-
Články
- Časopisy
- Kurzy
- Témy
- Kongresy
- Videa
- Podcasty
Regulation by the Noncoding RNA
The homeotic genes in Drosophila melanogaster are aligned on the chromosome in the order of the body segments that they affect. The genes affecting the more posterior segments repress the more anterior genes. This posterior dominance rule must be qualified in the case of abdominal-A (abd-A) repression by Abdominal-B (Abd-B). Animals lacking Abd-B show ectopic expression of abd-A in the epidermis of the eighth abdominal segment, but not in the central nervous system. Repression in these neuronal cells is accomplished by a 92 kb noncoding RNA. This “iab-8 RNA” produces a micro RNA to repress abd-A, but also has a second, redundant repression mechanism that acts only “in cis.” Transcriptional interference with the abd-A promoter is the most likely mechanism.
Published in the journal: Regulation by the Noncoding RNA. PLoS Genet 8(5): e32767. doi:10.1371/journal.pgen.1002720
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002720Summary
The homeotic genes in Drosophila melanogaster are aligned on the chromosome in the order of the body segments that they affect. The genes affecting the more posterior segments repress the more anterior genes. This posterior dominance rule must be qualified in the case of abdominal-A (abd-A) repression by Abdominal-B (Abd-B). Animals lacking Abd-B show ectopic expression of abd-A in the epidermis of the eighth abdominal segment, but not in the central nervous system. Repression in these neuronal cells is accomplished by a 92 kb noncoding RNA. This “iab-8 RNA” produces a micro RNA to repress abd-A, but also has a second, redundant repression mechanism that acts only “in cis.” Transcriptional interference with the abd-A promoter is the most likely mechanism.
Introduction
Genome wide surveys for RNA transcription units in a variety of eukaryotes have revealed a surprising number of transcripts that are not traditional messenger RNAs. A variety of functions have been suggested for these “noncoding” RNAs (ncRNAs), although the large majority have no known purpose (reviewed in ref [1]). In Drosophila melanogaster, primary transcripts cover at least 60% of the genome [2]. Many of these transcripts do not correspond to defined genes, but they are evolutionarily conserved. Particular attention has been given to ncRNAs in the bithorax complex (BX-C), a cluster of three homeobox-containing transcription factors required for segment identity (reviewed in ref. [3]). Although much of the ∼300 kb of BX-C DNA is transcribed, the BX-C contains only one other protein coding sequence [4]. Lipshitz et al. [5] first described apparent ncRNAs from the BX-C in the bithoraxoid regulatory region. They suggested that such transcripts could reflect nonspecific initiation by RNA polymerase near a strong enhancer, a possibility that still remains attractive. Several other ncRNAs in the BX-C have been identified by Northern blots or RNA in-situ hybridizations [6]–[8]. It has been suggested that such transcripts might block silencing by the Polycomb Group proteins [9], but this idea is not yet supported by the analysis of existing mutations. A readthrough product of the bithoraxoid ncRNA transcription unit may repress features of early transcription from the Ultrabithorax (Ubx) promoter [10], and the iab-4 and iab-8 ncRNAs are the likely precursors for micro RNAs (miRNAs) [11]–[13]. Otherwise, these ncRNAs still lack functions.
Prior indications of the iab-8 noncoding RNA
Here, we focus on the iab-8 ncRNA. Several lines of evidence have suggested the existence of a 90 kb-long transcription unit, extending between Abd-B and abd-A, with a likely start site within the iab-8 regulatory region. RNA in-situ hybridizations to embryos, using genomic DNA fragment from the iab-2 through the iab-8 regulatory regions as probes, detect an RNA in the 8th and 9th abdominal segments (parasegments 13 & 14). Strand-specific probes revealed that it is transcribed in a distal-to-proximal direction (from Abd-B towards abd-A) [7], [8], [14], [15]. This transcript is first seen at about stage 6 [11] in the epidermis, but from stage 14 onward (germband shortening), the RNA is detected only in the developing central nervous system (CNS). A promoter for an uncharacterized RNA was independently mapped to the iab-8 region, just downstream of the Abd-B transcription unit [16].
Additionally, a transcript starting in the iab-8 region has been suggested as the precursor for a micro RNA, called miR-iab-8 or miR-iab-4AS [11]–[13]. This miRNA is transcribed from the iab-3 regulatory region in the distal-to-proximal direction, and strand specific genomic probes from this region indicate that the precursor is made in the 8th and 9th abdominal segments, as described above. This miRNA is required for male and female fertility, and complementation tests with a series of rearrangement breakpoints suggest that the start site of this RNA is in the iab-8 regulatory region, downstream of Abd-B [11].
Here, we characterize the structure and function of the 92 kb long “iab-8 ncRNA”. This ncRNA represses the expression of the homeotic gene abd-A in the posterior CNS. This repression depends not only on the miR-iab-8 micro RNA, but also on transcriptional interference in the region of the abd-A promoter.
Results/Discussion
Repression of abd-A in the 8th abdominal segment
In wild type embryos, abd-A expression is detected in the epidermis and CNS of PS7 to PS12 but not in PS13 (Figure 1A). Abd-B is strongly expressed in PS13, and it was initially claimed that Abd-B represses abd-A in PS13 [17], just as abd-A represses Ubx and Ubx represses Antp [18]. This repression hierarchy can account for the dominance of posterior homeotic genes over anterior ones, often called “posterior prevalence” [19]. Indeed, embryos homozygous for Df(3R)C4, which removes Abd-B, show ABD-A expression extending throughout PS13 (Figure 1B). However, the Df(3R)C4 deficiency extends downstream of the ABD-B transcription unit, removing all of the iab-8 regulatory region and part of iab-7 (Figure 2). Surprisingly, embryos homozygous for an Abd-B null point mutation, Abd-BD16, show ABD-A derepression in PS13 of the epidermis, but not in the CNS (Figure 1C). Homozygotes for Abd-BD18, a deletion removing all of the Abd-B coding sequences (Figure 2), show the same ABD-A expression pattern (not shown). This unexpected repression of ABD-A in the CNS can be seen most dramatically in the Abd-BD14 mutation. Abd-BD14 deletes the promoter for the Abd-B “m” transcript [20], expressed from PS10 through PS13, but leaves the promoters for the “r” transcripts expressed in PS14. In the CNS of Abd-BD14 homozygotes, abd-A does not fill in the gap left by the absence of Abd-B in PS13 (Figure 1D). Clearly then, there must be some function deleted by Df(3R)C4 that is not affected by Abd-BD18 or more subtle Abd-B mutations. Our attention turned to the iab-8 ncRNA, which appeared to initiate in the iab-8 region deleted in Df(3R)C4.
