-
Články
- Časopisy
- Kurzy
- Témy
- Kongresy
- Videa
- Podcasty
Glutamate Utilization Couples Oxidative Stress Defense and the Tricarboxylic Acid Cycle in Phagosomal Escape
Intracellular bacterial pathogens have developed a variety of strategies to avoid degradation by the host innate immune defense mechanisms triggered upon phagocytocis. Upon infection of mammalian host cells, the intracellular pathogen Francisella replicates exclusively in the cytosolic compartment. Hence, its ability to escape rapidly from the phagosomal compartment is critical for its pathogenicity. Here, we show for the first time that a glutamate transporter of Francisella (here designated GadC) is critical for oxidative stress defense in the phagosome, thus impairing intra-macrophage multiplication and virulence in the mouse model. The gadC mutant failed to efficiently neutralize the production of reactive oxygen species. Remarkably, virulence of the gadC mutant was partially restored in mice defective in NADPH oxidase activity. The data presented highlight links between glutamate uptake, oxidative stress defense, the tricarboxylic acid cycle and phagosomal escape. This is the first report establishing the role of an amino acid transporter in the early stage of the Francisella intracellular lifecycle.
Published in the journal: Glutamate Utilization Couples Oxidative Stress Defense and the Tricarboxylic Acid Cycle in Phagosomal Escape. PLoS Pathog 10(1): e32767. doi:10.1371/journal.ppat.1003893
Category: Research Article
doi: https://doi.org/10.1371/journal.ppat.1003893Summary
Intracellular bacterial pathogens have developed a variety of strategies to avoid degradation by the host innate immune defense mechanisms triggered upon phagocytocis. Upon infection of mammalian host cells, the intracellular pathogen Francisella replicates exclusively in the cytosolic compartment. Hence, its ability to escape rapidly from the phagosomal compartment is critical for its pathogenicity. Here, we show for the first time that a glutamate transporter of Francisella (here designated GadC) is critical for oxidative stress defense in the phagosome, thus impairing intra-macrophage multiplication and virulence in the mouse model. The gadC mutant failed to efficiently neutralize the production of reactive oxygen species. Remarkably, virulence of the gadC mutant was partially restored in mice defective in NADPH oxidase activity. The data presented highlight links between glutamate uptake, oxidative stress defense, the tricarboxylic acid cycle and phagosomal escape. This is the first report establishing the role of an amino acid transporter in the early stage of the Francisella intracellular lifecycle.
Introduction
Francisella tularensis is a Gram-negative bacterium causing the disease tularemia in a large number of animal species. This highly infectious bacterial pathogen can be transmitted to humans in numerous ways [1], including direct contact with sick animals, inhalation, ingestion of contaminated water or food, or by bites from ticks, mosquitoes or flies. Four different subspecies (subsp.) of F. tularensis that differ in virulence and geographic distribution exist, designated subsps. tularensis, holarctica, mediasiatica and novicida, respectively. The tularensis subspecies is the most virulent causing a severe disease in humans [2], [3]. F. tularensis subsp. novicida (F. novicida) is rarely pathogenic to non-immuno-compromized humans but is fully virulent for mice and is therefore widely used as a model to study Francisella intracellular parasitism.
F. novicida has the capacity to evade host defenses and to replicate to high numbers within the cytosol of eukaryotic cells [4]. The bacterium is able to enter and to replicate inside a variety of cells, and in particular in macrophages. After a transient passage through a phagosomal compartment, bacteria are released within 30–60 minutes in the host cell cytosol where they undergo several rounds of active replication [1]. Upon Francisella entry into macrophages, the phagosomal compartment transiently acidifies and the activation of NADPH oxidase leads to the production of noxious oxygen reactive species [5]. Although several genes required for phagosomal escape have been identified ([6], [7] and references therein), the molecular mechanisms underlying this complex process are still very poorly understood.
Protection against oxidative stress includes the production of anti-oxidant molecules (such as glutathione and NADPH) and of enzymes (such as catalases, superoxide dismutases glutaredoxin-related protein and alkylhydroperoxide reductases). Francisella subspecies encode a whole set of such oxidative stress-related enzymes [8]. Inactivation of the corresponding genes generally leads to increased sensitivity to oxidative stress, defective intracellular multiplication, and attenuated virulence [9], [10], [11]. Protection against oxidative and other stress also involves a number of dedicated protein chaperones and chaperone complexes [12].
In contrast, the importance of acid-resistance mechanisms in Francisella intracellular survival remains controversial [13], [14], [15] and their possible contribution to pathogenesis still largely unknown. One of the best characterized acid-resistance systems in bacteria couples the glutamate:γ-aminobutyrate exchanger GadC with the glutamate decarboxylase(s) GadA and/or GadB [16]. The decarboxylase replaces the α-carboxyl group of its amino acid substrate with a proton that is consumed from the cytoplasmic pool [17]. The capacity to produce γ-aminobutyric acid (GABA) through glutamate decarboxylation has been observed in both Gram-negative and Gram-positive bacteria. The GadC/GadB glutamate decarboxylase (GAD) system has been shown to play an essential role in acid tolerance in food-borne bacterial pathogens that must survive the potentially lethal acidic environments of the stomach before reaching the intestine. Some bacteria possess a unique permease-decarboxylase pair whereas others, like Listeria monocytogenes [18], encode several paralogues of each component.
Recent genome sequence analyses and genome-scale genetic studies suggest that an important proportion of genes related to metabolic and nutritional functions participate to Francisella virulence [19]. However, the relationship between nutrition and the in vivo life cycle of F. tularensis remain poorly understood. Francisella is predicted to possess numerous nutrient uptake systems to capture its necessary host-derived nutrients, some of which are probably available in limiting concentrations. Notably, we showed very recently that an asparagine transporter of the major facilitator superfamily of transporters was specifically required for cytoslic multiplication of Francisella and its systemic dissemination [20].
The amino acid-polyamine-organocation family of transporters (APC) is specifically involved in amino acid transport [19]. Remarkably, eight of the 11 APC members have been identified at least once in earlier genetic studies, and are likely to be involved in bacterial virulence. In particular, the gene encoding the GadC permease has been identified in several different genome-wide screens, performed in either F. tularensis subsp. holarctica [21] or F. novicida [22], [23], [24].
In the present work, we elucidate the functional role of the GadC protein in Francisella pathogenesis. We show that glutamate uptake plays a critical role in Francisella oxidative stress defense in the phagosomal compartment. Strikingly, the activity of GadC influences the expression of metabolic genes and the production of tricarboxylic acid (TCA) cycle intermediates, unraveling a relationship between oxidative stress defense, metabolism and Francisella virulence.
Results
The Gad system of Francisella
F. tularensis subspecies possess a unique putative GAD system, composed of the antiporter GadC and a decarboxylase GadB (encoded by genes FTN_0571 and FTN_1701 in F. novicida and hereafter designated gadC and gadB, respectively for simplification) (Figure S1A). The transcription of gadC is initiated 27 nucleotides upstream of the translational start from a predicted σ70 promoter (Figure S1B). This genetic organization is highly conserved in all the available F. tularensis genomes (not shown). The gene gadC encodes a protein of 469 amino acids sharing 98.7%, 99.1% and 99.6% identity with its orthologues in the subspecies mediasiatica (FTM_1423), holartica (FTL_1583) and tularensis (FTT_0480c), respectively.
The Francisella GadC protein is predicted as a putative glutamate:γ-aminobutyric acid (GAD) antiporter (KEGG database). Although it shows only modest homology (approximately 25% amino acid identity) with GadC of E. coli [25], secondary structure prediction (using the method for prediction of transmembrane helices HMM available at the internet site www.cbs.dtu.dk) indicates that the GadC transporter of Francisella also comprises 12 transmembrane helixes and has its N and C-terminal ends facing the cytoplasm (not shown).
The gadB gene encodes a putative glutamate decarboxylase protein of 448 amino acid residues that is highly conserved in F. tularensis subsp. tularensis (98.7% amino acid identity with FTT_1722c). However, the corresponding protein is truncated at its C-terminal end in the subspecies holarctica (FTL_1863) and mediasiatica (FTM_1673, and noted as a pseudogene in the KEGG database).
Role of GadC in bacterial intracellular multiplication and virulence
We constructed a strain with chromosomal deletion of the entire gadC gene in F. novicida by allelic replacement [26]. We confirmed that the ΔgadC mutation did not have any polar effect on the downstream gene FTN_0570 by quantitative qRT-PCR (Figure S1C). The growth kinetics of the parental F. novicida strain and the ΔgadC mutant were indistinguishable in tryptic soya broth (TSB) and chemically defined medium (CDM) [27] liquid media at 37°C (Figure S2), indicating that inactivation of gadC had no impact on bacterial growth in broth.
We examined the ability of wild-type F. novicida, the ΔgadC mutant and a ΔgadC mutant strain complemented with a plasmid-encoded copy of wild-type gadC, to survive in murine and human macrophage cell lines and primary bone marrow-derived mouse macrophages, over a 24 h-period. The ΔgadC mutant showed a severe growth defect in J774.1 cells, comparable to that of a mutant deleted of the entire Francisella pathogenicity island (ΔFPI mutant), with more than a 30-fold reduction of intracellular bacteria after 10 h and a 1,000-fold reduction after 24 h (Figure 1A). Impaired multiplication of the ΔgadC mutant was also observed in THP-1 macrophages (Figure 1B) as well as in bone marrow-derived macrophages (Figure 1C). In all cell types tested, introduction of the complementing plasmid (pKK-gadC) restored bacterial viability to same level as in the wild-type parent, confirming the specific involvement of the gadC gene in intracellular survival.
Fig. 1. gadC inactivation affects intracellular survival and virulence. 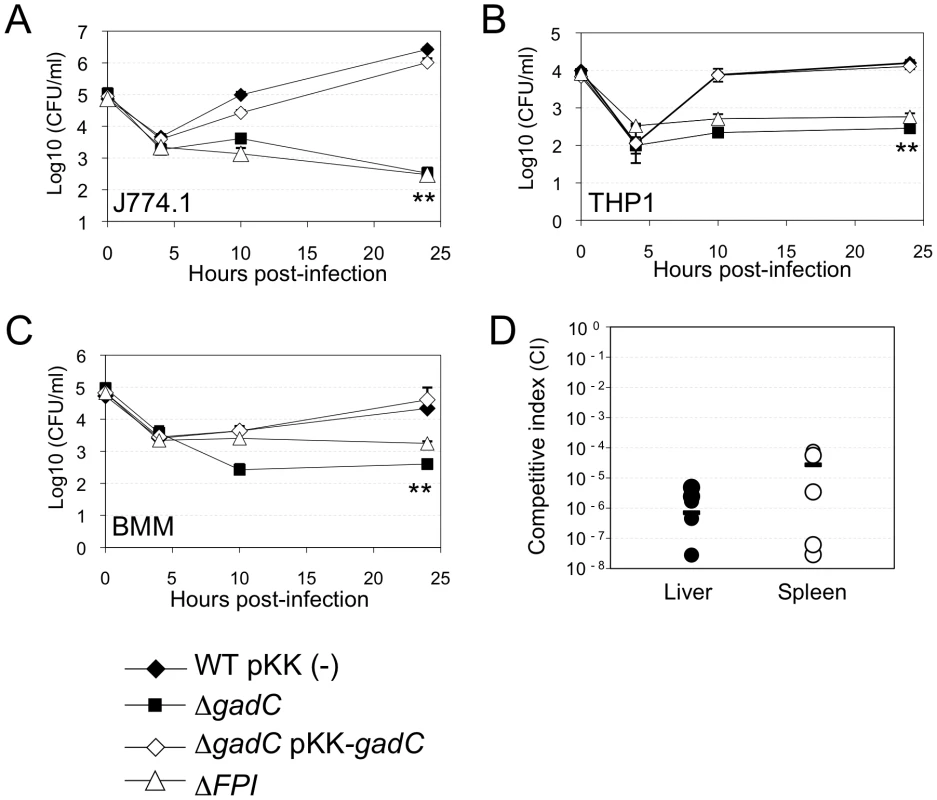
Intracellular replication of wild-type F. novicida (WT) carrying the empty plasmid pKK214 (WT/pKK(−)), of the ΔgadC mutant (ΔgadC) and complemented strain (ΔgadC/pKK-gadC), and of the ΔFPI mutant (ΔFPI), was monitored in J774.1 macrophage-like cells (A); in THP-1 human macrophages (B); and in bone marrow-derived macrophages (C), over a 24 h-period. Results are shown as the average of log10 cfu mL−1 ± standard deviation. Each experiment was performed in triplicate. **, p<0.01 (as determined by the Student's t-test). Competition assays (D). A group of five female BALB/c mice were infected i.p. with a 1∶1 mixture of wild-type F. novicida and ΔgadC mutant strains (100 colony forming units (cfu) of each). The data represent the competitive index (CI) value for cfu of mutant/wild-type in the liver (L: black diamonds, left column) and spleen (S: black circles, right column) of each mouse, 48 h after infection. Bars represent the geometric mean CI value. Next, in vivo competition assays in BALB/c mice were performed to determine if the GadC protein played a role in the ability of Francisella to cause disease. Five mice (6 - to 8-week old) were inoculated by the intraperitoneal (i.p.) route with a 1∶1 mixture of wild-type F. novicida and ΔgadC mutant strains. Bacterial multiplication in the liver and spleen was monitored at day 2 post-infection (Figure 1D). The Competition Index (CI), calculated for both organs, was extremely low (10−6) demonstrating that the gene gadC played an essential role in Francisella virulence in the mouse model.
