-
Články
- Časopisy
- Kurzy
- Témy
- Kongresy
- Videa
- Podcasty
Comprehensive Analysis of Transcriptome Variation Uncovers Known and Novel Driver Events in T-Cell Acute Lymphoblastic Leukemia
RNA-seq is a promising technology to re-sequence protein coding genes for the identification of single nucleotide variants (SNV), while simultaneously obtaining information on structural variations and gene expression perturbations. We asked whether RNA-seq is suitable for the detection of driver mutations in T-cell acute lymphoblastic leukemia (T-ALL). These leukemias are caused by a combination of gene fusions, over-expression of transcription factors and cooperative point mutations in oncogenes and tumor suppressor genes. We analyzed 31 T-ALL patient samples and 18 T-ALL cell lines by high-coverage paired-end RNA-seq. First, we optimized the detection of SNVs in RNA-seq data by comparing the results with exome re-sequencing data. We identified known driver genes with recurrent protein altering variations, as well as several new candidates including H3F3A, PTK2B, and STAT5B. Next, we determined accurate gene expression levels from the RNA-seq data through normalizations and batch effect removal, and used these to classify patients into T-ALL subtypes. Finally, we detected gene fusions, of which several can explain the over-expression of key driver genes such as TLX1, PLAG1, LMO1, or NKX2-1; and others result in novel fusion transcripts encoding activated kinases (SSBP2-FER and TPM3-JAK2) or involving MLLT10. In conclusion, we present novel analysis pipelines for variant calling, variant filtering, and expression normalization on RNA-seq data, and successfully applied these for the detection of translocations, point mutations, INDELs, exon-skipping events, and expression perturbations in T-ALL.
Published in the journal: Comprehensive Analysis of Transcriptome Variation Uncovers Known and Novel Driver Events in T-Cell Acute Lymphoblastic Leukemia. PLoS Genet 9(12): e32767. doi:10.1371/journal.pgen.1003997
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1003997Summary
RNA-seq is a promising technology to re-sequence protein coding genes for the identification of single nucleotide variants (SNV), while simultaneously obtaining information on structural variations and gene expression perturbations. We asked whether RNA-seq is suitable for the detection of driver mutations in T-cell acute lymphoblastic leukemia (T-ALL). These leukemias are caused by a combination of gene fusions, over-expression of transcription factors and cooperative point mutations in oncogenes and tumor suppressor genes. We analyzed 31 T-ALL patient samples and 18 T-ALL cell lines by high-coverage paired-end RNA-seq. First, we optimized the detection of SNVs in RNA-seq data by comparing the results with exome re-sequencing data. We identified known driver genes with recurrent protein altering variations, as well as several new candidates including H3F3A, PTK2B, and STAT5B. Next, we determined accurate gene expression levels from the RNA-seq data through normalizations and batch effect removal, and used these to classify patients into T-ALL subtypes. Finally, we detected gene fusions, of which several can explain the over-expression of key driver genes such as TLX1, PLAG1, LMO1, or NKX2-1; and others result in novel fusion transcripts encoding activated kinases (SSBP2-FER and TPM3-JAK2) or involving MLLT10. In conclusion, we present novel analysis pipelines for variant calling, variant filtering, and expression normalization on RNA-seq data, and successfully applied these for the detection of translocations, point mutations, INDELs, exon-skipping events, and expression perturbations in T-ALL.
Introduction
T-cell acute lymphoblastic leukemia (T-ALL) is an aggressive malignancy that accounts for approximately 15% of pediatric and 25% of adult ALL cases. Despite improved outcome over the years, about 25% of children and 50% of adults still fail to respond to intensive chemotherapy protocols or relapse [1]. Improved understanding of T-ALL biology through the identification and characterization of oncogenic lesions is expected to lead to a better prognostic classification and the development of new targeted therapeutic strategies.