Fig. 1. ABD-A expression in Abd-B mutant embryos. 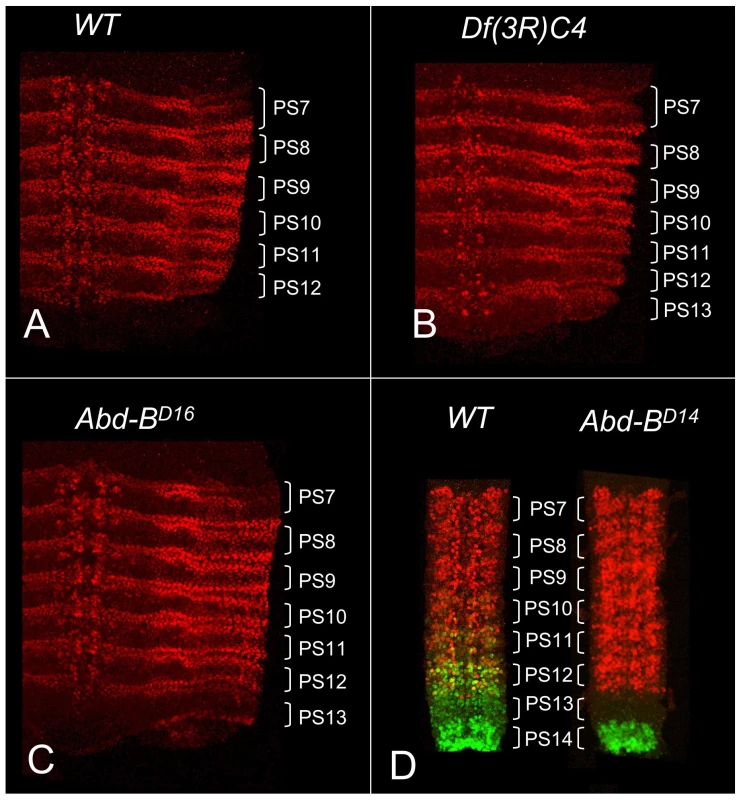
Stage 14 embryos in A–C were stained with antibody to ABD-A, opened along the dorsal midline and flattened for photography. ABD-A is absent from PS13 in wild type (A), but appears throughout PS13 in Df(3R)C4 homozygotes (B). In Abd-BD16 homozygotes, ABD-A is only in the lateral and dorsal epidermis of PS13 (C). Dissected CNS's from stage15 embryos in D were doubly stained for ABD-A (red) and ABD-B (green). In wild type, the expression domains overlap through PS10-12, with some nuclei expressing both proteins. In Abd-BD14 homozygotes, ABD-B expression is absent from PS10-13, but the ABD-A pattern is unchanged, leaving PS13 without either protein. Fig. 2. Map of the abdominal half of the bithorax complex. 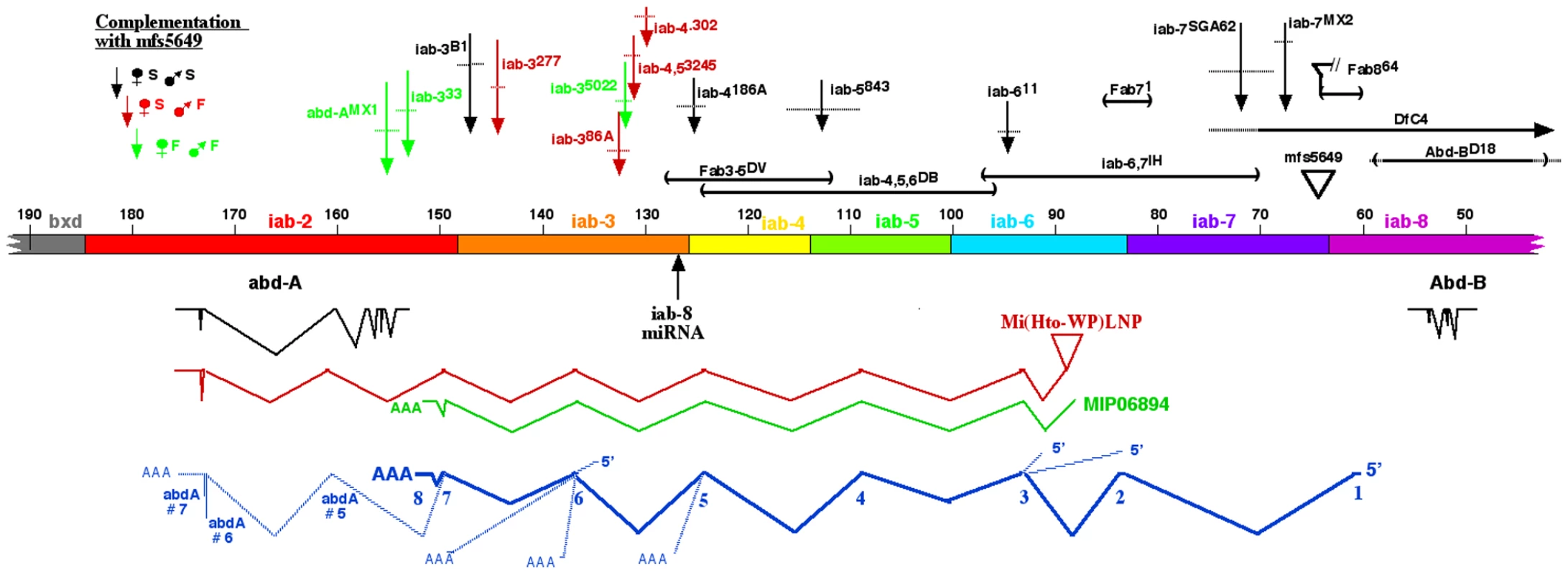
The horizontal bar indicates the DNA sequence map, numbered in kb according to Martin et al. [4] (Genbank U31961). Base #1 corresponds to base 12,809,162 on chromosome 3R in release 5.37 of the Drosophila genome. The coordinates proceed distal to proximal on chromosome 3R, which is opposite in orientation to the whole genome numbering. The regulatory domains iab-2 through iab-8 are color coded; the domain borders are defined by deletion mutations (Fab8 [22], ; Fab7, [41]; Mcp, [44]; iab-3/iab-4, L. Sipos personal communication), or inferred from the binding sites of the CTCF factor [45]. Below the DNA bar are shown the splicing patterns of abd-A and Abd-B (in black), a cDNA derived from the Mi(Hto-WP)LNP insertion (red), and the MIP06894 cDNA (green). At the bottom, the splicing pattern for the iab-8 ncRNA is shown in dark blue, with numbered exons, and alternate 5′ or 3′ extensions indicated with light blue lines. Mutant lesions are indicated above the DNA bar. The rearrangement breakpoints are color coded according to their phenotypes when heterozygous with the mfs5649 insertion. Mapping the iab-8 ncRNA exons
The spliced product of the iab-8 ncRNA was initially uncovered by a fortuitous insertion of an exon-trap mobile element. This element, a derivative of the Minos mobile element, is called Hostile takeover (Mi[Hto-WP]; Genbank #JN049642). An insertion was recovered in the iab-6 domain of the BX-C (“TA” target site bases 85,277 & 85,278), named Mi(Hto-WP)LNP or simply LNP, for short (Figure 2). 3′ RACE products were amplified with primers within LNP and within the 3′ exon of abd-A. The sequence of the product revealed the exon structure diagramed in Figure 2. The sequence included 5 novel exons before it spliced into abd-A, at the 5th exon of the predominant splice form of the abd-A mRNA [4], [17]. Many of these exons match those of a cDNA designated MIP06894 (Genbank BT099824.1)(Figure 2), identified by the Berkeley Drosophila Genome Project.