Upon Francisella entry into cells, Francisella initially resides in a phagosomal compartment that transiently acidifies and that acquires reactive oxygen species. We therefore examined the ability of wild-type and ΔgadC mutant strains to survive under acid or oxidative stress conditions. For this, bacteria were exposed either to pH 5.5 or to 500 µM H2O2 (Figure 2). Under the pH condition tested, the viability of two strains was unaffected (Figure 2A). It should be noted that at the lower pH of 2.5, the viability of both wild-type and ΔgadC mutant was equally reduced (approximately 2 logs, not shown). In contrast, the ΔgadC mutant strain appeared to be significantly more sensitive to oxidative stress than the wild-type strain in TSB (Figure 2B). After 40 min of exposure, it showed a 10-fold decrease in the number of viable bacteria and an approximately 50-fold decrease after 60 min of exposure to H2O2. Remarkably, in CDM, the wild-type and ΔgadC mutant strains were equally sensitive to H2O2 in the absence of glutamate supplementation (Figure 2C). However, upon glutamate supplementation, the wild-type strain showed increased resistance to H2O2 whereas the ΔgadC strain was unaffected (Figure 2D).
Fig. 2. Stress sensitivity. 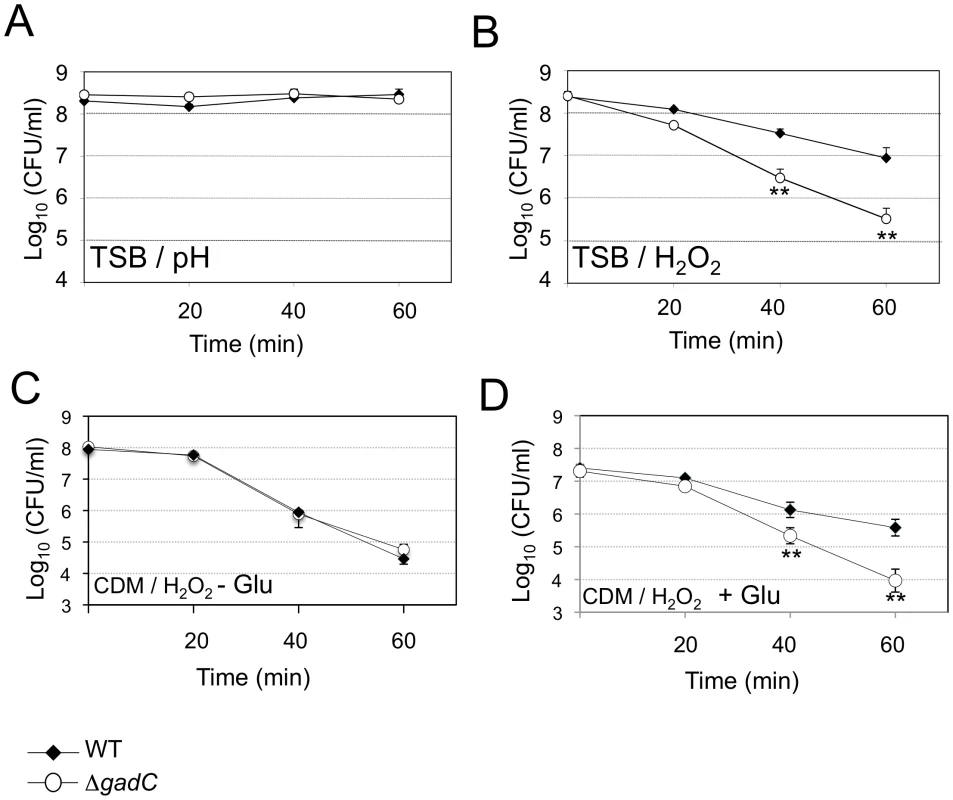
Exponential phase bacteria, diluted in TSB medium, were subjected: (A) to acidic stress (pH 5.5), or (B) to oxidative stress (500 µM H202). Exponential phase bacteria, diluted in chemically defined medium (CDM) (C), or CDM supplemented with 1 mM glutamate (D), were subjected to oxidative stress (500 µM H202). The bacteria were plated on chocolate agar plates at different times and viable bacteria were monitored 2 days after. Data are the average cfu mL−1 for three points. Experiments were realized twice. **, p<0.01 (as determined by the Student's t-test). GadC is involved in phagosomal escape
Confocal and electron microscopy analyses demonstrated that the ΔgadC mutant had lost the capacity to escape from the phagosomal compartment of infected macrophages.
Confocal microscopy
We used the differential solubilization process previously described [28] to follow the sub-cellular localization of the ΔgadC mutant in infected cells (Figure 3A). Briefly, the plasma membrane was selectively permeabilized with digitonin. This treatment allowed the detection of cytoplasmic bacteria and proteins. Subsequent treatment with saponin rendered intact phagosomes accessible to antibodies and allowed the detection of intra-phagosomal bacteria. Intracellular localization of the bacteria or LAMP-1 (used as a specific marker of phagosomes) was analyzed using specific antibodies and their co-localization was quantified with the Image J software (Figure 3B). Merging of LAMP-1 and bacteria was obtained at all three 3 time-point tested in cells infected with the ΔgadC or ΔFPI mutant strains. With both mutants, bacterial co-localization with LAMP-1 was elevated (70% and 74%, with ΔgadC and ΔFPI; respectively) after 1 h and remained very high after 4 h (75% and 64% with ΔgadC and ΔFPI; respectively) and after 10 h (63% and 64% with ΔgadC and ΔFPI; respectively). In contrast, co-localization of the wild-type strain with LAMP-1 was only around 24% after 1 h and remained in the same range throughout the infection (20% and 22%, after 4 h and 10 h, respectively). These results strongly suggest that the ΔgadC mutant is still trapped in the phagosomal compartment after 10 h, as the ΔFPI mutant, whereas the wild-type strain has already escaped into the cytosol after 1 h.
Fig. 3. Subcellular localization of the ΔgadC mutant. 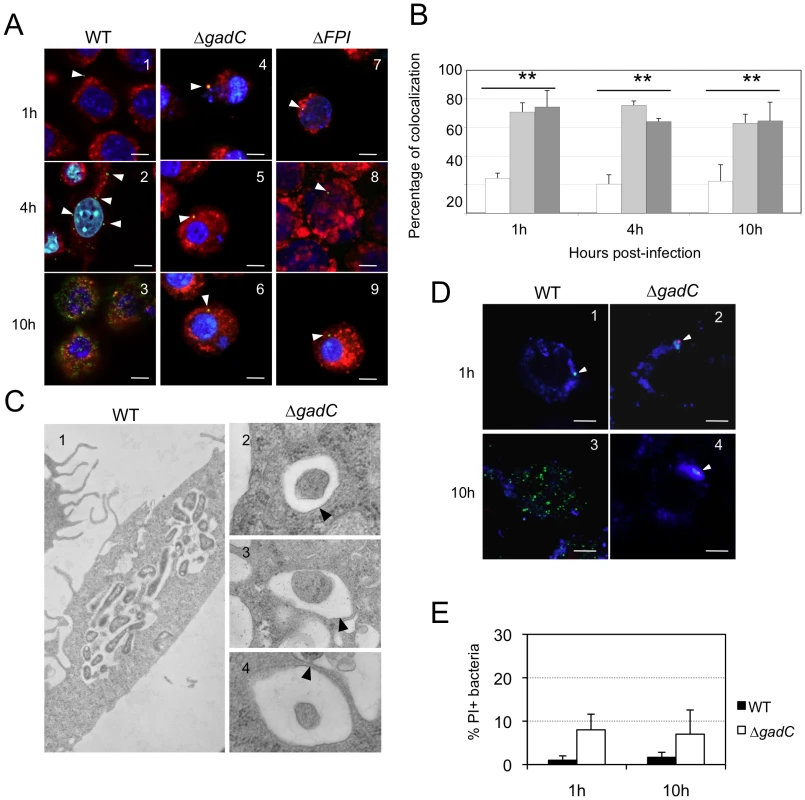
(A) Co-localization of wild-type F. novicida (1, 2, 3), ΔgadC (4, 5, 6) or ΔFPI mutant strain (7, 8, 9) with LAMP-1 was monitored by confocal microscopy, in J774.1 macrophage cells. Co-localization was monitored at 1 h (1, 4, 7), 4 h (2, 5, 8) and 10 h (3, 6, 9). Anti-Francisella antibody was used at a final dilution of 1∶500 and is shown in green. Anti-LAMP-1 antibody was used at a final dilution of 1∶100 and is shown in red. White arrowheads point to individual bacteria. Cell nuclei were labeled with DAPI (in blue). The images are representative from triplicate coverslips in three independent experiments. Scale bars at the bottom right of each panel correspond to 10 µM. (B) Quantification of co-localization between bacteria and LAMP-1 was obtained with Image J software. The graph results from the analysis of 4 different fields for each time of infection, in three independent experiments. **, p<0.01 (as determined by the Student's t-test). White bars, F. novicida U112 (WT); light grey bars, ΔgadC; dark grey bars, ΔFPI. (C) Transmission electron micrographs of thin sections of J774.1 macrophages, infected by wild-type F. novicida and ΔgadC mutant strains. Infections were monitored over a 10 h-period. At 10 h, active cytosolic multiplication of wild-type F. novicida was observed in most of the infected cells (1) whereas the ΔgadC mutant remains trapped into spacious phagosomes (2, 3, 4). Black arrowheads point to intact phagosomal membrane. (D) To evaluate the viability of intracellular Francisella, labeling with the cell-impermeant nucleic acid dye propidium iodide (PI) was performed. Confocal images of J774.1 cells, infected with wild-type F. novicida (1, 3) or ΔgadC mutant (2, 4) strain; after 1 h (1, 2) and 10 h of infection (3, 4). Intact bacteria are labeled in green. Bacteria with compromised membranes are labeled with PI and appear in red (or a red spot). Phagosomes are labeled in blue. Scale bars at the bottom right of each panel correspond to 10 µM. (E) Quantification of the percentage of dead bacteria. At least 100 bacteria per experiment were scored for PI labeling at 1 h and 10 h post infection. Data are means ± standard deviation from three independent assays. Electron microscopy
To confirm this result, we performed thin section electron microscopy of J774.1 cells infected either with wild-type F. novicida or with the ΔgadC mutant (Figure 3C). As expected, significant bacterial replication was observed in the cytosol of most infected cells 10 h post-infection with wild-type F. novicida whereas bacterial multiplication was severely impaired in cells infected with the ΔgadC mutant. Furthermore, mutant bacteria surrounded by intact phagosomal membrane were still observed after 10 h of infection.