T-ALL is caused by the accumulation of multiple oncogenic mutations that have been identified through characterization of chromosomal aberrations and candidate gene sequencing [2]. Chromosomal translocations in T-ALL frequently involve the T-cell receptor (TCR) loci, whereby TCR regulatory elements become juxtaposed to genes that are normally not expressed in T-cells [3], [4]. In this way, a specific set of recurrently over-expressed transcription factors (TFs) have been documented, including TLX1, TLX3, TAL1, LMO1, HOXA, and NKX family members [5]. T-ALL samples expressing each of these transcription factors show a distinctive gene expression signature and as such these transcription factors define distinct molecular subtypes in T-ALL [6]. Chromosomal rearrangements can also lead to large chromosomal deletions and amplifications; to focal gene deletions or amplifications, such as CDKN2A deletion and MYB duplication [7], [8]; and to in-frame fusion genes encoding chimeric proteins with oncogenic properties such as the constitutively active NUP214-ABL1 fusion kinase [9]. In addition, point mutations and small insertions/deletions (INDELs) have also been described leading to oncogenic events, such as mutations activating NOTCH1 that occur in more than 60% of T-ALL cases [10], or mutations in cytokine receptors and tyrosine kinases such as IL7R and JAK3 [11]–[17]. The latter may lead to new opportunities for molecularly tailored therapies with kinase inhibitors [12], [16], [18], [19].
With the advent of next generation sequencing (NGS) technologies, our sequencing capacity has significantly improved in the past five years. It is now possible to apply targeted re-sequencing, exome sequencing (Exome-seq), whole genome sequencing (WGS), whole transcriptome sequencing (RNA-seq) or a combination of these, to investigate individual genomes, especially those related to disease [20]. Also for T-ALL, these NGS approaches have recently proven their value in the discovery of novel driver genes [13], [14], [17], [21]. We previously identified a spectrum of new oncogenic driver genes using Exome-seq on 67 T-ALLs, and described clear differences between pediatric and adult patients [17]. In particular, we identified CNOT3 as a tumor suppressor mutated in 8% of adult T-ALL cases and mutations affecting the ribosomal proteins RPL5 and RPL10 in 10% of pediatric T-ALLs [17]. Similarly, whole genome sequencing of early T-cell precursor ALL cases led to the identification of mutations in several new oncogenes and tumor suppressor genes affecting cytokine signaling, T-cell development and histone-modifying genes [2], [13]. However, the potential of RNA-seq for the discovery of driver genes in T-ALL remains unexplored.
In the present study, we applied paired-end RNA-seq on 49 T-ALL samples (31 patients, 18 cell lines) to gain insights in the transcriptome landscape of T-ALL. First, we show that identification of somatic single nucleotide variants (SNV) and recurrently mutated driver genes is feasible on RNA-seq data, even without matched normal samples (e.g., germlines or remission DNA). We identify STAT5B, H3F3A, and PTK2B as candidate cancer genes in T-ALL. This becomes possible when (1) optimal read mapping and SNV calling procedures are applied; and (2) functional annotation, gene expression, or additional sequencing data from other cohorts is used to prioritize the true driver genes. Next, we optimized gene expression measurements using multiple normalization strategies, and showed that classical gene expression studies (e.g., clustering) are feasible on normalized RNA-seq data. We also detected new fusion genes (SSBP2-FER and TPM3-JAK2) and used gene expression data to determine the consequence of observed chromosomal rearrangements on the over-expression of key driver genes. Finally, we searched for significant alternative transcript events (ATE) but besides one coherent exon-skipping event in SUZ12, we found relatively few candidate ATEs in T-ALL. In conclusion, through a combination of the analysis of gene expression levels, fusion transcripts, SNVs, and INDELs, we could identify known and new driver alterations in T-ALLs and novel potential targets for therapy.
