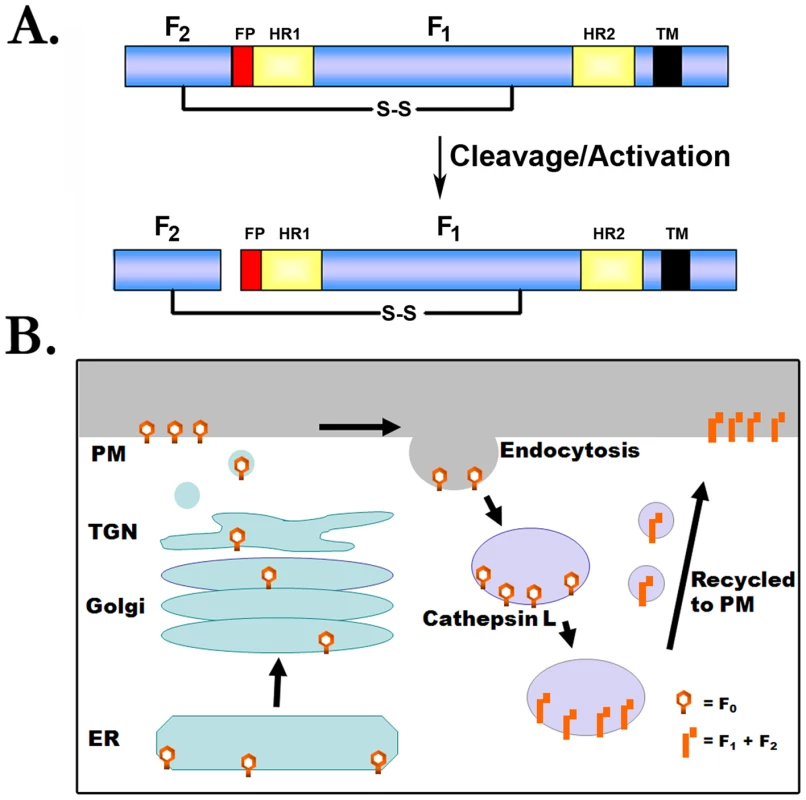
(A) Schematic of F protein cleavage/activation. FP, fusion peptide; HR1 and HR2, heptad repeat; S-S, disulfide bond; TM, transmembrane domain. (B) Model for proteolytic processing of the henipavirus F proteins. The Hendra or Nipah F proteins are synthesized in the endoplasmic reticulum (ER), transit through the secretory pathway to the plasma membrane (PM), and are subsequently endocytosed. Following interaction with cathepsin L at an undetermined point in the endocytic pathway, the cleaved protein is recycled to the plasma membrane. Analysis of HMPV: Novel Findings on Attachment, Fusion, and the Entry Pathway
While analysis of henipavirus entry mechanisms has broadened diversity related to receptor usage and proteolytic activation, study of HMPV entry has further illuminated differences between the Paramyxovirinae and Pneumovirinae sub-families (Figure 1). Specific attachment protein∶fusion protein interactions are needed for fusion promoted by Paramyxovirinae glycoproteins, with the exception of the parainfluenza virus 5 F protein, which can promote fusion in the absence of its attachment protein, though at a significantly decreased level [1]. In contrast, a recombinant RSV lacking the attachment protein can replicate in some types of cultured cells [5], indicating a significantly decreased requirement for this protein during attachment and entry. The altered role of the attachment protein is even more striking in HMPV, as recombinants lacking the attachment protein are competent for replication in African green monkeys, though there is decreased replication in the lower respiratory tract [16]. Studies of cell–cell fusion found that the HMPV F protein by itself was capable of promoting both binding and fusion, and no stimulation by the attachment protein was observed [17]. Combined with the viral studies, these results suggest that HMPV F can interact with a receptor(s) on the target cells, though the identity of the F receptor remains to be defined. These findings also raise the important question of what triggers the HMPV F protein to initiate fusion, as in this case fusion initiation is clearly not controlled by interactions with the attachment protein. Interestingly, recent studies indicate that low pH can serve as a fusion trigger [17] for at least a portion of the HMPV strains [18], and specific residues that could promote low-pH-induced conformational change through an electrostatic repulsion mechanism have been identified [19]. In addition, endocytosis has very recently been implicated in both HMPV [19] and RSV [20] entry, indicating that the virus will come in contact with the acidic pH of the endosomes during entry.