Bacterial death
We then determined whether the ΔgadC mutant bacteria, trapped in the phagosomal compartment of J774.1 macrophages, remained alive. For this, bacteria were subjected to an intracellular viability assay [29]. As illustrated in Figure 3D and quantified (Figure 3E), the majority of the replication-deficient ΔgadC mutant bacteria (>85%) were alive after 10 h of infection.
Infection of murine bone marrow derived macrophages (BMM) with wild-type F. novicida results in activation of the AIM2 inflammasome and pyroptosis [30], [31], [32], which can be monitored by real time incorporation of the membrane-impermeable dye propidium iodide [33]. Therefore, we compared the cell death kinetics of BMM infected either with the wild-type strain or with the ΔgadC mutant. As previously shown [33], infection with wild-type F. novicida triggered macrophage death after 6–8 h, while infection with the vacuolar mutant (ΔFPI mutant) had no effect on cellular viability during the time frame of the experiment. In agreement with its inability to escape from the vacuole and to replicate within host cell, the ΔgadC mutant behaved as a ΔFPI mutant and was unable to trigger host cell death (Figure S3).
F. tularensis gadC encodes a genuine glutamate transporter
Earlier phylogenetic studies have distinguished ten distinct subfamilies within the APC family of transporters, inferring possible substrate specificities. Consensus signature motifs were defined for each of them [34]. Inspection of the Francisella GadC protein reveals a signature sequence of the Glutamate-GABA antiporter subfamily in its N-proximal portion (Figure 4A), prompting us to test functional complementation of an E. coli gadC mutant by the Francisella gadC orthologue.
Fig. 4. GadC is a glutamic acid transporter. 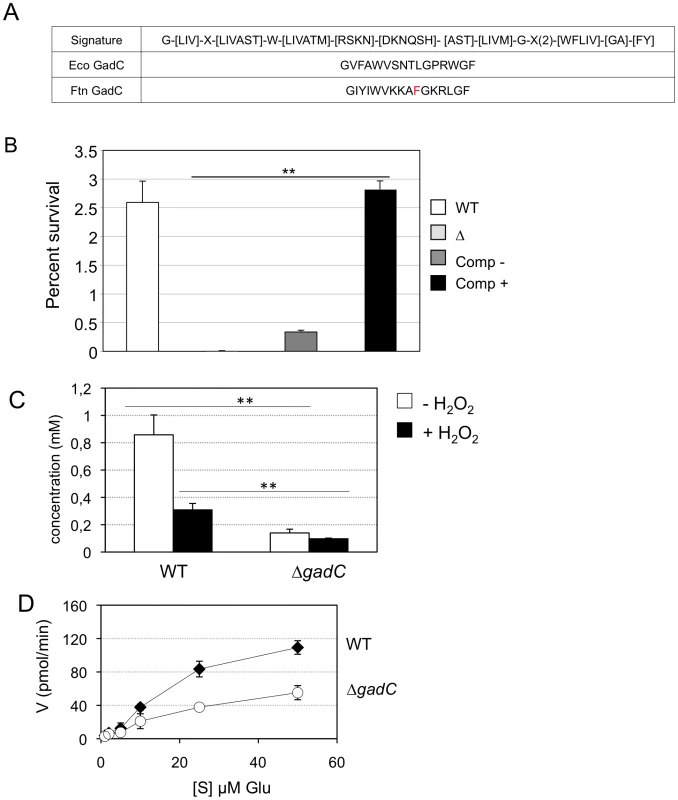
(A) The signature sequence for the Glutamate/GABA subfamily of APC transporters is shown in the upper line. Middle line, sequence of the motif present in GadC of E. coli; lower line, sequence of the motif present in GadC of Francisella (in red the only residue diverging from the consensus). (B) Functional complementation of E. coli gadC. Acid resistance assays were performed on E. coli recombinant strains. Δ: E. coli strain bearing an inactivated gadC allele. WT: complemented strain bearing the wild-type E. coli gadC gene on plasmid pCF348 [49]. Comp −: complemented strain bearing the wild-type Francisella gadC gene carried on plasmid pCR2.1-Topo and Comp +: complemented strain bearing the wild-type Francisella gadC gene carried on plasmid pCR2.1-Topo and cultivated with IPTG . **p<0.05 as determined by the Student's t-test. (C) Intracellular glutamate detection and quantification was assayed on exponentially grown bacteria by HPLC analysis. Wild-type F. novicida and ΔgadC mutant strains were grown in CDM supplemented with 1.5 mM of glutamate, in the absence or presence of H2O2 (500 µM). **, p<0.01 (as determined by the Student's t-test). (D) Glutamate transport. Kinetics of 14C-Glu uptake by wild-type F. novicida and ΔgadC mutant, at 14C-Glu concentrations ranging from 1 µM to 50 µM. Bacteria grown to mid-exponential phase in CDM were tested. Uptake was measured after 5 min incubation with 14C-Glu. Ordinate, pmol of glutamate taken up per min (per sample of app. 2.5×109 bacteria). Abscissa, final concentrations of glutamate tested. Functional complementation (Figure 4B) was determined by comparing the acid resistance (at pH 2.5) of a gadC-inactivated strain of E. coli (EF491) to the same strain carrying a plasmid-borne F. novicida gadC gene (pCRT-gadC). As a positive control, we used the E. coli gadC mutant complemented with the wild-type E. coli gadC gene (EF547). IPTG-induced expression of the Francisella gadC allele in the E. coli gadC mutant strain restored acid resistance to wild-type level, indicating that the Francisella GadC protein displays the acid-resistance function of the E. coli GadC protein.
To further support the role of GadC in glutamate entry, we quantified the amounts of intracellular glutamate by HPLC analysis, in the wild-type and ΔgadC strains grown in CDM supplemented with 1.5 mM of glutamate (in the presence or in the absence of hydrogen peroxide). As shown in Fig. 4C, the concentration of intracellular glutamate was significantly lower in the ΔgadC mutant than in the wild-type strain, both in the absence (84% reduction in concentration) or in the presence (31% reduction) of H2O2. We also quantified the amount of glutamate in culture supernatants of the two strains in the presence of H2O2 (not shown). External glutamate present in the culture medium of the wild-type strain was 39% lower than that of the ΔgadC mutant.
Altogether these data are compatible with a reduced capacity of the ΔgadC mutant to take up external glutamate.
We then directly evaluated the impact of gadC inactivation on glutamate uptake by live F. novicida. For this, we compared the uptake of radiolabeled glutamate (14C-Glu) by wild-type F. novicida to that of the ΔgadC mutant, over a broad range of glutamate concentrations (Fig. 4D). Incorporation of 14C-Glu was significantly affected in the ΔgadC mutant (representing only approximately 50% of the wild-type values at each concentration tested), confirming that GadC is a genuine glutamate transporter. The fact that glutamate uptake was not totally abolished in the ΔgadC mutant suggests that other transporter(s) allow the entry of glutamate in this strain.
The gadC mutant shows impaired control of ROS production
We compared the amount of reactive oxygen species (ROS) in J774.1 cells infected either with wild-type F. novicida, ΔgadC or the ΔFPI strain, over a 60 min period. For this, we used the H2DCF-DA assay (Sigma-Aldrich Co). H2DCF-DA is a non-fluorescent cell-permeable compound that has been widely used for the detection of ROS [35]. Once inside the cell, this compound is first cleaved by endogenous esterases to H2DCF. The de-esterified product becomes the highly fluorescent compound 2′,7′-dichlorofluorescein (DCF) upon oxidation by ROS. The ROS content increased by 25% after 60 min in cells infected with wild-type F. novicida (Figure 5). A comparable increase was recorded with the ΔFPI mutant. However, in cells infected with the ΔgadC mutant, the ROS content was significantly higher than that recorded with the two other strains at each time point (25% higher at 15 min, and 55% higher after 60 min). These results suggest that the ΔgadC mutant is affected in its ability to neutralize the production of ROS in the phagosomal compartment. Alternatively, the ΔgadC mutant may trigger an increased production of ROS.
Fig. 5. ROS dosage in infected J774.1 cells. 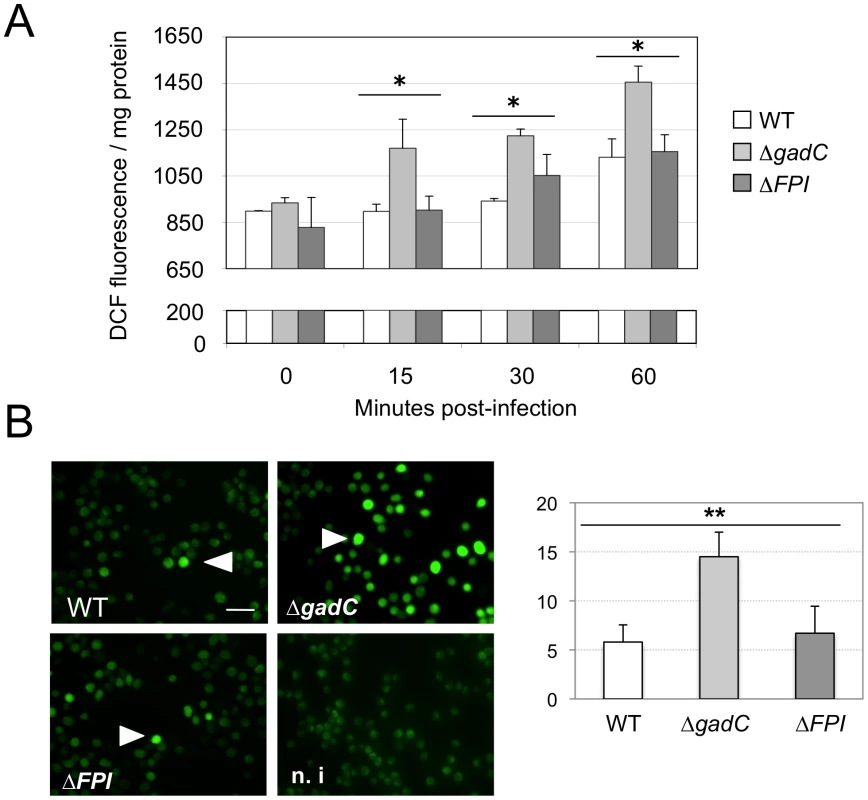
(A) ROS dosages. Generation of ROS was measured by the H2DCFDA assay in J774.1 cells infected with wild-type F. novicida (WT), the ΔgadC or the ΔFPI mutant strain. Results, normalized to the protein concentration in each well, are expressed per mg of total protein. The histogram is representative of three independent experiments. (B) Fluorescence microscopy. Left panel: DCFDA levels were also visualized using fluorescence microscopy. J774.1 cells were infected with wild-type (1), ΔgadC (2) or ΔFPI (3) bacteria. Non-infected J774.1 cells were used as negative control (4). White arrowheads indicate increased DCFDA levels. Scale bar is 50 µm. Images represent fluorescence after 1 h of DCFDA treatment. Typical fields were chosen for illustration. Right panel: Quantification of the percentage of fluorescent J774.1 cells. At least 500 cells per experiment were scored for DCFDA labeling after 1 h of DCFDA treatment. Data are means ± standard deviation from three independent assays. Impaired virulence of the gadC mutant is abrogated in NADPH oxidase KO mice
This result prompted us to evaluate the pathogenicity of the ΔgadC mutant in mice lacking a functional NADPH oxidase complex, both in vitro and in vivo.
In vitro
Intracellular replication of the ΔgadC mutant was monitored in bone marrow-derived macrophages from wild-type control mice or homozygotes gp91phox−/− in the same C57BL/6J background (designated WT and phox-KO BMMs, respectively) (Figure 6A, 6B). Multiplication of wild-type F. novicida and the ΔgadC/pKKgadC-complemented strain was similar at all time points tested in the two types of BMMs. In contrast, the ΔgadC mutant showed a significantly more severe intracellular multiplication defect in WT BMMs than in phox-KO BMMs. Indeed, the number of ΔgadC mutant bacteria was 10-fold lower than that of wild-type F. novicida already after 10 h in WT BMMs, and was 1,000-fold lower after 24 h (Fig. 6A). In BMM from phox-KO mice, multiplication of the mutant strain was 1/3 that of wild-type F. novicida after 10 h, and 100-fold higher than what was observed in WT macrophages after 24 h (Figure 6B). These results showed that multiplication of the ΔgadC mutant was mostly restored in BMMs having a defective NADPH oxidase.