Results
Correct SNV and INDEL calling on RNA-seq data depends on accurate read mapping
We performed paired-end RNA-seq on 31 T-ALL patients, 18 T-ALL cell lines, and 1 normal thymus sample. We obtained on average ∼110 million reads per sample, leading to an average coverage of ∼88× (Table S1.A). To assess the quality of detecting SNVs from the RNA-seq data, we compared the RNA-seq to Exome-seq data. For 16/18 of the cell lines and for 20/31 patient samples we had exome data available (previously generated [17] or obtained for this study, Table S2). For the exome data analysis, we followed the pipeline of mapping, SNV and somatic mutation detection that we validated previously [17] (using BWA, GATK, SomaticSniper, and Variant Effect Predictor (VEP)) [22]–[25]. For the RNA-seq data we used TopHat2 [26] for mapping, SAMTools [27] for SNV detection, and VEP [25] for variant annotation (Figure 1.A).
Fig. 1. RNA-seq data analysis pipelines for (A) variant calling and filtering to detect point mutations, (B) fusion detection and annotation, (C) gene expression analysis. 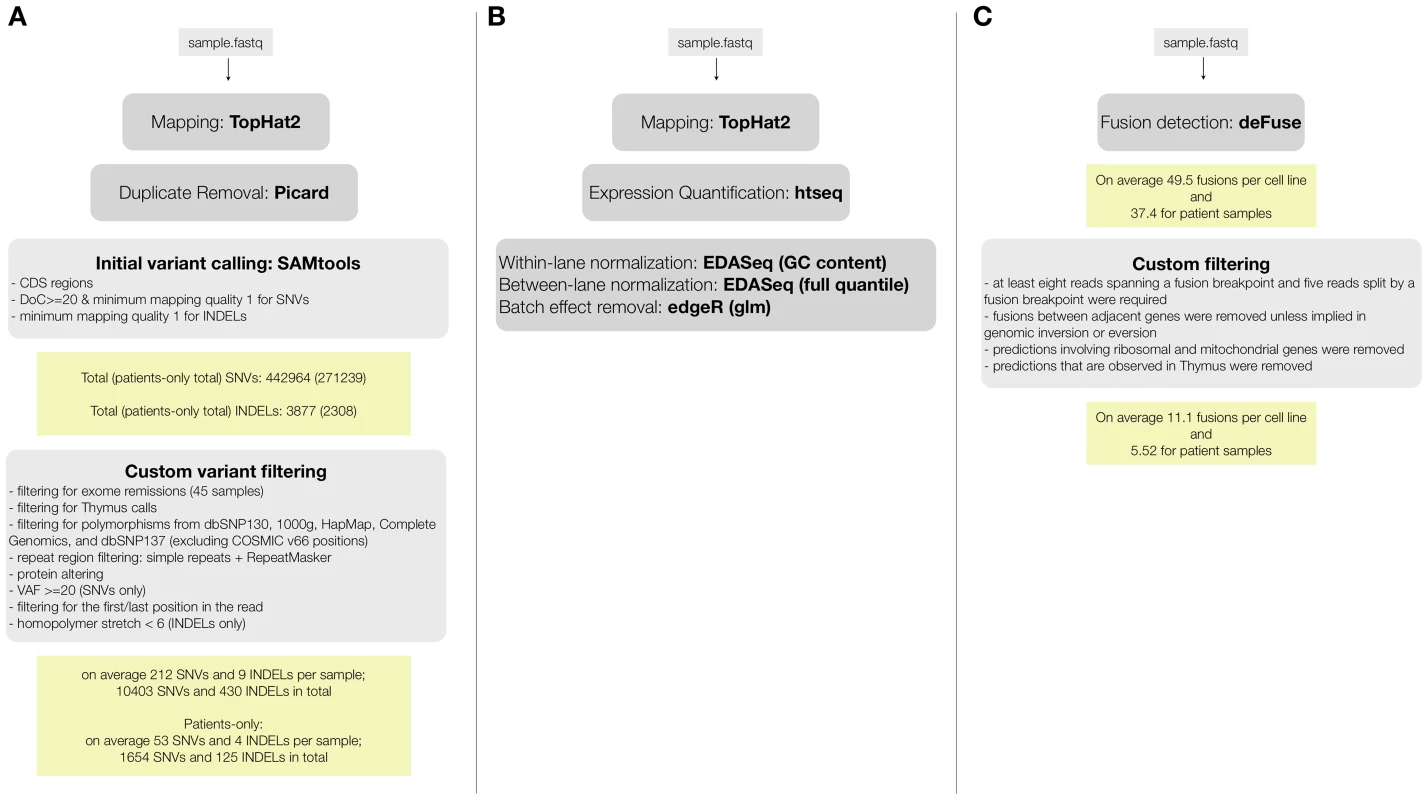
By comparing positions that had a coverage of at least 20× in both RNA-seq and Exome-seq, combined with Sanger re-sequencing of a subset of positions, we found that the accuracy of SNV calling in RNA-seq strongly depends on the read mapping, corroborating earlier observations [28], [29] (Figure S1). We found that mapping RNA-seq reads to the genome (as used by TopHat version 1.3.3) is prone to errors when dealing with paralogous genes, as observed by the prediction of false positive SNVs in KIF4A and GLUD1 due to erroneous mapping to KIF4B and GLUD2 (both pseudogenes with no introns) (Figure S1). However, these errors were resolved by mapping to the transcriptome. In the case of the RPMI8402 cell line, 877 SNVs were found by mapping to the genome, while this number was reduced to 283 SNVs when mapping to the transcriptome. Mapping to the transcriptome did not only reduce the number of RNA-seq exclusive calls but also increased the overlap with the Exome-seq calls (Figure 2, Figure S2).
Fig. 2. Comparison between RNA-seq and exome-seq. 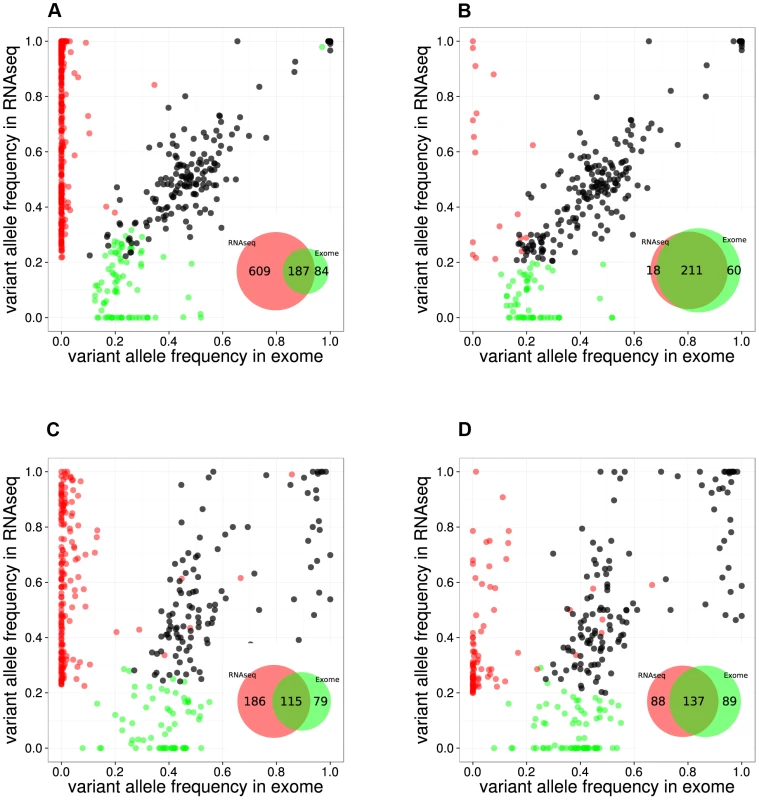