The exons of the LNP cDNA were used to generate primers for 5′ and 3′ RACE, using total RNA from Oregon R embryos. Figure 2 diagrams the predominant splicing product, which spans ∼92 kb. An RT/PCR product was recovered and sequenced that extended from exon 1 through exon 8, as well as one that extended from exon 1 through exon 7, and then included exons 5, 6, and 7 of the abd-A transcription unit. Figure 2 also shows three alternate 5′ exons and five alternate 3′ splicing patterns. RT/PCR products included extensions of exons #2 or 4, ending at sites of genomic poly(dA) stretches; these were likely derived from splicing intermediates. Rare clones were also recovered that skipped exons, splicing from exons 1, 2, or 6 into abd-A exons 5 or 6. Exon 1 had two start sites separated by 135 bases; the upstream start was ∼3 fold more abundant. Exon 4 included only 6 bases, although rare products included an alternate 3′ extension of 92 bases. Quantitative PCR was also used to show that termination at exon 8 was ∼500-fold more common than splicing into abd-A. The sequences of the predominant and alternate exons are given in Figure S1. Two recent genome-wide searches for novel non-coding transcripts in embryos have uncovered some of these same transcripts [2], [21]. Graveley et al. [2] also reported transcripts from adult males with most of the same exons, but an alternate start site, in the iab-6 region.
The promoter for the iab-8 RNA maps distal to the Fab-8 boundary, in the iab-8 regulatory region [22]. The iab-8 region should be under Polycomb Group repression in parasegments 1–12, which explains why the transcript is only expressed in PS13 and 14 [11] . Exons 1–7 appear to be evenly spaced across the abdominal region of the bithorax complex, with one in each of the iab cis-regulatory domains. A comparison with the genomic sequences of various Drosophila species suggests that the sequences of the exons are not more conserved than those of the introns. However, the existence of the exons does appear to be conserved, in that the splice junctions are among the most conserved features of the exons. This is illustrated in Figure 3A for exon 3, in the iab-6 region. The embryonic expression pattern is also conserved; expression is restricted to PS13 and 14 in D. pseudoobscura and D. virilis, as it is in D. melanogaster (see Figure S2).
Fig. 3. Evolutionary conservation. 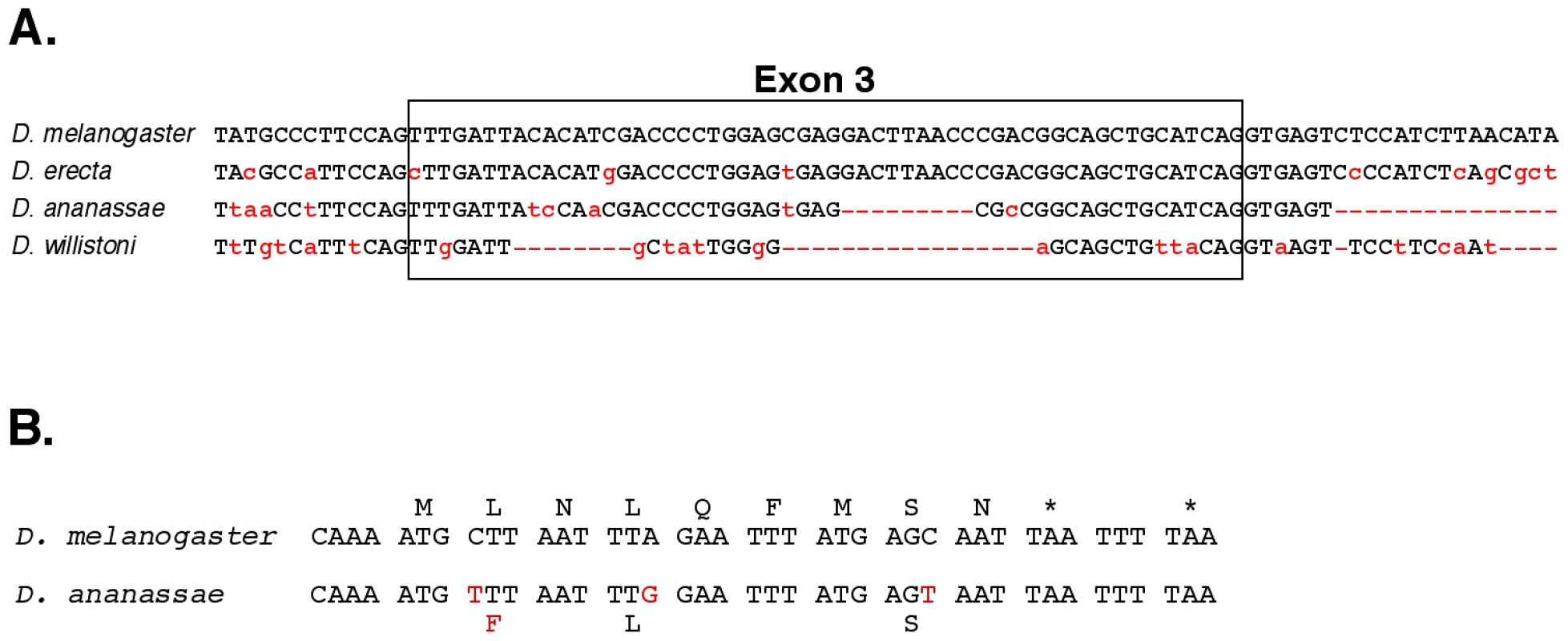
A. A comparison of exon 3 and neighboring bases with the homologous regions from the genomes of three other Drosophila species. B. Potential nine amino acid peptide within exon 8 of the iab-8 ncRNA. The D. melanogaster sequence is compared to that of D. ananassae. The initial methionine codon is preceded by a perfect translation start consensus sequence [46], and there are two stop codons after the 9th amino acid. The three bases altered in D. ananassae are highlighted in red; only one changes the predicted amino acid. The spliced product of the iab-8 RNA is non-coding by traditional criteria, but the possibility of small peptides [23], [24] cannot be ruled out. In particular, exon 8 includes a potential 9 amino acid peptide, with appropriate translation initiation and termination signals, and the coding potential for this peptide is well conserved in D. ananassae, D. pseudoobscura and D. willistoni (Figure 3B), although it is not found in D. virilis and more distantly related species.
abd-A repression by the iab-8 RNA
There are many chromosome rearrangements, mostly from the collection of E. B. Lewis, which interrupt the iab-8 ncRNA transcription unit. These can be used to test whether truncated versions of the iab-8 RNA can repress abd-A. Rearrangement breaks that truncate the iab-8 RNA near its start site cause a dramatic derepression of abd-A in the CNS of the 8th abdominal segment, indistinguishable from that seen in Df(3R)C4 homozygotes (Figure 1). Rearrangements with this effect include iab-7SGA62, iab-611, iab-5843, iab-42330, and iab-4186 (Figure 2 & 4). The same spread of ABD-A into the CNS of PS13 is seen with the Fab-864 deletion, which removes the iab-8 ncRNA promoter (Figure 2). Interestingly, embryos homozygous for chromosome breaks mapping closer to abd-A show a much more subtle derepression of abd-A. In iab-35022 homozygotes, for example, weak misexpression is limited to a few cells (Figure 4). Similar weak misexpression is seen in homozygotes of iab-4,53245 and iab-4302 (Figure 2). Finally, embryos homozygous for the iab-333 rearrangement show abd-A misexpression in only a very few CNS cells in the most anterior part of the 8th abdominal segment (Figure 4). This break lies downstream of the poly(A) addition site of the major iab-8 transcript, but upstream of the abd-A transcription start site.