Fig. 6. Intracellular survival and virulence in NADPH oxidase KO mice. 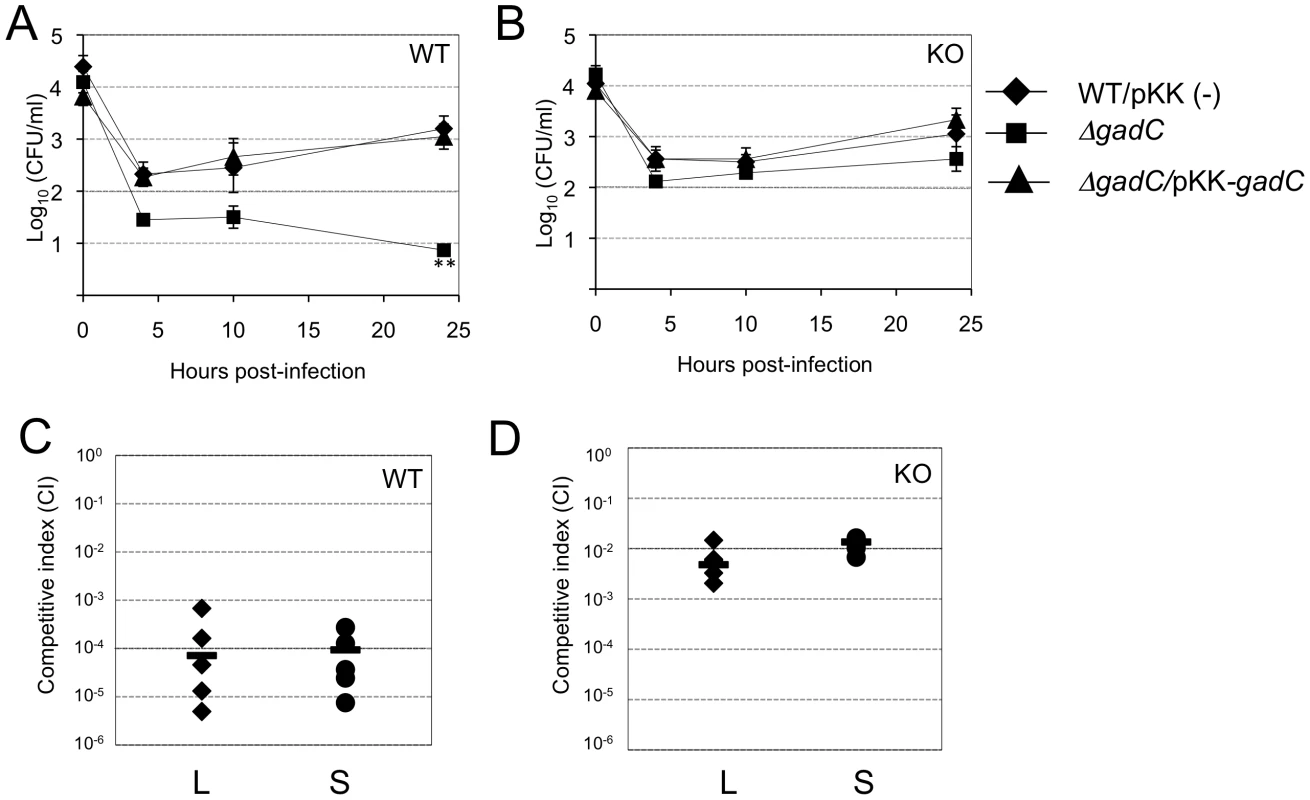
(A, B) Intracellular replication of wild-type F. novicida (carrying the empty plasmid pKK214 (WT/pKK(−)), ΔgadC mutant and complemented strain (ΔgadC/pKK-gadC), and ΔFPI mutant (ΔFPI), was monitored in BMM from either (A) C57BL/6J control mice (WT) or (B) phox-KO mice (homozygotes gp91phox−/−; KO), over a 24-h period. Results are shown as the average of log10 cfu mL−1 ± standard deviation. At all time points tested, the differences between the wild-type and ΔgadC mutant values were not statistically different (p>0.1, as determined by the Student's t-test). (C, D) Competition assays were performed by infecting intra-peritoneally: a group of five C57BL/6J control mice (WT, C); or a group of five phox-KO mice (KO, D), with a 1∶1 mixture of wild-type F. novicida and ΔgadC mutant strains (100 cfu of each). The data represent the competitive index (CI) value for cfu of mutant/wild-type in the liver (L: black diamonds, left column) and spleen (S: black circles, right column) of each mouse, 48 h after infection. Bars represent the geometric mean CI value. In vivo
We also performed in vivo competition assays in these mice (Figure 6C, 6D), as described above for the BALB/c mice. The Competition Index (CI) calculated for both target organs was 10−4 in WT mice (Figure 6C). In contrast, the CI was 100-fold higher in phox-KO mice (10−2, Figure 6D). This result comforts the data obtained in BMM cells and demonstrates that, in mice that are unable to produce ROS in the phagosome, the multiplication defect of the ΔgadC mutant is partially suppressed. However, the fact that the ΔgadC multiplication defect was not completely abolished suggests that oxidative stress may not be the only host restrictive factor.
Altogether, these in vitro and in vivo data obtained in phox-KO mice strongly suggest that GadC specifically contributes to the ROS defense of Francisella in the phagosomal compartment.
Impact of gadC inactivation in glutamate on metabolism
Intracellular glutamate plays a central role in a wide range of metabolic processes in bacteria. In order to evaluate the potential impact of the gadC inactivation on bacterial glutamate metabolism, we first quantitatively monitored the transcription of selected genes connecting glutamate utilization to either the TCA cycle or to glutathione biogenesis. This analysis was done for wild-type F. novicida and for the ΔgadC mutant strain, grown in broth with or without H2O2 (Figure 7A).
Fig. 7. Glutamate transport and metabolism. 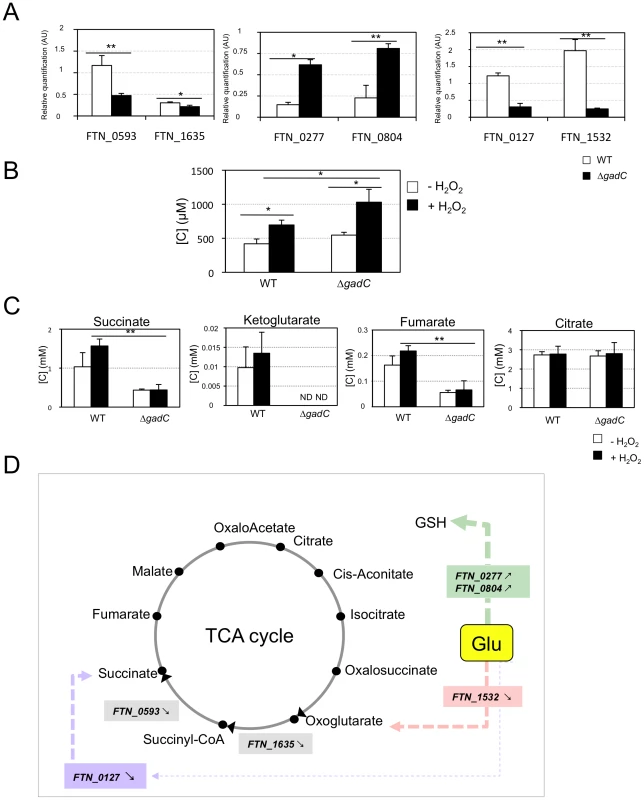
(A) qRT-PCR of metabolic genes ± H202. Bacteria were grown in TSB, in the absence or in the presence of H2O2 (500 µM). qRT-PCR analyses were performed on selected genes, in wild-type F. novicida and in the ΔgadC mutant. For each gene, the data are represented as the ratios of the value recorded under H2O2 stress versus non-stress condition. Left panel: two genes encoding enzymes of the TCA cycle (sucA, FTN_1635; sucD, FTN_0593); middle panel: two genes encoding enzymes, converting glutamate (Glu) to glutathione (GSH) (gshA, FTN_0277; gshB, FTN_0804); right panel: two genes encoding enzymes, converting Glu to TCA cycle intermediates (gdhA, FTN_1532; gabD, FTN_0127). (B) Dosage of glutathione. The effect of oxidative stress on the cytoplasmic content of glutathione was evaluated in wild-type F. novicida and ΔgadC mutant strains. Bacteria were cultivated for 30 min, with or without H2O2 (500 µM), in CDM supplemented with glutamate (1.5 mM). Reduced glutathione was quantified by HPLC analysis. Concentrations [C] are expressed in µM. *p<0.05 as determined by the Student's t-test. (C) Dosage of TCA intermediates. The effect of oxidative stress on the cytoplasmic contents of TCA cycle intermediates was monitored in wild-type F. novicida and ΔgadC mutant strains. Bacteria were cultivated for 30 min, with or without H2O2 (500 µM), in CDM supplemented with glutamate (1.5 mM). Succinate, fumarate, citrate and oxoglutarate, were quantified by gas chromatography coupled with mass spectrometry. Concentrations [C] are expressed in mM. *p<0.05, **p<0.01, as determined by the Student's t-test. (D) Schematic representation of selected genes involved in glutamate metabolism. The impact of gadC inactivation on the oxidative stress response of the target genes is indicated ( means the ratio (H2O2-treated/non-treated) is lower in the mutant strain than in the wild-type strain; ↗ the ratio (H2O2-treated/non-treated) is higher in the mutant strain than in the wild-type strain. In the absence of external glutamate (e.g. in standard chemically defined medium), the pool of glutamate present in the bacterial cytoplasm may be synthesized either from oxoglutarate, glutamine, GSH or even proline (according to KEGG metabolic pathways). Expression of FTN_0593 (sucD), FTN_0127 (gabD) and FTN_1532 (gdhA), was significantly decreased in the ΔgadC mutant under oxidative stress, whereas their expression was moderately increased in the wild-type strain. Expression of FTN_0277(gshA) and FTN_0804 (gshB) was reduced in both strains, under oxidative stress. However, the decrease was significantly less important (app. 4-fold) in the ΔgadC mutant than in the wild-type strain. Expression of FTN_1635 (sucA) was significantly decreased in both strains under oxidative stress. These data indicate that the absence of gadC affects the expression of several genes linked to glutamate metabolism under oxidative stress. The fact that expression of the gadC gene itself was significantly upregulated (approximately 10-fold) in the wild-type strain exposed to H2O2 stress (not shown) supports the importance of the GadC transporter in oxidative stress defense.
Direct quantification of TCA cycle intermediates present in the cytoplasm of the wild-type and ΔgadC strains, by gas chromatography coupled with mass spectrometry (see Materials and methods for details), revealed that gadC inactivation significantly affected succinate, fumarate, and oxoglutarate contents (Figure 7C). Indeed, in the ΔgadC mutant, the concentrations of succinate and fumarate were reduced ca. 60% as compared to the wild-type strain, whereas oxoglutarate was below the detection threshold of the assay. The concentrations of the three molecules increased up to 40% in the wild-type strain exposed to oxidative stress, suggesting an activation of the TCA under this condition. The concentrations of succinate and fumarate were not significantly modified in the ΔgadC mutant upon oxidative stress and oxoglutarate production was still below detection. The concentration of citrate was similar in the wild-type and the ΔgadC mutant and did not vary upon oxidative stress, in any of the two strains. The intracellular concentrations of glutathione were also almost similar in the wild-type and ΔgadC mutant (Figure 7B). Remarkably, under oxidative stress, the intracellular concentration of glutathione increased in both strains but only reached 65% of the level of the ΔgadC mutant in the wild-type strain.
These observations prompted us to evaluate the impact of supplementation with different TCA cycle intermediates on survival of the ΔgadC mutant in response to H2O2 challenge. For this, exponential phase wild-type and ΔgadC mutant strains, diluted in CDM supplemented with glutamate, were subjected to oxidative stress, in the presence or absence of either fumarate, succinate or oxoglutarate (Figure S5). The sensitivity to H2O2 of the ΔgadC mutant was not modified neither by fumarate nor by oxoglutarate. In contrast, supplementation with succinate increased significantly the survival of the ΔgadC mutant, to nearly wild-type level.