Overall, the study of entry and membrane fusion of recently identified paramyxoviruses has broadened the paradigms of receptor usage, F protein proteolytic activation, and membrane fusion triggering. Future work will continue to define how these variations modulate infectivity and pathogenicity in this important viral family.
Zdroje
1. LambRA
ParksGD
2007 Paramyxoviridae: the viruses and their replication.
KnipeDM
HowleyPM
Fields Virology: Lippincott, Williams and Wilkins 1449 1496
2. EatonBT
BroderCC
MiddletonD
WangLF
2006 Hendra and Nipah viruses: different and dangerous. Nat Rev Microbiol 4 23 35
3. EatonBT
MackenzieJS
WangL-F
2007 Henipaviruses.
KnipeDM
HowleyPM
Fields Virology: Lippincott, Williams and Wilkins 1587 1600
4. van den HoogenBG
de JongJC
GroenJ
KuikenT
de GrootR
2001 A newly discovered human pneumovirus isolated from young children with respiratory tract disease. Nat Med 7 719 724
5. CollinsPL
CroweJE
2007 Respiratory syncytial virus and metapneumovirus.
KnipeDM
HowleyPM
Fields Virology: Lippincott, Williams and Wilkins 1601 1646
6. SmithEC
PopaA
ChangA
MasanteC
DutchRE
2009 Viral entry mechanisms: the increasing diversity of paramyxovirus entry. FEBS J
7. AguilarHC
AtamanZA
AspericuetaV
FangAQ
StroudM
2009 A novel receptor-induced activation site in the Nipah virus attachment glycoprotein (G) involved in triggering the fusion glycoprotein (F). J Biol Chem 284 1628 1635
8. BonaparteMI
DimitrovAS
BossartKN
CrameriG
MungallBA
2005 Ephrin-B2 ligand is a functional receptor for Hendra virus and Nipah virus. Proc Natl Acad Sci U S A 102 10652 10657
9. NegreteOA
LevroneyEL
AguilarHC
Bertolotti-CiarletA
NazarianR
2005 EphrinB2 is the entry receptor for Nipah virus, an emergent deadly paramyxovirus. Nature 436 401 405
10. XuK
RajashankarKR
ChanYP
HimanenJP
BroderCC
2008 Host cell recognition by the henipaviruses: crystal structures of the Nipah G attachment glycoprotein and its complex with ephrin-B3. Proc Natl Acad Sci U S A 105 9953 9958
11. BossartKN
TachedjianM
McEachernJA
CrameriG
ZhuZ
2008 Functional studies of host-specific ephrin-B ligands as Henipavirus receptors. Virology 372 357 371
12. PagerCT
DutchRE
2005 Cathepsin L is involved in proteolytic processing of the Hendra virus fusion protein. J Virol 79 12714 12720
13. MeulendykeKA
WurthMA
McCannRO
DutchRE
2005 Endocytosis plays a critical role in proteolytic processing of the Hendra virus fusion protein. J Virol 79 12643 12649
14. DiederichS
MollM
KlenkHD
MaisnerA
2005 The nipah virus fusion protein is cleaved within the endosomal compartment. J Biol Chem 280 29899 29903
15. WhitmanSD
SmithEC
DutchRE
2009 Differential rates of protein folding and cellular trafficking for the Hendra virus F and G proteins: implications for F-G complex formation. J Virol 83 8998 9001
16. BiacchesiS
PhamQN
SkiadopoulosMH
MurphyBR
CollinsPL
2005 Infection of nonhuman primates with recombinant human metapneumovirus lacking the SH, G, or M2-2 protein categorizes each as a nonessential accessory protein and identifies vaccine candidates. J Virol 79 12608 12613
17. SchowalterRM
SmithSE
DutchRE