Variant Allele Frequency plots for evaluating two RNA-seq mapping strategies for two example samples, namely the RPMI8402 cell line (A, B) and the TLE79 patient sample (C, D). On the left are the results of mapping with TopHat 1.3.3. (A,C), while on the right are the results of mapping with TopHat 2.0.5 with forced re-mapping of all reads to the genome. The SNVs that have at least 20 reads in exome-seq and RNA-seq are plotted. Red and green dots represent the SNVs that are detected only in RNA-seq and only in exome-seq, respectively, while black dots represent the SNVs that are called in both. Venn diagrams are produced from the points represented in the graphs. However, transcriptome mapping also has limitations as it relies on current gene and isoform annotation. We observed that a combination of transcriptome and genome mapping provides the best solution. It is important that all reads are mapped twice to the genome, independently of each other; once as entire read and once as split read. This has become possible in TopHat2 by setting the option “read-realign-edit-dist” to zero. Our analysis reveals that this mapping approach results in the best overlap of SNVs compared to exomes (Figure 2, Figure S3). This mapping strategy not only improves the alignment accuracy by preventing misalignment to pseudogenes, but also leads to identification of the most likely isoform structure of a gene by mapping the reads independently both to the transcriptome and to the genome and then selecting the best possible alignment.
Using the optimized mapping and filtering strategy we identified 436,974 SNVs across 49 samples. By using samples for which both the exome and the transcriptome were sequenced several aspects of SNV detection in RNA-seq data can be evaluated, such as sensitivity, specificity, and allelic imbalance. Regarding sensitivity, we found that on average, 32% of the SNVs that are called in Exome-seq were also called by the RNA-seq (Table S3). Similar ratios were observed when comparing validated somatic SNVs from Exome-seq/WGS to RNA-seq SNVs: 36% in a triple negative breast cancer study [30], and 41% in a lymphoma study [31]. We observed that the sensitivity varies considerably between samples, and strongly correlates with the average depth of coverage of the sample (Figure S4). Regarding specificity, we found that the remaining RNA-seq-only and Exome-seq-only SNVs (for positions where both have at least 20× coverage) are found mainly with a low variant allele frequency (VAF) and are therefore likely due to arbitrary VAF and coverage