Discussion
Intracellular pathogenic bacteria have adapted a variety of strategies and specific intracellular niches for survival and multiplication within their host [36]. Some reside in a vacuolar compartment whereas others have evolved to gain access to the host cytosol for multiplication. In mammalian host cells, Francisella intracellular replication occurs exclusively in the cytosolic compartment. We show here that inactivation of the GadC permease of Francisella prevents phagosomal escape, thus severely altering bacterial intracellular multiplication and virulence.
The data presented suggest that the GadC protein of Francisella is required to resist to the oxidative burst triggered by the NADPH oxidase in the phagosomal compartment of infected macrophages. We propose that GadC-mediated entry of glutamate contributes to fuel the tricarboxylic acid cycle and modulates the redox status of the bacterium. This work thus provides insights into the possible links between oxidative stress resistance, metabolism, and bacterial intracellular parasitism.
GadC is involved in oxidative stress defense and required for phagosomal escape
Inactivation of gadC in F. novicida led to a severe growth defect in all cell types tested and in vivo assays further demonstrated the importance of GadC in Francisella virulence. Confocal and electron microscopy analyses revealed that the severe intracellular growth defect of the mutant was due to its inability to escape from the phagosomal compartment of infected macrophages.
Interestingly, most of the mutant bacteria that remained trapped within the phagosome were still alive for at least 10 h post-infection, indicating that impaired escape was not due to bacterial death. Since the ΔgadC mutant showed increased susceptibility to oxidative stress in broth and failed to efficiently neutralize reactive oxygen species production in cells, it is likely that ROS may predominantly affect bacterial escape rather than survival.
F. tularensis produces enzymes that can metabolize and neutralize ROS, such as a superoxide dismutases (SodB, SodC), a catalase (KatG), a glutathione peroxidase and a peroxireductase [9], [10]. Acid phosphatases have also been implicated in the resistance of intracellular Francisella to H2O2 generated in the phagosomal compartment by the NADPH oxidase ([37], [38] and references therein). However, inactivation of the major phosphatase acpA in F. tularensis subsp. tularensis, had no impact on the activity of the NADPH oxidase in human neutrophils [5], thus confirming that other Francisella factors were involved in NADPH oxidase inhibition.
We show here that Francisella GadC is an important player specifically involved in oxidative stress defense. The existence of several paralogues of both the transporter GadC and the decarboxylase GadB in some bacterial species (for example in L. monocytogenes) might account for the fact that these have not yet been found to contribute to oxidative stress resistance and intracellular survival in standard genetic screens. Indeed, if functional paralogues exist, they must be simultaneously inactivated to observe a possible phenotypic defect. In addition, isofunctional antiporters with no significant amino acid sequence similarity to the GadC protein might exist in these bacteria.
The contribution of the GAD system to intracellular survival critically depends on the cellular compartment where bacterial survive and multiply. Indeed, bacteria residing in vacuolar compartments (such as Salmonella, Mycobacteria, Legionella, Brucella and Chlamydia) encounter different types of stresses (pH, oxidative, nutritional,…) than bacteria able to multiply in the host cell cytosol (such as Francisella, Listeria, Shigella and Rickettsia).
L-glutamate is very abundant in the intracellular compartment (reported concentrations vary between 2 and 20 mM) when compared to the extracellular compartment (app. 20 µM) [39]. Human macrophages have both the cystine/glutamate transporter and the Na-dependent high-affinity glutamate transporters (excitatory amino acid transporters, EAATs) that transport glutamate and aspartate. To maintain their intracellular pool of glutamate, macrophages may use either these transporters to import glutamate from the extracellular milieu or enzymatically convert cytosolic glutamine (via glutaminase) and aspartate (via aspartate transpeptidase) to glutamate. Glutamate might also be produced spontaneously intracellularly from pyroglutamate. Currently, nothing is known with respect to the content of glutamate in the phagosomal compartment. This might prove extremely difficult to establish, especially for pathogens such as Francisella or Listeria that reside only very transiently in this compartment.
A limited number of bacterial species have been shown to possess a GAD system [40]. These include E. coli, Lactobacillus, L. monocytogenes and Shigella species, in which the GAD system plays a major role in acid tolerance. It has been suggested that the GAD system is important for pathogenic microorganisms that, upon oral infection of mammalian hosts, need to survive the low pH of the stomach. However, some enteric pathogens like Salmonella do not possess a function GAD system and must thus rely on other anti-acidic pH strategies. Interestingly, the GAD system has been also found to contribute to oxidative stress defense in yeast and plant [41]. In bacteria, molecules such as the NADPH and NADH pools and glutathione (GSH), contribute to oxidative stress defense. Reduced GSH, present at mM concentrations, maintain a strong reducing environment in the cell. Specific enzymes are also dedicated to control the levels of reactive oxygen species (ROS).
Remarkably, the ΔgadC mutant was still outcompeted by wild-type bacteria in phox-KO mice. The different environments and the immune pressure, encountered by the bacterium during its systemic dissemination, are probably far more complex than in culture systems. In vivo, Francisella GadC is thus likely to contribute to other functions than combat ROS in the phagosomal compartment. It may, for instance, fulfill classical nutritional functions during bacterial cytosolic multiplication (in macrophages and/or in other infected non-phagocytic cells). Alternatively, GadC may be required during the bacterial blood stage multiplication and dissemination of the bacterium.
A link between oxidative stress and bacterial metabolism
In E. coli, GABA produced by the glutamate decarboxylase is metabolized via the GABA shunt pathway. This leads to the production of succinate via the consecutive action of two enzymes: a GABA/oxoglutarate amino-transferase (GabT) that removes the amino group from GABA to form succinic semialdehyde (SSA) and Glu; and a succinic semialdehyde dehydrogenase (GabD) that oxidises SSA to form succinate. Very recently, Karatzas and co-workers have shown [42] that L. monocytogenes also possessed functional GabT and GabD homologues that could provide a possible route for succinate biosynthesis in L. monocytogenes. The GABA shunt pathway, allowing the bypass of two enzymatic steps of the TCA (from oxoglutarate to succinate; Figure 7 and S6), is thought to play a role in glutamate metabolism, anaplerosis and antioxidant defense. However, its physiological role in pathogenesis is yet poorly understood. Francisella genomes possess a gabD orthologue but lack gabT. The GABA shunt pathway may therefore be non-functional in Francisella. Interestingly, the isogenic glutamate decarboxylase ΔgadB mutant (Figure S4) that we constructed, showed a much less severe intracellular multiplication defect than the ΔgadC mutant, and as well as no (or only a very mild) attenuation of virulence. If the glutamate imported via GadC would serve to produce GadB-mediated GABA, one would expect gadB inactivation to cause the same defect as gadC inactivation. As already mentioned in the Introduction, the gadB orthologue encodes a truncated protein in the subspecies holarctica. Altogether, these data support the notion that GadC and GadB of Francisella do not function in concert, unlike in several other bacterial species, and that GABA production plays a marginal role in Francisella pathogenesis. Further work will be required to understand the exact contribution of GadB in Francisella metabolism.
Our data indicate that GadC of Francisella encodes a genuine glutamate transporter involved in oxidative stress, unlike most other GadC orthologues described thus far. Glutamate can be converted in the bacterial cytoplasm into a number of compounds (Figure 7), such as glutamine, glutathione, GABA or the TCA cycle intermediate oxoglutarate. Oxoglutarate is known to be a potent anti-oxidant molecule that can be converted, in absence of any enzymatic reaction, into succinate in the presence of H2O2. In addition, conversion of glutamate to oxoglutarate by the glutamate dehydrogenase GdhA increases the production of NADPH, which might also contribute to the anti-oxidant effect of glutamate acquisition.
Quantitative analyses of the intra-bacterial content of TCA cycle intermediates (Figure 7B) revealed a significant reduction of succinate and fumarate in the gadC mutant, as compared to wild-type F. novicida, and a striking decrease of oxoglutarate. These data support the notion that reduced entry of glutamate directly affects the production of these TCA cycle intermediates. In contrast, the amount of citrate remained unchanged in the mutant, suggesting refueling of the TCA cycle via other entry points (such as glycolysis or amino acid conversion).
Of note, mutants in genes gdhA (FTN_1533) and gabD (FTN_0127) were identified as required for replication in D. melanogaster S2 cells in a recent screen, supporting a role for these genes in intracellular bacterial survival [43]. The production and utilization of oxoglutarate by Francisella may thus constitute an efficient mean to modulate its cytoplasmic concentration of ROS.
In the absence of external glutamate, the pool of intracellular glutamate may be synthesized either from oxoglutarate, glutamine, GSH or even proline (according to KEGG metabolic pathways). Therefore, we evaluated the impact of gadC inactivation on the expression of genes involved in glutamate metabolism, under oxidative stress conditions. qRT-PCR analyses were performed in wild-type F. novicida and in the ΔgadC mutant, grown in chemically defined medium containing glutamate, in the absence or in the presence of H2O2 (Figure 7A). These assays revealed that gadC inactivation led to an important down-regulation of the genes involved the conversion of glutamate to oxoglutarate and succinate, upon oxidative stress (FTN_1532 and FTN_0127, respectively). Conversely, gadC inactivation only moderately decreased the expression of gshA and gshB, the two genes involved in glutathione biosynthesis (FTN_0277 and FTN_0804), upon oxidative stress whereas the expression of these genes was severely decreased in the wild-type strain.
These data are compatible with the notion that, under oxidative stress, the wild-type strain may favor the conversion of a fraction of its cytoplasmic pool of glutamate (neosynthesized and imported) to produce oxoglutarate and succinate rather than GSH. In contrast, when the cytosolic pool of glutamate is restricted to neosynthesized glutamate (i.e. in a gadC mutant or in a glutamate-depleted medium), the production of oxaloglutarate and succinate may be decreased to favor that of other molecules (including GSH).
In conclusion, we identified a glutamate transporter as a novel Francisella virulence attribute that suggests links between the oxidative stress response and the TCA cycle during the early stage of the bacterial intracellular life cycle. The importance of the TCA cycle in the homeostasis of reactive oxygen species has just started to be considered in pathogenic bacterial species [12], [44], [45], [46]. The development of specific inhibitors of transport systems involved in intracellular adaptation might constitute interesting anti-bacterial therapeutic targets.
Materials and Methods
Ethics statement
All experimental procedures involving animals were conducted in accordance with guidelines established by the French and European regulations for the care and use of laboratory animals (Decree 87–848, 2001–464, 2001–486 and 2001–131 and European Directive 2010/63/UE) and approved by the INSERM Ethics Committee (Authorization Number: 75-906).
Bacterial strains, media, and chemicals
F. tularensis subsp. novicida (F. novicida) strain U112, its ΔFPI derivative, and all the mutant strains constructed in this work, were grown as described in Supplementary Material. E. coli strains (kindly provided by John Foster, University of South Alabama, USA) were grown as described in Supplementary Material. All bacterial strains, plasmids, and primers used in this study are listed in Supplemental Table 1.
Details of the construction and characterization of mutant and complemented strains; macrophage preparation and infections, are described in Supplementary Material. Quantitative (q)RT-PCR (real-time PCR) was performed with gene-specific primers (Supplemental Table 1), using an ABI PRISM 7700 and SYBR green PCR master mix (Applied Biosystems, Foster city, CA, USA).
Electron and confocal microscopy complete descriptions; real time cell death and phagosome permeabilization assays, are described in Supplementary Material.