2006 Characterization of human metapneumovirus F protein-promoted membrane fusion: critical roles for proteolytic processing and low pH. J Virol 80 10931 10941
18. HerfstS
MasV
VerLS
WierdaRJ
OsterhausAD
2008 Low pH induced membrane fusion mediated by human metapneumoviruses F protein is a rare, strain dependent phenomenon. J Virol 82 8891 8895
19. SchowalterRM
ChangA
RobachJG
BuchholzUJ
DutchRE
2009 Low-pH triggering of human metapneumovirus fusion: essential residues and importance in entry. J Virol 83 1511 1522
20. KolokoltsovAA
DenigerD
FlemingEH
RobertsNJJr
KarpilowJM
2007 Small interfering RNA profiling reveals key role of clathrin-mediated endocytosis and early endosome formation for infection by respiratory syncytial virus. J Virol 81 7786 7800
Štítky
Hygiena a epidemiológia Infekčné lekárstvo Laboratórium
Článek Insight into the Mechanisms of Adenovirus Capsid Disassembly from Studies of Defensin NeutralizationČlánek Serum-Dependent Selective Expression of EhTMKB1-9, a Member of B1 Family of Transmembrane KinasesČlánek Host Cell Invasion and Virulence in Sepsis Is Facilitated by the Multiple Repeats within FnBPA
Článok vyšiel v časopisePLOS Pathogens
Najčítanejšie tento týždeň
2010 Číslo 6- Parazitičtí červi v terapii Crohnovy choroby a dalších zánětlivých autoimunitních onemocnění
- Očkování proti virové hemoragické horečce Ebola experimentální vakcínou rVSVDG-ZEBOV-GP
- Koronavirus hýbe světem: Víte jak se chránit a jak postupovat v případě podezření?
-
Všetky články tohto čísla
- Cognitive Dysfunction Is Sustained after Rescue Therapy in Experimental Cerebral Malaria, and Is Reduced by Additive Antioxidant Therapy
- DNA Watermarking of Infectious Agents: Progress and Prospects
- Self-Protection against Gliotoxin—A Component of the Gliotoxin Biosynthetic Cluster, GliT, Completely Protects Against Exogenous Gliotoxin
- Modifies the Tsetse Salivary Composition, Altering the Fly Feeding Behavior That Favors Parasite Transmission
- Requirement of NOX2 and Reactive Oxygen Species for Efficient RIG-I-Mediated Antiviral Response through Regulation of MAVS Expression
- The Enteropathogenic Effector EspF Targets and Disrupts the Nucleolus by a Process Regulated by Mitochondrial Dysfunction
- Epithelial p38α Controls Immune Cell Recruitment in the Colonic Mucosa
- Coexpression of PD-1, 2B4, CD160 and KLRG1 on Exhausted HCV-Specific CD8+ T Cells Is Linked to Antigen Recognition and T Cell Differentiation
- Insight into the Mechanisms of Adenovirus Capsid Disassembly from Studies of Defensin Neutralization
- EspA Acts as a Critical Mediator of ESX1-Dependent Virulence in by Affecting Bacterial Cell Wall Integrity
- An RNA Element at the 5′-End of the Poliovirus Genome Functions as a General Promoter for RNA Synthesis
- Tetherin Restricts Productive HIV-1 Cell-to-Cell Transmission
- Epstein-Barr Virus-Encoded LMP2A Induces an Epithelial–Mesenchymal Transition and Increases the Number of Side Population Stem-like Cancer Cells in Nasopharyngeal Carcinoma
- Protein Kinase A Dependent Phosphorylation of Apical Membrane Antigen 1 Plays an Important Role in Erythrocyte Invasion by the Malaria Parasite
- Requirement for Ergosterol in V-ATPase Function Underlies Antifungal Activity of Azole Drugs