thresholds. For example, on the RPMI8402 and TLE79 samples, many RNA-seq-only SNVs (9/18 and 61/88 respectively) have a VAF below 40%. Regarding allelic imbalance, we found that of all heterozygous Exome SNVs with more than 20× coverage, the majority (2,914/4,043 or 72%) were also heterozygous SNVs in RNA-seq. Of the remaining SNVs, many (988/4,043) are homozygous reference in the RNA-seq (i.e., not detected). A small fraction we can almost certainly attribute to allelic imbalance, namely the 141/4,043 SNVs (3.5%) that are homozygous variant in the RNA-seq, indicating that for those only the variant allele is expressed (or the gene is only expressed in cancer cells that harbor the variant).
Next we asked whether small insertions and deletions (INDELs) can be detected from RNA-seq data. As with the SNVs, we used the Exome-seq data for assessing the quality of our INDEL detection strategy. On average, 47.5% of the INDELs that were detected by RNA-seq were also found in the Exome-seq (unfiltered) INDEL calls. However, only 4% of the Exome-seq INDELs (for which the region containing INDEL is covered by at least 3 reads in RNAseq data) were found back in the RNA-seq calls (Table S3). To investigate this sensitivity issue, we evaluated ten validated INDELs that we previously identified with Exome-Seq [17](Table S4). Three of the ten INDELs were also identified in the RNA-seq data using the default SAMTools parameters (see Materials and Methods). Of the seven missed INDELS, two are found in a gene that is not expressed; another two are clearly present in the RNA-seq data when inspected manually with IGV, but did not reach the default threshold (see Materials and Methods); and the last three are effectively discordant between RNA-seq and Exome-seq, as they show only reads with reference sequence (Figure S5). Re-mapping of the reads with BWA [22] on the transcriptome followed by BLAT [32] on the genome improved the INDEL identification, now revealing the KDM6A INDEL in TLE87 and PTEN INDEL in TLE92, which were previously missed (Figure S6.A–B). It is notable that the combination of TopHat2 (to transcriptome only) and BLAT does not correctly detect these two INDELS (Figure S6.C–D). We conclude that INDEL detection on RNA-seq data is feasible, yet technically challenging and that the fraction of INDELs compared to SNVs is moderate (see also the next Section and Figure 3).