Multiplication in macrophages
J774.1 macrophage-like cells (ATCC Number: TIB-67) were propagated in Dulbecco's Modified Eagle's Medium (DMEM) containing 10% fetal calf serum, whereas human monocyte-like cell line THP-1 (ATCC Number: TIB-202) and bone marrow-derived macrophages (BMM) from BALB/c were propagated in RPMI Medium 1640 containing 10% fetal calf serum, respectively. J774.1 and BMM were seeded at a concentration of ∼2×105 cells per well in 12-well cell tissue plates and monolayers were used 24 h after seeding. THP-1 were seeded at a concentration of ∼2×105 cells per well in 12-well cell tissue plates 48 h before infection, and supplemented with phorbol myristate acetate (PMA) to induce cell differentiation (200 ng/ml). J774.1, BMM and THP-1 were incubated for 60 min at 37°C with the bacterial suspensions (approximately multiplicities of infection 100) to allow the bacteria to enter. After washing (time zero of the kinetic analysis), the cells were incubated in fresh culture medium containing gentamicin (10 µg mL−1) to kill extracellular bacteria. At several time-points, cells were washed three times in DMEM or RPMI, macrophages were lysed by addition of water and the titer of viable bacteria released from the cells was determined by spreading preparations on Chocolate agar plates. For each strain and time in an experiment, the assay was performed in triplicate. Each experiment was independently repeated at least three times and the data presented originate from one typical experiment.
Isolation of total RNA and reverse transcription
Bacteria were centrifuged for 2 min in a microcentrifuge at room temperature and the pellet was quickly re-suspended in Trizol solution (Invitrogen, Carlsbad, CA, USA). Samples were either processed immediately or frozen and stored at −80°C. Samples were treated with chloroform and the aqueous phase was used in the RNeasy Clean-up protocol (Qiagen, Valencia, CA, USA) with an on-column DNase digestion of 30 min [47].
RNA Reverse transcription (RT)-PCR experiments were carried out with 500 ng of RNA and 2 pmol of specific reverse primers. After denaturation at 65°C for 5 min, 6 µL of the mixture containing 4 µL of 5× first strand buffer and 2 µL of 0,1 M DTT were added. Samples were incubated 2 min at 42°C and, then, 1 µL of Superscript II RT (Thermo Scientific) was added. Samples were incubated for 50 min at 42°C, heated at 70°C for 15 min and chilled on ice. Samples were diluted with 180 µL of H2O and stored at −20°C.
The following pair of primers was used to amplify the mRNA corresponding to the transcript of FTN_0570 (p13/p14), FTN_0571 (p15/p16), FTN_1700 (p27/9p28), FTN_1701 (p29/p30), FTN_1702 (p31/p32), FTN_1532 (p33/p34), FTN_0127 (p35/p36), FTN_0277 (p37/p38), FTN_0804 (p39/p40), FTN_0593 (p41/p42), FTN_1434 (p43/p44) and FTN_1635 (p45/p46) (Supplemental Table 1).
Quantitative real-time RT-PCR
Wild-type F. novicida and mutant strains were grown at 37°C from OD600 ∼0.1. After 4 h of incubation, samples were harvested and RNA was isolated. For oxidative stress tests, samples were cultivated 30 min more with or without H2O2 (500 µM). The 25 µL reaction consisted of 5 µL of cDNA template, 12.5 µL of Fastart SYBR Green Master (Roche Diagnostics), 2 µL of 10 µM of each primer and 3.5 µL of water. qRT-PCR was performed according manufacturer's protocol on Applied Biosystems - ABI PRISM 7700 instrument (Applied Biosystems, Foster City, CA). To calculate the amount of gene-specific transcript, a standard curve was plotted for each primer set using a series of diluted genomic DNA from wild-type F. novicida. The amounts of each transcript were normalized to helicase rates (FTN_1594).
Oxidative and pH stress survival assays
Stationary-phase bacterial cultures were diluted at a final OD600 of 0.1 in TSB broth or CDM with or without glutamate (1.5 mM final). Exponential-phase bacterial cultures were diluted to a final concentration of 108 bacteria mL−1 and subjected to either 500 µM H2O2 or pH 5.5.
Oxidative stress response was also tested in CDM supplemented with glutamate, in the presence or absence of the TCA cycle intermediates: oxoglutarate, succinate or fumarate (1.5 mM final). The number of viable bacteria was determined by plating appropriate dilutions of bacterial cultures on Chocolate Polyvitex plates at the start of the experiment and after the indicated durations. Cultures (5 mL) were incubated at 37°C with rotation (100 rpm) and aliquots were removed at indicated times, serially diluted and plated immediately. Bacteria were enumerated after 48 h incubation at 37°C. Experiments were repeated independently at least twice and data represent the average of all experiments.
Confocal experiments
J774.1 cells were infected with wild-type F. novicida, ΔgadC or ΔFPI strains for 1 h, 4 h and 10 h at 37°C, and were washed in KHM (110 mM potassium acetate, 20 mM Hepes, 2 mM MgCl2). Cells were incubated for 1 min with digitonin (50 µg/mL) to permeabilize membranes. Then cells were incubated for 15 min at 37°C with primary anti F. novicida mouse monoclonal antibody (1/500 final dilution, Immunoprecise). After washing, cells were incubated for 15 min at 37°C with secondary antibody (Ab) (Alexa Fluor 488-labeled GAM, 1/400 final dilution, Abcam) in the dark. After washing, cells were fixed with PFA 4% for 15 min at room temperature (RT) and incubated for 10 min at RT with 50 mM NH4Cl to quench residual aldehydes. After washing with PBS, cells were incubated for 30 min at RT with primary anti-LAMP1 Ab (1/100 final dilution, Abcam) in a mix with PBS, 0.1% saponine and 5% goat serum. After washing with PBS, cells were incubated for 30 min at RT with secondary anti-rabbit Ab (alexa 546-labeled, 1/400 dilution, Abcam). DAPI was added (1/5,000 final dilution) for 1 min. After washing, the glass coverslips were mounted in Mowiol. Cells were examined using an X63 oil-immersion objective on a LeicaTSP SP5 confocal microscope. Co-localization tests were quantified by using Image J software; and mean numbers were calculated on more then 500 cells for each condition. Confocal microscopy analyses were performed at the Cell Imaging Facility (Faculté de Médecine Necker Enfants-Malades).
Electron microscopy
Infection of J774.1 cells was followed by thin section electron microscopy as previously described [48].
Determination of intracellular bacterial viability
To evaluate the viability of F. tularensis, labelings were adapted to use the cell-impermeant nucleic acid dye propidium iodide (PI). J774.1 macrophage-like cells were seeded at 5.105cells/ml on glass coverslips in 12-well bottom flat plates. Next day, cells were infected for 10 h with wild-type F. novicida or ΔgadC strain. After infection, cells were first permeabilized with digitonin for 1 min, washed three times with KHM and incubated for 12 min at 37°C with 2.6 µM PI (Life technologies, L7007) in KHM buffer to label compromised bacteria in permeabilized cells. Cells were washed three times with KHM and incubated for 15 min at 37°C with primary anti F. novicida mouse monoclonal antibody (1/500 final dilution). After washing, cells were incubated for 15 min at 37°C with secondary antibody (Ab) (Alexa Fluor 488-labeled GAM, 1/400 final dilution) in the dark. After washing, cells were fixed with PFA 4% for 15 min at room temperature (RT) and incubated for 10 min at RT with 50 mM NH4Cl to quench residual aldehydes. After washing with PBS, cells were incubated for 30 min at RT with primary anti-LAMP1 Ab (1/100 final dilution) in a mix with PBS, 0.1% saponine and 5% goat serum. After washing with PBS, cells were incubated for 30 min at RT with secondary anti-rabbit Ab (alexa 405-labeled, 1/400 dilution). After washing, the glass coverslips were mounted in Mowiol. Cells were examined using an X63 oil-immersion objective on a LeicaTSP SP5 confocal microscope. Analysis of cell fluorescence was performed with Image J software (http://rsb.info.nih.gov/ij).
ROS detection assay
Intracellular reactive oxygen species (ROS) were detected by using the oxidation-sensitive fluorescent probe dye, 2′,7′-dichlorodihydrofluorescein diacetate (H2DCF-DA) as recommended by the manufacturer (CM-H2DCF-DA, Molecular Probes, Eugene, OR). J774.1 cells were seeded at 5.104 cells/well. Cells were infected with bacteria for 15 min (MOI of 100∶1), washed three times with PBS and incubated with H2DCF-DA diluted in PBS (concentration). DCF fluorescence was measured with the Victor2 D fluorometer (Perkin-Elmer, Norwal, CT) with the use of excitation and emission wavelengths of 480 nm and 525 nm, respectively. Values were normalized by protein concentration in each well (Bradford). Samples were tested in triplicates in three experiments.
Determination of ROS generation via fluorescent microscopy
J774.1 cells were seeded at 5.104 cells/well. Cells were infected with bacteria for 15 min (MOI of 1,000∶1), washed three times with PBS and incubated with H2DCF-DA diluted in PBS for 1 h (5 µM). Images of the cells were captured with an Olympus CKX41 microscope and treated with Image J software. Cell counts were performed over 10 images of approximately 50 cells.
Acid resistance assay
Acid resistance tests in E. coli were performed at pH 2.5 as described previously [49], by comparing the number of survival treated cells after 1 h of treatment over the number of cells at T0. We compared survival of wild-type E. coli strain (WT) with E. coliΔgadC (Δ) and E. coliΔgadC complemented with the F. novicida gadC gene (PCR-amplified gene FTN-0571 introduced into plasmid pCR2.1-Topo, in the correct orientation downstream of the plac promoter) (Comp). Acid challenge was performed by diluting 1∶100 the overnight (22 h) culture in LB, supplemented (Comp +) or not (Comp −) with 10−4 M final IPTG.
Glutamate assays
Glutamate detection and quantification was done by using HPLC analysis. Wild-type F. novicida and ΔgadC strains were tested in CDM supplemented with 1.5 mM of glutamate, with or without H2O2 (500 µM). For each condition, three independent cultures were prepared by overnight growth in CDM. The overnight cultures were diluted with 50 mL of fresh medium to OD600 of 0.05 and cultivated to an OD600 of app. 0.35. Bacteria were harvested by centrifugation at 4,000× g for 20 min, resuspended in 25 mL of pre-warmed appropriated medium and cultivated for 30 min. For extracellular glutamate dosage, 100 µL of each supernatant were resuspended with 400 µL of cold methanol and centrifuged at 12,000× g for 5 min. 20 µL of each preparation were derivatized with 80 µL of OPA. For intracellular dosage, each sample were resuspended in 600 µL of cold methanol. The bacterial suspensions were sonicated thrice for 30 sec at 4.0 output, 70% pulsed (Branson Sonifier 250). Lysates were then centrifuged at 3,300× g for 8 min, to remove debris.
Following steps were done with the standard procedure of Agilent using ZORBAX Eclipse AAA high as HPLC column. An amount equivalent to 2 µL of each sample was injected on a Zorbax Eclipse-AAA column, 5 µm, 150×4.6 mm (Agilent), at 40°C, with fluorescence detection. Aqueous mobile phase was 40 mM NaH2PO4, adjusted to pH 7.8 with NaOH, while organic mobile phase was acetonitrile/methanol/water (45/45/10 v/v/v). The separation was obtained at a flow rate of 2 mL min−1 with a gradient program that allowed for 2 min at 0% B followed by a 16-min step that raised eluent B to 60%. Then washing at 100% B and equilibration at 0% B was performed in a total analysis time of 38 min. To evaluate glutamate concentration, glutamate standard curve was made in parallel.
Glutathione assays
The procedure for the measurement of GSH was previously described [50]. Briefly, GSH were separated by HPLC, equipped with a Shimadzu Prominence solvent delivery system (Shimadzu Corp., Kyoto, Japan), using a reverse-phase C18 Kromasil (5 µm; 4.6×250 mm), obtained from AIT (Paris, Fr). The mobile phase for isocratic elution consisted of 25 mmol L−1 monobasic sodium phosphate, 0.3 mmol L−1 of the ion-pairing agent 1-octane sulfonic acid, 4% (v/v) acetonitrile, pH 2.7, adjusted with 85% phosphoric acid. The flow rate was 1 mL min−1. Under these conditions, the separation of aminothiol was completed in 20 min. Deproteinated samples were injected directly onto the column using a Shimadzu autosampler (Shimadzu Corp.). Following HPLC separation, GSH was detected with a model 2465 electrochemical detector (Waters, MA, USA) equipped with a 2 mm Glassy carbon (GC) analytical cell and potential of +700 mV were applied.