Fig. 4. ABD-A expression in rearrangements truncating the iab-8 ncRNA. 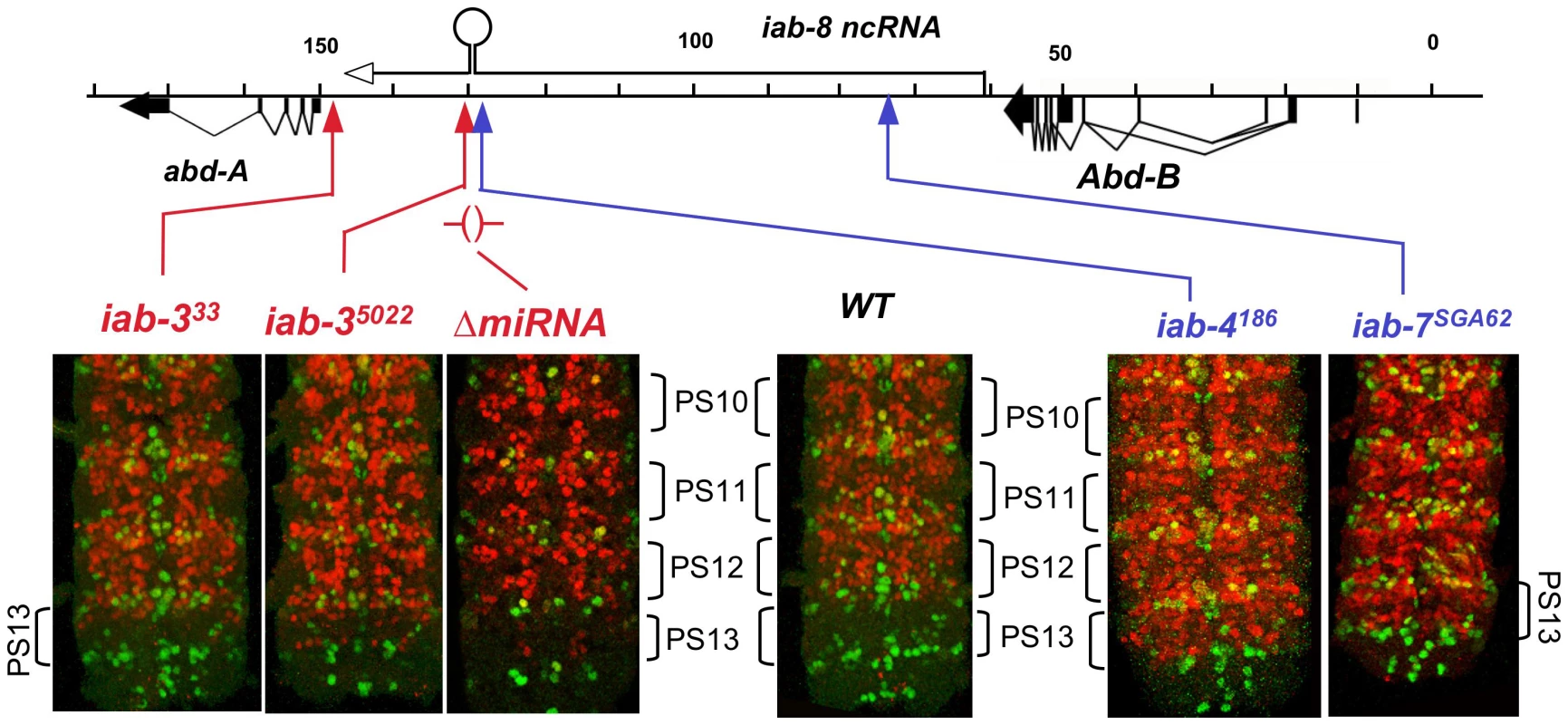
Embryos homozygous for the indicated mutations were doubly stained for ENGRAILED (green) and ABD-A (red), and the CNS's were dissected and photographed. The posterior end of each CNS is shown; the ENGRAILED stripes mark the parasegmental boundaries. The iab-4 and iab-7 breaks cause widespread misexpression of ABD-A in PS13, but iab-3 breaks show only subtle misexpression in a few nuclei. Embryos homozygous for a deletion of the iab-8 miRNA also show misexpression in only a few nuclei. The difference between the two classes of breakpoints seems to be the expression of miR-iab-8. The iab-4186 break, maps just upstream (within 3 kb) of the miR-iab-8 coding region and shows complete loss of abd-A repression in PS13. In contrast, the iab-35022 break maps ∼5 kb downstream of miR-iab-8 and shows only slight misexpression. Thus, one might guess that miR-iab-8 is responsible for most of the repression of abd-A, especially since the 3′ UTR of abd-A includes sequences homologous to the “seed” region of miR-iab-8 [12], [13]. However, embryos homozygous for a deletion of miR-iab-8 (ΔmiR-iab-8) do not show a dramatic misexpression of ABD-A in the PS13 CNS [11]. A closer examination of these homozygous embryos does reveal a weak misexpression of abd-A in a small number of nuclei in anterior PS13 (Figure 4), but clearly not the strong and widespread misexpression of iab-4186. Thus, it appears that miR-iab-8 does repress abd-A in the PS13 CNS, but there must be a second, redundant function of the iab-8 RNA to completely represses abd-A. UBX expression in embryos is apparently not affected by this second function; its expression pattern in iab-7SGA62 homozygous embryos is the same as that in miR-iab-8 deletion homozygotes (not shown).
Fertility function of the iab-8 ncRNA
A deletion of the miR-iab-8 causes sterility in both sexes [11]. Thus, we expected that any combination of alleles that failed to make the miR-iab-8 micro RNA would be sterile, including, for example, an iab-7 break (iab-7MX2 or iab-7SGA62) heterozygous with ΔmiR-iab-8 [11]. There is an insertion of the “PZ” P element ∼4.2 kb downstream of the iab-8 RNA start site, designated mfs(3)05649 (here called mfs5649; Figure 2). Homozygotes are sterile in both sexes, and the females show the same phenotype (blockage of the oviduct) as is seen in ΔmiR-iab-8 homozygotes [25]. We assume the mfs5649 insertion truncates the iab-8 RNA, since it fails to complement with ΔmiR-iab-8 for the sterility phenotype. The Fab864 deletion (derived from the mfs5649 P element; Figure 2; [22]) is also sterile as a homozygote or as a heterozygote with ΔmiR-iab-8.
We tested rearrangement breakpoints in the iab-2,3, and 4 regions, downstream of the miR-iab-8 template, for fertility when heterozygous with the mfs5649 P element. Surprisingly, many rearrangement breakpoints 3′ to the miR iab-8 template have a female sterility phenotype when heterozygous with mfs5649 (Figure 2); males of these genotypes are fertile. These sterile females show a failure of mature oocytes to move through the oviduct, much like mfs5649 homozygotes or the ΔmiR-iab-8 homozygotes. It does not seem likely that breakpoints downstream of the miR-iab-8 template interfere with the proper processing of the micro RNA, because these same breakpoints are fertile when heterozygous to ΔmiR-iab-8. It is possible that the subtle misexpression of ABD-A in PS13 seen in iab-3 breaks is responsible for the female sterility, especially if the misexpression it is more dramatic at later times in development. Not all breakpoints give this female sterility phenotype, and there is no apparent order to the fertile and sterile breakpoint alleles (Figure 2). Some of the rearrangements may fuse the iab-3 region with novel transcription units, restoring the repression of abd-A in the critical cells.