Fig. 3. Point mutations and gene fusions organized into functional categories. 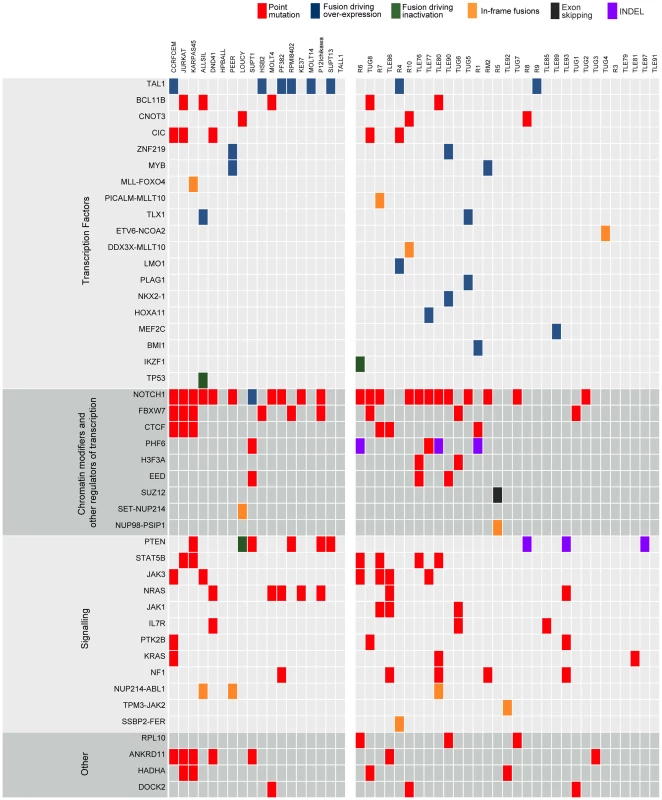
Protein altering mutations and INDELs, alternative splicing events and validated fusions are shown. Red boxes indicate protein-altering mutations (i.e. nonsense, missense and splice site mutations); purple boxes indicate frame-shift INDELs whereas blue, green and orange boxes represent fusion events resulting in over-expression of the partner gene, inactivation of the partner gene or generation of a chimeric protein, respectively, and finally black boxes indicating alternative splicing events. Leveraging diagnosis-only RNA-seq data with the T-ALL body of knowledge to identify mutated cancer genes
Our next aim was to select candidate driver genes using the collected SNVs and INDELS. To remove germline variants we initially removed all SNPs present in dbSNP [33], 1000genomes [34], the Complete Genomics genomes [35], and those detected in our own exome data from normal samples (39 from our earlier work [17] and 6 from this study). We, however, retained those variants also present in the COSMIC [36] database, since SNP databases are known to contain also some disease-specific SNVs. Some examples of SNVs that are likely driver mutations, but that are also present in polymorphism databases are: JAK3 A572V in R7, and FBXW7 R425C in TUG1. With this filtering, we obtained a final list of 10,403 protein-altering SNVs and 430 protein-altering INDELs, with a median of 63 SNVs and 4 INDELS per sample (Table S1.B). Cell lines harbored significantly more mutations than patient samples (Mann-Whitney test p-value = 1.095E-05), as previously also observed by Exome-seq [17].