Glutamate transport assays
Cells were grown in Chamberlain medium to mid-exponential phase and then harvested by centrifugation and washed twice with Chamberlain without amino acid. The cells were suspended at a final OD600 of 0.5 in the same medium containing 50 mg/ml of chloramphenicol. After 15 min of pre-incubation at 25°C, uptake was started by the addition of L-[U-14C] glutamic acid (Perkin Elmer), at various concentrations (14C-Glu ranging from 1 to 50 µM). The radiolabeled 14C-Glu was at a specific activity of 9.25 GBq.mmol −1. Samples (100 µL of bacterial suspension) were removed after 5 min and collected by vacuum filtration on membrane filters (Millipore type HA, 25 mm, 0.22 mm) and rapidly washed with Chamberlain without amino acid (2×5 mL). The filters were transferred to scintillation vials and counted in a Hidex 300 scintillation counter. The counts per minute (c.p.m.) were converted to picomoles of amino acid taken up per sample, using a standard derived by counting a known quantity of the same isotope under similar conditions.
Quantification of TCA cycle intermediates
Succinate, fumarate, citrate and oxoglutarate were quantified by gas chromatography coupled with mass spectrometry (GC/MS). Wild-type F. novicida and ΔgadC strains were grown as for glutamate quantification. Briefly, after an overnight preculture in CDM, three independent cultures of wild-type and ΔgadC mutant were cultivated in 50 mL CDM to an OD600 of app. 0.35. Bacteria were harvested by centrifugation at 4,000× g for 15 min, resuspended to the same OD600 in pre-warmed CDM supplemented with glutamate (1.5 mM) and cultivated for 30 min±500 µM H2O2.
Metabolite measurements were normalized by checking that each sample contained equal amounts of total proteins.
Protein dosage
After centrifugation at 4,000× g for 20 min, each sample was resuspended in 2 mL of ice-cold 5 mM Tris.HCl (pH8). The bacterial suspensions were sonicated thrice for 30 sec at 4.0 output, 70% pulsed (Branson Sonifier 25). Lysates (three per strain per condition) were then centrifuged at 3,300× g for 8 min, to remove debris. Protein concentration of the different samples was determined by the BCA assay (Pierce), following the manufacturer's recommendation. The values recorded for the wild-type and the ΔgadC mutant, in the absence or in the presence of H2O2, were not significantly different (1,000 mg mL−1±10 mg mL−1; p value≥0.5). The assay was repeated twice.
Metabolite measurements
After oxidative stress, bacterial samples were centrifugated at 4,000× g for 20 min, resuspended in 600 µl of cold methanol. The bacterial suspensions were sonicated thrice for 30 sec at 4.0 output, 70% pulsed (Branson Sonifier 250). Lysates were then centrifuged at 3,300× g for 8 min, to remove debris. Then, samples were purified through SPE column (Strata-X, 30 mg mL−1, Phenomenex, California, USA). After elution, complete lyophilization and derivatization with methoxymation and syliation steps, 1 µl of each sample were injected on GC column coupled to detection mass. Analyses were performed on Shimadzu GC2/MS-2010 (Columbia, MD). Data represent the average of three independent cultures for each condition.
Virulence determination
Wild-type F. novicida and ΔgadC mutant strains were grown in TSB to exponential growth phase and diluted to the appropriate concentrations. 6 to 8-week-old female BALB/c mice (Janvier, Le Genest St Isle, France) were intra-peritoneally (i.p.) inoculated with 200 µl of bacterial suspension. The actual number of viable bacteria in the inoculum was determined by plating appropriate dilutions of bacterial cultures on Chocolate Polyvitex plates. For competitive infections, wild-type F. novicida and mutant bacteria were mixed in 1∶1 ratio and a total of 100 bacteria were used for infection of each of five mice. After two days, mice were sacrificed. Homogenized spleen and liver tissue from the five mice in one experiment were mixed, diluted and spread on to chocolate agar plates. Kanamycin selection to distinguish wild-type and mutant bacteria were performed. Competitive index (CI) [(mutant output/WT output)/(mutant input/WT input)]. Statistical analysis for CI experiments was as described in [51]. Macrophage experiments were analyzed by using the Student's t-test.
Supporting Information
Zdroje
1. Sjostedt A, editor (2011) Francisella tularensis and tularemia: Fontiers Media SA.
2. OystonPC, SjostedtA, TitballRW (2004) Tularaemia: bioterrorism defence renews interest in Francisella tularensis. Nat Rev Microbiol 2 : 967–978.
3. KeimP, JohanssonA, WagnerDM (2007) Molecular epidemiolgy, evolution, and ecology of Francisella. Ann N Y Acad Sci 1105 : 30–66.
4. CelliJ, ZahrtTC (2013) Mechanisms of Francisella tularensis intracellular pathogenesis. Cold Spring Harb Perspect Med 3: a010314.
5. McCaffreyRL, SchwartzJT, LindemannSR, MorelandJG, BuchanBW, et al. (2010) Multiple mechanisms of NADPH oxidase inhibition by type A and type B Francisella tularensis. J Leukoc Biol 88 : 791–805.
6. MeibomKL, CharbitA (2010) The unraveling panoply of Francisella tularensis virulence attributes. Curr Opin Microbiol 13 : 11–17.
7. NapierBA, MeyerL, BinaJE, MillerMA, SjostedtA, et al. (2012) Link between intraphagosomal biotin and rapid phagosomal escape in Francisella. Proc Natl Acad Sci USA 109 : 18084–18089.
8. MelilloAA, BakshiCS, MelendezJA (2010) Francisella tularensis antioxidants harness reactive oxygen species to restrict macrophage signaling and cytokine production. J Biol Chem 285 : 27553–27560.
9. LindgrenH, ShenH, ZingmarkC, GolovliovI, ConlanW, et al. (2007) Resistance of Francisella strains against reactive nitrogen and oxygen species with special reference to the role of KatG. Infect Immun 75 : 1303–1309.
10. MelilloAA, MahawarM, SellatiTJ, MalikM, MetzgerDW, et al. (2009) Identification of Francisella tularensis live vaccine strain CuZn superoxide dismutase as critical for resistance to extracellularly generated reactive oxygen species. J Bacteriol 191 : 6447–6456.
11. BakshiCS, MalikM, ReganK, MelendezJA, MetzgerDW, et al. (2006) Superoxide dismutase B gene (sodB)-deficient mutants of Francisella tularensis demonstrate hypersensitivity to oxidative stress and attenuated virulence. J Bacteriol 188 : 6443–6448.
12. DieppedaleJ, GesbertG, RamondE, ChhuonC, DubailI, et al. (2013) Possible links between stress defense and the tricarboxylic acid cycle in Francisella pathogenesis. Mol Cell Proteomics 12 : 2278–2292.
13. ClemensDL, LeeBY, HorwitzMA (2004) Virulent and avirulent strains of Francisella tularensis prevent acidification and maturation of their phagosomes and escape into the cytoplasm in human macrophages. Infect Immun 72 : 3204–3217.
14. ClemensDL, LeeBY, HorwitzMA (2009) Francisella tularensis phagosomal escape does not require acidification of the phagosome. Infect Immun 77 : 1757–1773.
15. SanticM, AsareR, SkrobonjaI, JonesS, Abu KwaikY (2008) Acquisition of the vacuolar ATPase proton pump and phagosome acidification are essential for escape of Francisella tularensis into the macrophage cytosol. Infect Immun 76 : 2671–2677.
16. De BiaseD, PennacchiettiE (2012) Glutamate decarboxylase-dependent acid resistance in orally acquired bacteria: function, distribution and biomedical implications of the gadBC operon. Mol Microbiol 86 (4) 770–86.
17. FosterJW (2004) Escherichia coli acid resistance: tales of an amateur acidophile. Nat Rev Microbiol 2 : 898–907.
18. KaratzasKA, BrennanO, HeavinS, MorrisseyJ, O'ByrneCP (2010) Intracellular accumulation of high levels of gamma-aminobutyrate by Listeria monocytogenes 10403S in response to low pH: uncoupling of gamma-aminobutyrate synthesis from efflux in a chemically defined medium. Appl Environ Microbiol 76 : 3529–3537.
19. MeibomKL, CharbitA (2010) Francisella tularensis metabolism and its relation to virulence. Front Microbiol 1 : 140.
20. GesbertG, RamondE, RigardM, FrapyE, DupuisM, et al. (2013) Asparagine assimilation is critical for intracellular replication and dissemination of Francisella. Cell Microbiol [epub ahead of print].
21. MaierTM, CaseyMS, BeckerRH, DorseyCW, GlassEM, et al. (2007) Identification of Francisella tularensis Himar1-based transposon mutants defective for replication in macrophages. Infect Immun 75 : 5376–5389.
22. WeissDS, BrotckeA, HenryT, MargolisJJ, ChanK, et al. (2007) In vivo negative selection screen identifies genes required for Francisella virulence. Proc Natl Acad Sci USA 104 : 6037–6042.
23. PengK, MonackDM (2010) Indoleamine 2,3-dioxygenase 1 is a lung-specific innate immune defense mechanism that inhibits growth of Francisella tularensis tryptophan auxotrophs. Infect Immun 78 : 2723–2733.
24. KraemerPS, MitchellA, PelletierMR, GallagherLA, WasnickM, et al. (2009) Genome-wide screen in Francisella novicida for genes required for pulmonary and systemic infection in mice. Infect Immun 77 : 232–244.
25. MaD, LuP, YanC, FanC, YinP, et al. (2012) Structure and mechanism of a glutamate-GABA antiporter. Nature 483 : 632–636.
26. LaurianoCM, BarkerJR, NanoFE, ArulanandamBP, KloseKE (2003) Allelic exchange in Francisella tularensis using PCR products. FEMS Microbiol Lett 229 : 195–202.
27. ChamberlainRE (1965) Evaluation of Live Tularemia Vaccine Prepared in a Chemically Defined Medium. Appl Microbiol 13 : 232–235.
28. BarelM, MeibomK, CharbitA (2010) Nucleolin, a shuttle protein promoting infection of human monocytes by Francisella tularensis. PLoS One 5: e14193.
29. ChongA, WehrlyT, ChildR, HansenB, HwangS, et al. (2012) Cytosolic clearance of replication-deficient mutants reveals Francisella tularensis interactions with the autophagic pathway. Autophagy 8 : 1342–1356.
30. Fernandes-AlnemriT, YuJW, JulianaC, SolorzanoL, KangS, et al. (2010) The AIM2 inflammasome is critical for innate immunity to Francisella tularensis. Nat Immunol 11 : 385–393.
31. JonesJW, BrozP, MonackDM (2011) Innate immune recognition of Francisella tularensis: activation of type-I interferons and the inflammasome. Front Microbiol 2 : 16.
32. RathinamVA, JiangZ, WaggonerSN, SharmaS, ColeLE, et al. (2010) The AIM2 inflammasome is essential for host defense against cytosolic bacteria and DNA viruses. Nat Immunol 11 : 395–402.
33. PieriniR, JurujC, PerretM, JonesCL, MangeotP, et al. (2012) AIM2/ASC triggers caspase-8-dependent apoptosis in Francisella-infected caspase-1-deficient macrophages. Cell Death Differ 19 : 1709–1721.
34. JackDL, PaulsenIT, SaierMH (2000) The amino acid/polyamine/organocation (APC) superfamily of transporters specific for amino acids, polyamines and organocations. Microbiology 146 (Pt 8) 1797–1814.
35. TakanashiT, OguraY, TaguchiH, HashizoeM, HondaY (1997) Fluorophotometric quantitation of oxidative stress in the retina in vivo. Invest Ophthalmol Vis Sci 38 : 2721–2728.
36. KwaikYA, BumannD (2013) Microbial Quest for Food in vivo: “Nutritional virulence” as an emerging paradigm. Cell Microbiol 15 : 882–890.
37. MohapatraNP, SoniS, ReillyTJ, LiuJ, KloseKE, et al. (2008) Combined deletion of four Francisella novicida acid phosphatases attenuates virulence and macrophage vacuolar escape. Infect Immun 76 : 3690–3699.
38. MohapatraNP, SoniS, RajaramMV, StrandbergKL, GunnJS (2013) Type A Francisella tularensis Acid Phosphatases Contribute to Pathogenesis. PLoS One 8: e56834.