Mechanism of repression
The iab-8 ncRNA could make a product, such as another miRNA, that represses abd-A. Indeed, there is a secondary structure hairpin in exon 6 of the spliced transcript that could serve as a miRNA precursor. The iab-8 ncRNA could also code for tiny peptides, as noted above (Figure 3B). These possibilities prompted us to misexpress the iab-8 ncRNA spliced product. A cDNA cassette, representing the major splicing product (Figure 2) plus 236 bp of genomic DNA downstream of the poly(A) addition site, was cloned into the pUAST vector [26]. P element transgenes were recovered and crossed to flies expressing the yeast GAL4 activator in abdominal segments 3–8 (parasegments 8–13). However, embryos containing both the GAL4 activator and the UAS/iab-8 cDNA target showed no apparent reduction in the ABD-A levels in the segments expressing GAL4 (not shown).
The cDNA misexpression experiment does not rule out a product made from an intron, such as an RNA component of a diffusible repressive complex, as alleged for non-coding RNAs in mammalian HOX complexes [27]. If the putative second repressor involves a diffusible molecule, it should be able to act on both chromosomes, even if it is only produced by one. The miR-iab-8 micro RNA should be diffusible in this way, and so, to examine the second repressive function, we needed to test genotypes lacking miR-iab-8. Specifically, heterozygotes were made with the ΔmiR-iab-8 deletion on one chromosome, and with a mutation truncating the iab-8 RNA upstream of the miRNA template (mfs5649, iab-7SGA62, or iab-5845) on the other chromosome. Such embryos make the iab-8 RNA from only one chromosome, and cannot make the micro RNA from either. As shown in Figure 5, these embryos showed strong ABD-A misexpression in the CNS of PS13 (the 8th abdominal segment), suggesting that the iab-8 RNA can only repress the copy of abd-A on the chromosome from which it is transcribed. To control for a potential effect of haploinsufficiency of the iab-8 RNA, the ΔmiR-iab-8 deletion was also tested over DfP9, a deletion that removes the entire bithorax complex. These ΔmiR-iab-8/DfP9 embryos show no apparent misexpression of ABD-A in PS13. Thus, the second iab-8 RNA repressive function must act only in cis.
Fig. 5. Test for trans repression by the iab-8 ncRNA. 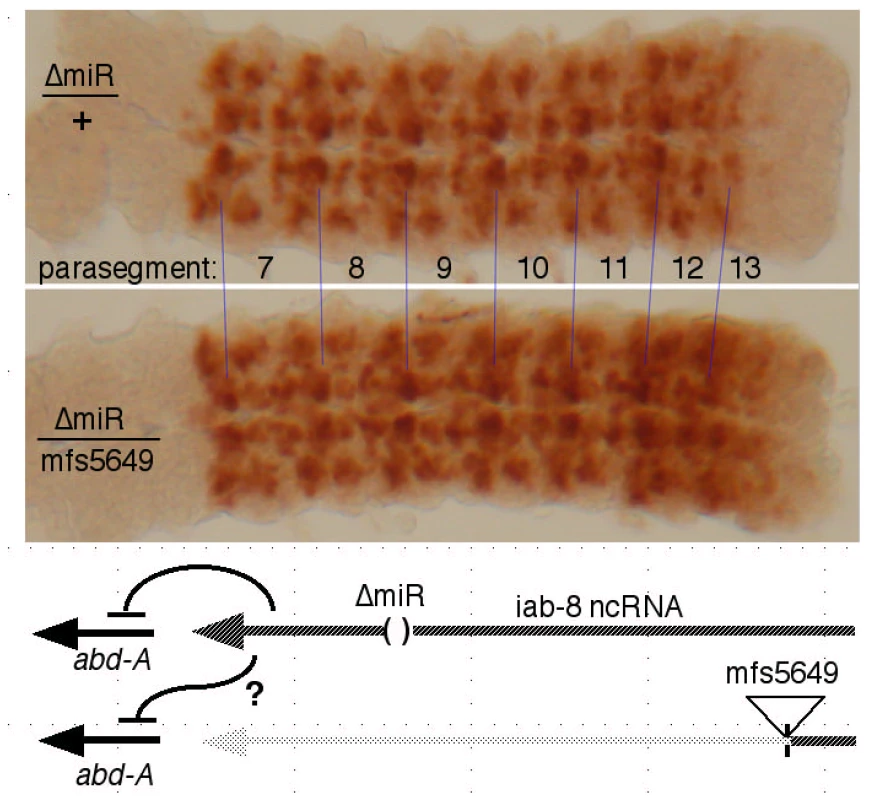
CNS's, stained for ABD-A, were dissected from embryos of the indicated genotypes. In ΔmiR/mfs5649 embryos, only one chromosome makes a full length iab-8 ncRNA, and neither chromosome produces the iab-8 miRNA, as diagrammed. The strong expression of ABD-A in PS13 in this genotype shows that the abd-A gene on the mfs5649 chromosome is not repressed, i.e. the iab-8 ncRNA acts only in cis. In a similar test, we employed a duplication for the proximal two thirds of the complex, Dp(3∶2)D109, which extends into the iab-5 region (at ∼110 kb; [28]). This duplication includes abd-A, but lacks the iab-8 RNA promoter. Embryos homozygous for the ΔmiR-iab-8 deletion but containing this duplication also show ABD-A misexpression in the PS13 CNS (Figure S3). Thus, there are two mechanisms by which the iab-8 RNA represses abd-A, first, through production of the iab-8 miRNA (acting in trans), and second, a repressive function acting only in cis. The Supplementary Table S1 summarizes which genotypes supply which repressive functions.
The cis-repression of one transcription unit by another is often termed transcriptional interference. This term, however, encompasses several possible molecular mechanisms [29]. An example of a long, ncRNA involved in transcriptional cis-repression is the XIST RNA, involved in mammalian X chromosome inactivation [30] (A recent report suggests that the XIST RNA can also work in trans [31]). Nascent transcripts are involved in repression in RNAi silencing of heterochromatin is fission yeast [32] and in RNA-directed DNA methylation in Arabidopsis [33]. By analogy to these systems, the iab-8 RNA could recruit gene silencing machinery to the site of its transcription. The RNA sequences required for such recruitment might be mapped by examination of deletions in the BX-C. Ideally, the iab-8 miRNA should be removed to have a clear assay for the cis repressor. We have checked embryos homozygous for the Fab3,5DV deletion (Figure 2), which covers the site of the iab-8 miRNA precursor; they still show abd-A repression in the posterior CNS. Likewise, a double deletion chromosome, with ΔmiR-iab-8 and Fab71, also retains the cis repression. The Fab71 deletion (Figure 2) was tested because it removes a Polycomb Response Element [34], [35] which is coincident with exon 2 of the cDNA. Two other deletions have been examined which span the iab-4 through iab-7 regions, although both retain the iab-8 miRNA (iab-4,5,6DB and iab-6,7IH; Figure 2). In these, we looked for more subtle misexpression, such as that seen in iab-3 breaks (Figure 4), but no such misexpression was seen. This analysis does not yet cover the iab-2 and iab-3 regions, nor does it exclude the possibility of multiple redundant sequences throughout the transcription unit that could recruit repressive factors.