As a first approach to identify candidate T-ALL driver genes, we selected all genes that contained a protein-altering mutation in at least two of the 31 patient samples (for recurrence we did not take cell lines into account). This process resulted in the selection of 213 genes (Table S5). We found that this list is strongly enriched for genes related to T-ALL and to cancer in general, with “precursor T-cell lymphoblastic leukemia-lymphoma” as the most highly enriched function (p-value = 1.35E-11 by Ingenuity Pathway Analysis) (Table S6). The list of 213 candidates contained many known T-ALL driver genes (Figure 3), such as NOTCH1, BCL11B, FBXW7, IL7R, JAK1 and JAK3; and it also contained the drivers CNOT3 and RPL10, recently identified in our exome re-sequencing study [17]; and CTCF, which was recently reported to be recurrently mutated in ETP-ALL [13]. In addition, the candidate list contained two established cancer driver genes involved in other cancer types, but not yet reported to be mutated in T-ALL, namely H3F3A and CIC. These genes were reported recently by Vogelstein [37] to be true cancer drivers. We identified two patient samples (TLE76 and TUG6) with H3F3A mutations both on the K28 residue that is a mutational hotspot in glioblastoma [38]. This mutation was confirmed somatic in the TUG6 sample. Sequencing of this hotspot in additional T-ALL samples indicated a low frequency of H3F3A K28 mutation in T-ALL (detected in 3 of 102 cases).
Next we asked if we could identify additional genes in the candidate list that could be linked to T-ALL. We wanted to utilize the genes that are known to be involved in T-ALL as a guide for identifying additional candidates. To this end we used our gene prioritization approach ENDEAVOUR [39], which scores candidate genes based on a set of training genes. It builds a profile based on the training genes (integrating information on protein-protein interactions, genetic interactions, gene expression, text-mining, sequence homology, Gene Ontology, and protein domains) and then prioritizes the candidate genes for their similarity to the derived profile. As training set we used all known drivers, and as test set we used all the 213 candidates with at least two patient mutations (excluding the genes that are in the training set). We reasoned that this would reveal the genes with strong similarity to the known drivers and such genes would be good candidate drivers. We found 45 significantly ranked genes with two interesting genes at the top of the ranking, namely PTK2B and STAT5B that are involved in JAK/STAT signaling (Table S7). Furthermore, the list contained genes for which we had identified single T-ALL cases with a somatic mutation in our previous exome study: ANKRD11, CTCF, DOCK2, H3F3A, and HADHA. We did not select these genes before in our Exome-seq cohort [17] because they were only mutated in one of the 39 samples we analyzed. Now, with the RNA-seq cohort, we thus found additional samples with mutations in these genes.
Optimized gene expression measurements and batch effect removal from RNA-seq data identify co-expression modules and T-ALL subtypes
T-ALL is characterized by the overexpression of transcription factors (TFs), such as TLX1, TLX3, TAL1, and the HOXA family members [6]. Therefore, identifying and analyzing expression perturbations in a T-ALL cohort is highly relevant. To obtain accurate gene expression levels from the mapped RNA-seq reads, we followed the procedure outlined in Figure 1.B, including read aggregation, GC-normalization, length normalization, and between-sample normalization (see Materials and Methods). In addition, we removed a batch effect that was clearly present in the data set using a Generalized Linear Model (GLM, see Materials and Methods) (Figure S7). It is notable that transcript-based expression analysis conducted with cufflinks revealed the same batch effect linked to the origin of the sample, thereby confirming a technical bias in the data set (Figure S7.B, see Materials and Methods).
We next looked at the expression values of TLX1, TLX3, TAL1, and other important TFs in T-ALL. Clustering of TLX1, TLX3, and TAL1 expressing samples confirmed that the correct samples (based on karyotyping and molecular analysis) showed over-expression of the respective TF (Figure 4.A). Indeed, 8 samples that harbored a STIL-TAL1 rearrangement showed high TAL1 expression (Figure 4.D). Note that also other samples with high TAL1 expression were detected. This fits with a previously reported observation of TAL1 over-expression in the absence of a translocation in T-ALL [6], [40].