39. NewsholmeP, ProcopioJ, LimaMM, Pithon-CuriTC, CuriR (2003) Glutamine and glutamate–their central role in cell metabolism and function. Cell Biochem Funct 21 : 1–9.
40. FeehilyC, KaratzasKA (2012) Role of glutamate metabolism in bacterial responses towards acid and other stresses. J Appl Microbiol 114 : 11–24.
41. CaoJ, BarbosaJM, SinghNK, LocyRD (2013) GABA shunt mediates thermotolerance in Saccharomyces cerevisiae by reducing reactive oxygen production. Yeast 30 : 129–144.
42. KaratzasKA, SuurL, O'ByrneCP (2012) Characterization of the intracellular glutamate decarboxylase system: analysis of its function, transcription, and role in the acid resistance of various strains of Listeria monocytogenes. Appl Environ Microbiol 78 : 3571–3579.
43. AsareR, AkimanaC, JonesS, Abu KwaikY (2010) Molecular bases of proliferation of Francisella tularensis in arthropod vectors. Environ Microbiol 12 : 2587–2612.
44. MaillouxRJ, BeriaultR, LemireJ, SinghR, ChenierDR, et al. (2007) The tricarboxylic acid cycle, an ancient metabolic network with a novel twist. PLoS One 2: e690.
45. EohH, RheeKY (2013) Multifunctional essentiality of succinate metabolism in adaptation to hypoxia in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A 110 : 6554–6559.
46. RichardsonAR, PayneEC, YoungerN, KarlinseyJE, ThomasVC, et al. (2011) Multiple targets of nitric oxide in the tricarboxylic acid cycle of Salmonella enterica serovar typhimurium. Cell Host Microbe 10 : 33–43.
47. ThompsonLJ, MerrellDS, NeilanBA, MitchellH, LeeA, et al. (2003) Gene expression profiling of Helicobacter pylori reveals a growth-phase-dependent switch in virulence gene expression. Infect Immun 71 : 2643–2655.
48. AlkhuderK, MeibomKL, DubailI, DupuisM, CharbitA (2009) Glutathione provides a source of cysteine essential for intracellular multiplication of Francisella tularensis. PLoS Pathog 5: e1000284.
49. Castanie-CornetMP, PenfoundTA, SmithD, ElliottJF, FosterJW (1999) Control of acid resistance in Escherichia coli. J Bacteriol 181 : 3525–3535.
50. RebrinI, KamzalovS, SohalRS (2003) Effects of age and caloric restriction on glutathione redox state in mice. Free Radic Biol Med 35 : 626–635.
51. BrotckeA, WeissDS, KimCC, ChainP, MalfattiS, et al. (2006) Identification of MglA-regulated genes reveals novel virulence factors in Francisella tularensis. Infect Immun 74 : 6642–6655.
52. CowingDW, BardwellJC, CraigEA, WoolfordC, HendrixRW, et al. (1985) Consensus sequence for Escherichia coli heat shock gene promoters. Proc Natl Acad Sci U S A 82 : 2679–2683.
Štítky
Hygiena a epidemiológia Infekčné lekárstvo Laboratórium
Článek Serotonin Signaling in : A Serotonin–Activated G Protein-Coupled Receptor Controls Parasite MovementČlánek Regulators of Cell Cycle Progression and Differentiation Identified Using a Kinome-Wide RNAi ScreenČlánek IFNγ/IL-10 Co-producing Cells Dominate the CD4 Response to Malaria in Highly Exposed ChildrenČlánek Functions of CPSF6 for HIV-1 as Revealed by HIV-1 Capsid Evolution in HLA-B27-Positive SubjectsČlánek Decreases in Colonic and Systemic Inflammation in Chronic HIV Infection after IL-7 Administration
Článok vyšiel v časopisePLOS Pathogens
Najčítanejšie tento týždeň
2014 Číslo 1- Parazitičtí červi v terapii Crohnovy choroby a dalších zánětlivých autoimunitních onemocnění
- Očkování proti virové hemoragické horečce Ebola experimentální vakcínou rVSVDG-ZEBOV-GP
- Koronavirus hýbe světem: Víte jak se chránit a jak postupovat v případě podezření?
-
Všetky články tohto čísla
- HIV-1 Accessory Proteins Adapt Cellular Adaptors to Facilitate Immune Evasion
- Ranaviruses: Not Just for Frogs
- Effectors and Effector Delivery in
- Plasmacytoid Dendritic Cell Dynamics Tune Interferon-Alfa Production in SIV-Infected Cynomolgus Macaques
- Lu/BCAM Adhesion Glycoprotein Is a Receptor for Cytotoxic Necrotizing Factor 1 (CNF1)
- A Substrate-Fusion Protein Is Trapped inside the Type III Secretion System Channel in
- Parvovirus-Induced Depletion of Cyclin B1 Prevents Mitotic Entry of Infected Cells
- Red Blood Cell Invasion by : Structural Basis for DBP Engagement of DARC
- NsrR, GadE, and GadX Interplay in Repressing Expression of the O157:H7 LEE Pathogenicity Island in Response to Nitric Oxide
- Loss of Circulating CD4 T Cells with B Cell Helper Function during Chronic HIV Infection
- TREM-1 Deficiency Can Attenuate Disease Severity without Affecting Pathogen Clearance
- Origin, Migration Routes and Worldwide Population Genetic Structure of the Wheat Yellow Rust Pathogen f.sp.
- Glutamate Utilization Couples Oxidative Stress Defense and the Tricarboxylic Acid Cycle in Phagosomal Escape
- Serotonin Signaling in : A Serotonin–Activated G Protein-Coupled Receptor Controls Parasite Movement
- Recovery of an Antiviral Antibody Response following Attrition Caused by Unrelated Infection
- Regulators of Cell Cycle Progression and Differentiation Identified Using a Kinome-Wide RNAi Screen
- Absence of Intestinal PPARγ Aggravates Acute Infectious Colitis in Mice through a Lipocalin-2–Dependent Pathway
- Induction of a Stringent Metabolic Response in Intracellular Stages of Leads to Increased Dependence on Mitochondrial Metabolism
- CTCF and Rad21 Act as Host Cell Restriction Factors for Kaposi's Sarcoma-Associated Herpesvirus (KSHV) Lytic Replication by Modulating Viral Gene Transcription
- Gammaherpesviral Gene Expression and Virion Composition Are Broadly Controlled by Accelerated mRNA Degradation
- The Arabidopsis Silencing Pathway Modulates PAMP- and Effector-Triggered Immunity through the Post-transcriptional Control of Disease Resistance Genes
- Inflammatory Stimuli Reprogram Macrophage Phagocytosis to Macropinocytosis for the Rapid Elimination of Pathogens
- Alphavirus Mutator Variants Present Host-Specific Defects and Attenuation in Mammalian and Insect Models
- Phosphopyruvate Carboxylase Identified as a Key Enzyme in Erythrocytic Carbon Metabolism
- IFNγ/IL-10 Co-producing Cells Dominate the CD4 Response to Malaria in Highly Exposed Children
- Electron Tomography of HIV-1 Infection in Gut-Associated Lymphoid Tissue
- Characterisation of a Multi-ligand Binding Chemoreceptor CcmL (Tlp3) of
- Single Cell Stochastic Regulation of Pilus Phase Variation by an Attenuation-like Mechanism
- Cell Tropism Predicts Long-term Nucleotide Substitution Rates of Mammalian RNA Viruses
- Functions of CPSF6 for HIV-1 as Revealed by HIV-1 Capsid Evolution in HLA-B27-Positive Subjects
- RNA-seq Analysis of Host and Viral Gene Expression Highlights Interaction between Varicella Zoster Virus and Keratinocyte Differentiation
- Kaposi's Sarcoma Associated Herpesvirus Tegument Protein ORF75 Is Essential for Viral Lytic Replication and Plays a Critical Role in the Antagonization of ND10-Instituted Intrinsic Immunity
- DAMP Molecule S100A9 Acts as a Molecular Pattern to Enhance Inflammation during Influenza A Virus Infection: Role of DDX21-TRIF-TLR4-MyD88 Pathway
- Variable Suites of Non-effector Genes Are Co-regulated in the Type III Secretion Virulence Regulon across the Phylogeny
- Reengineering Redox Sensitive GFP to Measure Mycothiol Redox Potential of during Infection
- Preservation of Tetherin and CD4 Counter-Activities in Circulating Alleles despite Extensive Sequence Variation within HIV-1 Infected Individuals
- KSHV 2.0: A Comprehensive Annotation of the Kaposi's Sarcoma-Associated Herpesvirus Genome Using Next-Generation Sequencing Reveals Novel Genomic and Functional Features
- Nutrient Limitation Governs Metabolism and Niche Adaptation in the Human Nose
- Decreases in Colonic and Systemic Inflammation in Chronic HIV Infection after IL-7 Administration
- Investigation of Acetylcholine Receptor Diversity in a Nematode Parasite Leads to Characterization of Tribendimidine- and Derquantel-Sensitive nAChRs
- Intranasal Vaccination Promotes Detrimental Th17-Mediated Immunity against Influenza Infection
- -Mediated Inhibition of Iron Export Promotes Parasite Replication in Macrophages
- Variation in RNA Virus Mutation Rates across Host Cells
- A Single Amino Acid in the Stalk Region of the H1N1pdm Influenza Virus HA Protein Affects Viral Fusion, Stability and Infectivity
- Group B Engages an Inhibitory Siglec through Sialic Acid Mimicry to Blunt Innate Immune and Inflammatory Responses
- Synthesis and Biological Properties of Fungal Glucosylceramide
- HIV Protective KIR3DL1/S1-HLA-B Genotypes Influence NK Cell-Mediated Inhibition of HIV Replication in Autologous CD4 Targets
- Recruitment of PfSET2 by RNA Polymerase II to Variant Antigen Encoding Loci Contributes to Antigenic Variation in
- Human and Plant Fungal Pathogens: The Role of Secondary Metabolites
- Lyme Disease: Call for a “Manhattan Project” to Combat the Epidemic
- Enhancing Virus-Specific Immunity by Combining Therapeutic Vaccination and PD-L1 Blockade in Chronic Hepadnaviral Infection
- Suppression of Interferon Lambda Signaling by SOCS-1 Results in Their Excessive Production during Influenza Virus Infection
- Inflammation Fuels Colicin Ib-Dependent Competition of Serovar Typhimurium and in Blooms
- Host-Specific Enzyme-Substrate Interactions in SPM-1 Metallo-β-Lactamase Are Modulated by Second Sphere Residues
- STING-Dependent Type I IFN Production Inhibits Cell-Mediated Immunity to
- From Scourge to Cure: Tumour-Selective Viral Pathogenesis as a New Strategy against Cancer
- Lysine Acetyltransferase GCN5b Interacts with AP2 Factors and Is Required for Proliferation
- Narrow Bottlenecks Affect Populations during Vertical Seed Transmission but not during Leaf Colonization
- Targeted Cytotoxic Therapy Kills Persisting HIV Infected Cells During ART
- Murine Gammaherpesvirus M2 Protein Induction of IRF4 via the NFAT Pathway Leads to IL-10 Expression in B Cells
- iNKT Cell Production of GM-CSF Controls
- Malaria-Induced NLRP12/NLRP3-Dependent Caspase-1 Activation Mediates Inflammation and Hypersensitivity to Bacterial Superinfection
- Detection of Host-Derived Sphingosine by Is Important for Survival in the Murine Lung
- PLOS Pathogens
- Archív čísel
- Aktuálne číslo
- Informácie o časopise
Najčítanejšie v tomto čísle- Lyme Disease: Call for a “Manhattan Project” to Combat the Epidemic
- Origin, Migration Routes and Worldwide Population Genetic Structure of the Wheat Yellow Rust Pathogen f.sp.
- IFNγ/IL-10 Co-producing Cells Dominate the CD4 Response to Malaria in Highly Exposed Children
- Human and Plant Fungal Pathogens: The Role of Secondary Metabolites
Prihlásenie#ADS_BOTTOM_SCRIPTS#Zabudnuté hesloZadajte e-mailovú adresu, s ktorou ste vytvárali účet. Budú Vám na ňu zasielané informácie k nastaveniu nového hesla.
- Časopisy