- The SOCS-Box of HIV-1 Vif Interacts with ElonginBC by Induced-Folding to Recruit Its Cul5-Containing Ubiquitin Ligase Complex
- A Crucial Role for Infected-Cell/Antibody Immune Complexes in the Enhancement of Endogenous Antiviral Immunity by Short Passive Immunotherapy
- Modulation of the Arginase Pathway in the Context of Microbial Pathogenesis: A Metabolic Enzyme Moonlighting as an Immune Modulator
- Role of Abl Kinase and the Wave2 Signaling Complex in HIV-1 Entry at a Post-Hemifusion Step
- Serum-Dependent Selective Expression of EhTMKB1-9, a Member of B1 Family of Transmembrane Kinases
- Epigenetic Repression of by Latent Epstein-Barr Virus Requires the Interaction of EBNA3A and EBNA3C with CtBP
- Host Cell Invasion and Virulence in Sepsis Is Facilitated by the Multiple Repeats within FnBPA
- Role of PKR and Type I IFNs in Viral Control during Primary and Secondary Infection
- A Kinome RNAi Screen Identified AMPK as Promoting Poxvirus Entry through the Control of Actin Dynamics
- Cryptococcal Cell Morphology Affects Host Cell Interactions and Pathogenicity
- NleG Type 3 Effectors from Enterohaemorrhagic Are U-Box E3 Ubiquitin Ligases
- Human Cytomegalovirus UL29/28 Protein Interacts with Components of the NuRD Complex Which Promote Accumulation of Immediate-Early RNA
- Complement Receptor 1 Is a Sialic Acid-Independent Erythrocyte Receptor of
- A Viral microRNA Down-Regulates Multiple Cell Cycle Genes through mRNA 5′UTRs
- Immunotoxin Complementation of HAART to Deplete Persisting HIV-Infected Cell Reservoirs
- Protein Expression Redirects Vesicular Stomatitis Virus RNA Synthesis to Cytoplasmic Inclusions
- Entry and Fusion of Emerging Paramyxoviruses
- Paramyxovirus Entry and Targeted Vectors for Cancer Therapy
- Suppressing Glucose Transporter Gene Expression in Schistosomes Impairs Parasite Feeding and Decreases Survival in the Mammalian Host
- Formation of Complexes at Plasmodesmata for Potyvirus Intercellular Movement Is Mediated by the Viral Protein P3N-PIPO
- Fungal Cell Gigantism during Mammalian Infection
- Two Novel Point Mutations in Clinical Reduce Linezolid Susceptibility and Switch on the Stringent Response to Promote Persistent Infection
- Rotavirus Structural Proteins and dsRNA Are Required for the Human Primary Plasmacytoid Dendritic Cell IFNα Response
- The Epigenetic Landscape of Latent Kaposi Sarcoma-Associated Herpesvirus Genomes
- PLOS Pathogens
- Archív čísel
- Aktuálne číslo
- Informácie o časopise
Najčítanejšie v tomto čísle- Requirement of NOX2 and Reactive Oxygen Species for Efficient RIG-I-Mediated Antiviral Response through Regulation of MAVS Expression
- Formation of Complexes at Plasmodesmata for Potyvirus Intercellular Movement Is Mediated by the Viral Protein P3N-PIPO
- Insight into the Mechanisms of Adenovirus Capsid Disassembly from Studies of Defensin Neutralization
- Two Novel Point Mutations in Clinical Reduce Linezolid Susceptibility and Switch on the Stringent Response to Promote Persistent Infection
Prihlásenie#ADS_BOTTOM_SCRIPTS#Zabudnuté hesloZadajte e-mailovú adresu, s ktorou ste vytvárali účet. Budú Vám na ňu zasielané informácie k nastaveniu nového hesla.
- Časopisy