A more likely represion mechanism, perhaps, is that the RNA polymerase transcribing the iab-8 RNA somehow interfers with the abd-A promoter. Examples of this type of transcriptional interference come from budding yeast, where the GAL7 gene is repressed by the upstream GAL10 transcript [36], and the SER3 gene is repressed by the upstream, noncoding SRG1 transcript [37]. In these cases, the 3′ ends of the upstream transcripts are close to the downstream promoters, suggesting repression by occlusion of the downstream promoters or their proximal enhancers. If the iab-8 RNA interferes with an abd-A enhancer, that enhancer must lie downstream of the iab-4186 breakpoint, since abd-A is totally derepressed in the PS13 CNS in embryos homozygous for this break (Figure 4). The abd-A promoter seems like the most likely target of interference, since the major poly(A) site of the iab-8 RNA lies only 1.1 kb upstream of the initiation site of abd-A, and the iab-8 RNA primary transcript likely continues past its poly(A) addition site [38]. In any case, minor splice variants clearly do continue past the poly(A) site and into the abd-A transcription unit (Figure 2).
The function of the iab-8 ncRNA fits with the rule of posterior dominance - it blocks expression of a more anterior homeotic gene in more posterior segments. The repression of Ubx by the bxd ncRNA [10], although subtle, fits the same pattern. The novel aspect, here, is that this posterior repression can be accomplished by noncoding transcription units, in addition to DNA binding proteins. The mechanism of transcriptional interference would fix the arrangement of these ncRNAs in the bithorax complex. It seems possible that the ancestral HOX complex turned off anterior genes by readthrough transcripts of more posterior genes, or by noncoding RNAs initiating from their posterior enhancers. Such a method of repression would dictate the linear order of the HOX genes, 3′ to 5′, anterior to posterior.
Materials and Methods
Drosophila strains
Wild type stocks were Canton S or Oregon R. Mutations included abd-AMX1, iab-3277, iab-4302, iab-5843, iab-7SGA62, iab-7MX2, Abd-BD16, Abd-BD14, Df(3R)C4, Df(3)P9 (ref [39]); iab-333, iab-3B1 (ref [17]) iab-611 ref [40]); Fab71 ref [41]); mfs(3)05649 (ref [25], Fab864 ref [22]; Fab3-5DV ref [42]; iab-386A, iab-35022, iab-4186, iab-42330, iab4,53245, iab-5843, ΔmiR-iab-8 (ref [11]);); T(3∶2)DpD109 [28] and Mi[Hto-WP] (described here).
Antibody staining
Embryos were fixed, stained, and mounted as described by [17]. Primary antibodies used were mouse anti-ADB-B (1∶2 dilution, developed by S. Celniker, Developmental Studies Hybridoma Bank), mouse anti-UBX (1∶10, developed by R. White, Developmental studies Hybridoma Bank), rabbit anti-ß-galactosidase (1∶1500, Cappel/MP Biomedicals), mouse anti-ß-galactosidase (1∶1000, Promega), rabbit anti-En (1∶500, Santa Cruz Biotechnology), mouse anti-ABD-A (1∶500, 6A18.12, gift of I. Duncan), and goat anti-ABD-A (1∶100, Santa Cruz Biotechnology). Secondary antibodies were donkey anti-mouse, donkey anti-goat, and donkey anti-rabbit, coupled to either Alexa 488 or Alexa 555 (1∶500, Invitrogen), and HRP coupled goat anti-mouse (1∶1000, Bio-Rad).
The CNS's were hand dissected with tungsten needles and placed on a glass slide in a drop of Immu-Mount (for HRP staining, Shandon) or Vectashield with DAPI (for fluorescence, Vector Laboratories), and then gently flattened under a coverslip. Fluorescence images were taken with a Leica SP2 AOBS confocal microscope; the fluorescence pictures show free projection averages of stacks of images, after scanning through the depth of the tissue. Homozygous embryos were identified by the absence of lacZ staining from the TM3 ftz-LacZ balancer.
Fertility tests
Each of ten mutant virgin females was placed in a vial with three wild type males. Likewise ten mutant males were mated, each with three wild type virgin females. Vials were maintained at 25° for five days, and then examined for the presence of larvae.
cDNA analysis
Adults heterozygous for Mi[Hto-WP] and Hsp70-Gal4 (Bloomington stock #1799) were heat shocked for 45 min. at 37° to induce GAL4 expression, and then left to express the LNP transcript at room temperature for 4 h. RNA was then isolated using TRI reagent (Sigma) and reverse transcribed with MMLV reverse transcriptase (Promega) using an adaptor primer (GAAGACAGACACCGGACT18V). PCR was then performed using a forward primer in Hto and a reverse primer in the 6th exon of abd-A. The resulting amplicon was sequenced to identify the splicing pattern.
Total RNA from Oregon R embryos was prepared using the RNAqueous-4PCR kit (Ambion), and 3′ RACE and RNA ligase-mediated 5′ RACE reactions were performed using the FirstChoice RLM-RACE kit (Ambion). The 5′ RACE procedure was designed to recover only capped 5′ ends. Gel-isolated products were sequenced directly, or cloned first into the PCR-Blunt vector (Invitrogen). Quantitative PCR reactions used cDNA prepared from 6–12 h old embryos. The initial cDNA products were compared to measured dilutions of amplified cDNA products covering the relevant exons.
RNA in situ hybridization and embryo staining
The production of digoxigenin-labeled probes and the hybridization of embryos was as described by Fitzgerald and Bender [11], except that acetone treatment [43] was used instead of proteinase K for permeabilization of the embryos. Clones spanning exon 8 from D. melanogaster, D. pseudobscura and D. virils were recovered after PCR reaction on genomic DNAs with the following pairs of oligonucleotiedes: D. melanogaster 5′CGCTCGAGAGATTACAAACG3′ and 5′GGTGTATTACGGTCAAGGGGG3′ generating a fragment of 1013 bp; D. pseudobscura 5′CAGGCATTCAGTAAACACGGC3′ and 5′GGATGTGTCGAGTGGTGTGG3′ generating a fragment of 1477 bp; D.virilis 5′CTTTCGGTCCTATTCAACGG3′ and 5′CCGATCCTGCTGGTGTC3′ generating a fragment of 1364 bp.
Supporting Information
Zdroje
1. PontingCPOliverPLReikW 2009 Evolution and functions of long noncoding RNAs. Cell 136 629 641
2. GraveleyBRBrooksANCarlsonJWDuffMOLandolinJM 2011 The developmental transcriptome of Drosophila melanogaster. Nature 471 473 479
3. MaedaRKKarchF 2006 The ABC of the BX-C: the bithorax complex explained. Development 133 1413 1422
4. MartinCHMayedaCADavisCAEricssonCLKnafelsJD 1995 Complete sequence of the bithorax complex of Drosophila. Proc Natl Acad Sci U S A 92 8398 8402 Issn: 0027–8424
5. LipshitzHDPeattieDAHognessDS 1987 Novel transcripts from the Ultrabithorax domain of the bithorax complex. Genes Dev 1 307 322
6. CumberledgeSZaratzianASakonjuS 1990 Characterization of two RNAs transcribed from the cis-regulatory region of the abd-A domain within the Drosophila bithorax complex. Proc Natl Acad Sci U S A 87 3259 3263
7. Sanchez-HerreroEAkamM 1989 Spatially ordered transcription of regulatory DNA in the bithorax complex of Drosophila. Development 107 321 329
8. BaeECalhounVCLevineMLewisEBDrewellRA 2002 Characterization of the intergenic RNA profile at abdominal-A and -B in the Drosophila bithorax complex. Proc Natl Acad Sci U S A 99 16847 16852
9. SchmittSPrestelMParoR 2005 Intergenic transcription through a polycomb group response element counteracts silencing. Genes Dev 19 697 708
10. PetrukSSedkovYRileyKMHodgsonJSchweisguthF 2006 Transcription of bxd noncoding RNAs promoted by trithorax represses Ubx in cis by transcriptional interference. Cell 127 1209 1221
11. BenderW 2008 MicroRNAs in the Drosophila bithorax complex. Genes Dev 22 14 19
12. StarkABushatiNJanCHKheradpourPHodgesE 2008 A single Hox locus in Drosophila produces functional microRNAs from opposite DNA strands. Genes Dev 22 8 13
13. TylerDMOkamuraKChungWJHagenJWBerezikovE 2008 Functionally distinct regulatory RNAs generated by bidirectional transcription and processing of microRNA loci. Genes Dev 22 26 36
14. BenderWFitzgeraldDP 2002 Transcription activates repressed domains in the Drosophila bithorax complex. Development 129 4923 4930
15. RankGPrestelMParoR 2002 Transcription through intergenic chromosomal memory elements of the Drosophila bithorax complex correlates with an epigenetic switch. Mol Cell Biol 22 8026 8034
16. ZhouJAsheHBurksCLevineM 1999 Characterization of the transvection mediating region of the Abdominal - B locus in Drosophila. Development 126 3057 3065
17. KarchFBenderWWeiffenbachB 1990 abdA expression in Drosophila embryos. Genes Dev 4 1573 1587
18. HardingKWedeenCMcGinnisWLevineM 1985 Spatially regulated expression of homeotic genes in Drosophila. Science 229 1236 1242
19. DubouleD 1991 Patterning in the vertebrate limb. Curr Opin Genet Dev 1 211 216
20. ZavortinkMSakonjuS 1989 The morphogenetic and regulatory functions of the Drosophila Abdominal - B gene are encoded in overlapping RNAs transcribed from separate promoters. Genes Dev 3 1969 1981
21. EnderleDBeiselCStadlerMBGerstungMAthriP Polycomb preferentially targets stalled promoters of coding and noncoding transcripts. Genome Res 21 216 226
22. BargesSMihalyJGalloniMHagstromKMüllerM 2000 The Fab-8 boundary defines the distal limit of the bithorax complex iab-7 domain and insulates iab-7 from initiation elements and a PRE in the adjacent iab-8 domain. Development 127 779 790
23. KondoTPlazaSZanetJBenrabahEValentiP 2010 Small peptides switch the transcriptional activity of Shavenbaby during Drosophila embryogenesis. Science 329 336 339
24. GalindoMIPueyoJIFouixSBishopSACousoJP 2007 Peptides encoded by short ORFs control development and define a new eukaryotic gene family. PLoS Biol 5 e106 doi:10.1371/journal.pbio.0050106
25. LinHSpradlingAC 1993 Germline stem cell division and egg chamber development in transplanted Drosophila germaria. Dev Biol 159 140 152
26. BrandAHPerrimonN 1993 Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118 401 415
27. RinnJLKerteszMWangJKSquazzoSLXuX 2007 Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129 1311 1323
28. HopmannRDuncanDDuncanI 1995 Transvection in the iab-5,6,7 region of the bithorax complex of Drosophila: homology independent interactions in trans. Genetics 139 815 833 Issn: 0016–6731
29. ShearwinKECallenBPEganJB 2005 Transcriptional interference–a crash course. Trends Genet 21 339 345
30. PennyGDKayGFSheardownSARastanSBrockdorffN 1996 Requirement for Xist in X chromosome inactivation. Nature 379 131 137
31. JeonYLeeJT 2011 YY1 tethers Xist RNA to the inactive X nucleation center. Cell 146 119 133
32. HalicMMoazedD 2010 Dicer-independent primal RNAs trigger RNAi and heterochromatin formation. Cell 140 504 516
33. WierzbickiATHaagJRPikaardCS 2008 Noncoding transcription by RNA polymerase Pol IVb/Pol V mediates transcriptional silencing of overlapping and adjacent genes. Cell 135 635 648
34. HagstromKMullerMSchedlP 1997 A Polycomb and GAGA dependent silencer adjoins the Fab-7 boundary in the Drosophila bithorax comple. Genetics 146 1365 1380
35. MihalyJHoggaIGauszJGyurkovicsHKarchF 1997 In situ dissection of the Fab-7 region of the bithorax complex into a chromatin domain boundary and a Polycomb-response element. Development 124 1809 1820
36. GregerIHProudfootNJ 1998 Poly(A) signals control both transcriptional termination and initiation between the tandem GAL10 and GAL7 genes of Saccharomyces cerevisiae. EMBO J 17 4771 4779
37. MartensJAWuPYWinstonF 2005 Regulation of an intergenic transcript controls adjacent gene transcription in Saccharomyces cerevisiae. Genes Dev 19 2695 2704
38. BuratowskiS 2005 Connections between mRNA 3′ end processing and transcription termination. Curr Opin Cell Biol 17 257 261
39. KarchFWeiffenbachBPeiferMBenderWDuncanI 1985 The abdominal region of the bithorax complex. Cell 43 81 96
40. CelnikerSESharmaSKeelanDJLewisEB 1990 The molecular genetics of the bithorax complex of Drosophila: cis - regulation in the Abdominal-B domain. Embo J 9 4277 4286
41. GyurkovicsHGauszJKummerJKarchF 1990 A new homeotic mutation in the Drosophila bithorax complex removes a boundary separating two domains of regulation. Embo J 9 2579 2585 Issn: 0261 : 4189
42. MihalyJBargesSSiposLMaedaRCleardF 2006 Dissecting the regulatory landscape of the Abd-B gene of the bithorax complex. Development 133 2983 2993
43. NagasoHMurataTDayNYokoyamaKK 2001 Simultaneous detection of RNA and protein by in situ hybridization and immunological staining. J Histochem Cytochem 49 1177 1182
44. KarchFGalloniMSiposLGauszJGyurkovicsH 1994 Mcp and Fab-7: molecular analysis of putative boundaries of cis-regulatory domains in the bithorax complex of Drosophila melanogaster. Nucleic Acids Res 22 3138 3146 Issn: 0305–1048
45. HolohanEEKwongCAdryanBBartkuhnMHeroldM 2007 CTCF genomic binding sites in Drosophila and the organisation of the bithorax complex. PLoS Genet 3 e112 doi:10.1371/journal.pgen.0030112
46. CavenerDRRaySC 1991 Eukaryotic start and stop translation sites. Nucleic Acids Res 19 3185 3192
Štítky
Genetika Reprodukčná medicína
Článek Functional Centromeres Determine the Activation Time of Pericentric Origins of DNA Replication inČlánek Dynamic Deposition of Histone Variant H3.3 Accompanies Developmental Remodeling of the TranscriptomeČlánek Integrin α PAT-2/CDC-42 Signaling Is Required for Muscle-Mediated Clearance of Apoptotic Cells inČlánek Prdm5 Regulates Collagen Gene Transcription by Association with RNA Polymerase II in Developing BoneČlánek Acquisition Order of Ras and p53 Gene Alterations Defines Distinct Adrenocortical Tumor Phenotypes
Článok vyšiel v časopisePLOS Genetics
Najčítanejšie tento týždeň
2012 Číslo 5- Gynekologové a odborníci na reprodukční medicínu se sejdou na prvním virtuálním summitu
- Je „freeze-all“ pro všechny? Odborníci na fertilitu diskutovali na virtuálním summitu
-
Všetky články tohto čísla
- Slowing Replication in Preparation for Reduction
- Chromosome Pairing: A Hidden Treasure No More
- Loss of Imprinting Differentially Affects REM/NREM Sleep and Cognition in Mice
- Six Novel Susceptibility Loci for Early-Onset Androgenetic Alopecia and Their Unexpected Association with Common Diseases
- Regulation by the Noncoding RNA
- UDP-Galactose 4′-Epimerase Activities toward UDP-Gal and UDP-GalNAc Play Different Roles in the Development of
- Deletion of PTH Rescues Skeletal Abnormalities and High Osteopontin Levels in Mice
- Karyotypic Determinants of Chromosome Instability in Aneuploid Budding Yeast
- Genome-Wide Copy Number Analysis Uncovers a New HSCR Gene:
- MicroRNA-277 Modulates the Neurodegeneration Caused by Fragile X Premutation rCGG Repeats
- Functional Centromeres Determine the Activation Time of Pericentric Origins of DNA Replication in
- Dynamic Deposition of Histone Variant H3.3 Accompanies Developmental Remodeling of the Transcriptome
- Scientist Citizen: An Interview with Bruce Alberts
- YY1 Regulates Melanocyte Development and Function by Cooperating with MITF
- Congenital Heart Disease–Causing Gata4 Mutation Displays Functional Deficits
- Recombination Drives Vertebrate Genome Contraction
- KATNAL1 Regulation of Sertoli Cell Microtubule Dynamics Is Essential for Spermiogenesis and Male Fertility
- Re-Patterning Sleep Architecture in through Gustatory Perception and Nutritional Quality
- Using Whole-Genome Sequence Data to Predict Quantitative Trait Phenotypes in
- Genome-Wide Analysis of GLD-1–Mediated mRNA Regulation Suggests a Role in mRNA Storage
- Meiotic Chromosome Pairing Is Promoted by Telomere-Led Chromosome Movements Independent of Bouquet Formation
- LINT, a Novel dL(3)mbt-Containing Complex, Represses Malignant Brain Tumour Signature Genes
- The H3K27 Demethylase UTX-1 Is Essential for Normal Development, Independent of Its Enzymatic Activity
- Suppresses Senescence Programs and Thereby Accelerates and Maintains Mutant -Induced Lung Tumorigenesis
- Genome-Wide Association of Pericardial Fat Identifies a Unique Locus for Ectopic Fat
- An Essential Role for Katanin p80 and Microtubule Severing in Male Gamete Production
- Identification of Genes That Promote or Antagonize Somatic Homolog Pairing Using a High-Throughput FISH–Based Screen
- Principles of Carbon Catabolite Repression in the Rice Blast Fungus: Tps1, Nmr1-3, and a MATE–Family Pump Regulate Glucose Metabolism during Infection
- Integrin α PAT-2/CDC-42 Signaling Is Required for Muscle-Mediated Clearance of Apoptotic Cells in
- Histone H3 Localizes to the Centromeric DNA in Budding Yeast
- Collapse of Telomere Homeostasis in Hematopoietic Cells Caused by Heterozygous Mutations in Telomerase Genes
- Hypersensitive to Red and Blue 1 and Its Modification by Protein Phosphatase 7 Are Implicated in the Control of Arabidopsis Stomatal Aperture
- Extent, Causes, and Consequences of Small RNA Expression Variation in Human Adipose Tissue
- TBC-8, a Putative RAB-2 GAP, Regulates Dense Core Vesicle Maturation in
- Regulating Repression: Roles for the Sir4 N-Terminus in Linker DNA Protection and Stabilization of Epigenetic States
- Common Genetic Determinants of Intraocular Pressure and Primary Open-Angle Glaucoma
- Prdm5 Regulates Collagen Gene Transcription by Association with RNA Polymerase II in Developing Bone
- Fitness Landscape Transformation through a Single Amino Acid Change in the Rho Terminator
- Repeated, Selection-Driven Genome Reduction of Accessory Genes in Experimental Populations
- Allelic Variation and Differential Expression of the mSIN3A Histone Deacetylase Complex Gene Promote Mammary Tumor Growth and Metastasis
- DNA Demethylation and USF Regulate the Meiosis-Specific Expression of the Mouse
- Knowledge-Driven Analysis Identifies a Gene–Gene Interaction Affecting High-Density Lipoprotein Cholesterol Levels in Multi-Ethnic Populations
- A Duplication CNV That Conveys Traits Reciprocal to Metabolic Syndrome and Protects against Diet-Induced Obesity in Mice and Men
- EMT Inducers Catalyze Malignant Transformation of Mammary Epithelial Cells and Drive Tumorigenesis towards Claudin-Low Tumors in Transgenic Mice
- Inactivation of a Novel FGF23 Regulator, FAM20C, Leads to Hypophosphatemic Rickets in Mice
- Genome-Wide Association for Abdominal Subcutaneous and Visceral Adipose Reveals a Novel Locus for Visceral Fat in Women
- Stratifying Type 2 Diabetes Cases by BMI Identifies Genetic Risk Variants in and Enrichment for Risk Variants in Lean Compared to Obese Cases
- New Insight into the History of Domesticated Apple: Secondary Contribution of the European Wild Apple to the Genome of Cultivated Varieties
- Activated Cdc42 Kinase Has an Anti-Apoptotic Function
- The Region Is Critical for Birth Defects and Electrocardiographic Dysfunctions Observed in a Down Syndrome Mouse Model
- COP9 Signalosome Integrity Plays Major Roles for Hyphal Growth, Conidial Development, and Circadian Function
- Bmps and Id2a Act Upstream of Twist1 To Restrict Ectomesenchyme Potential of the Cranial Neural Crest
- Psip1/Ledgf p52 Binds Methylated Histone H3K36 and Splicing Factors and Contributes to the Regulation of Alternative Splicing
- The Number of X Chromosomes Causes Sex Differences in Adiposity in Mice
- Target Gene Analysis by Microarrays and Chromatin Immunoprecipitation Identifies HEY Proteins as Highly Redundant bHLH Repressors
- Acquisition Order of Ras and p53 Gene Alterations Defines Distinct Adrenocortical Tumor Phenotypes
- ELK1 Uses Different DNA Binding Modes to Regulate Functionally Distinct Classes of Target Genes
- Histone H1 Depletion Impairs Embryonic Stem Cell Differentiation
- IDN2 and Its Paralogs Form a Complex Required for RNA–Directed DNA Methylation
- Separation of DNA Replication from the Assembly of Break-Competent Meiotic Chromosomes
- Genomic Hypomethylation in the Human Germline Associates with Selective Structural Mutability in the Human Genome
- PLOS Genetics
- Archív čísel
- Aktuálne číslo
- Informácie o časopise
Najčítanejšie v tomto čísle- Inactivation of a Novel FGF23 Regulator, FAM20C, Leads to Hypophosphatemic Rickets in Mice
- Genome-Wide Association of Pericardial Fat Identifies a Unique Locus for Ectopic Fat
- Slowing Replication in Preparation for Reduction
- An Essential Role for Katanin p80 and Microtubule Severing in Male Gamete Production
Prihlásenie#ADS_BOTTOM_SCRIPTS#Zabudnuté hesloZadajte e-mailovú adresu, s ktorou ste vytvárali účet. Budú Vám na ňu zasielané informácie k nastaveniu nového hesla.
- Časopisy