Fig. 4. Validation and discovery using gene expression data, and SUZ12 ATE. 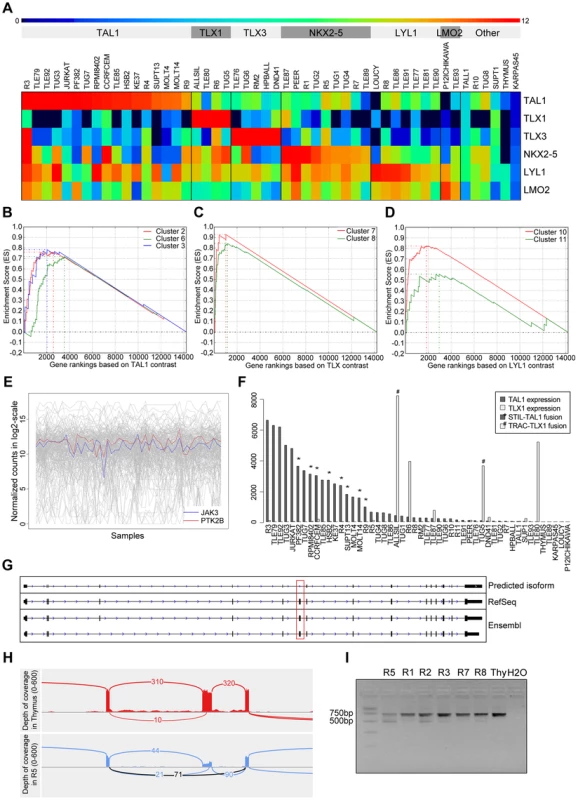
(A) Classification of the samples using the TFs that are known to be overexpressed in T-ALL. Using the expression patterns of TAL1, TLX1, TLX3, NKX2-5, LYL1 and LMO2 we could discriminate the samples in to six distinct clusters. The heatmap is plotted with the normalized log2(count) values. Gene set enrichment analysis curves are displayed for (B) enrichment of TAL1 associated clusters 2, 6 and 3 in TAL1 based ranking, (C) enrichment of TLX associated clusters 7 and 8 in TLX based ranking, and (D) enrichment of LYL1 associated clusters 10 and 11 in LYL1 based ranking of the genes. (E) Expression of JAK3 and PTK2B across samples is significantly correlated (with PTM p-value = 1E-05). (F) Normalized expression values of TAL1 and TLX1 with translocations affecting these genes indicated. The samples with a translocation have elevated expression of the affected gene, showing the driver potential of the fusion event. There are additional samples with high expression of TLX1 and TAL1 without the indicated fusions, pointing to other mechanisms of activating these genes. (G) Predicted SUZ12 transcript aligned with the known SUZ12 isoforms. Dotted red box indicates the location of the exon-skipping event. (H) The sashimi plot shows the junction (in black) supporting the exon-skipping event in patient sample R5 with respect to Thymus. (I) Agarose gel electrophoresis of the RT-PCR products for validation of SUZ12 exon skipping event. The two isoforms are clearly detected in R5 and to a minor extent in the other T-ALL samples while Thymus shows only the canonical transcript. To assess the accuracy of our expression values obtained after normalization, batch effect removal and clustering, we tested whether previously published gene signatures associated with TAL1, TLX (TLX1 and TLX3) and LYL1 can be detected also in our data set [41]. We used 13 gene signatures obtained by Soulier et al using a microarray study on 92 primary T-ALL samples [41]. Gene set enrichment analysis shows that our TAL1 expressing cases are significantly associated with TAL1 signatures, whereas our TLX over-expressing cases are associated with the TLX signature [7], [8] and the LYL1 cases with the LYL1 signature [10], [11]. This analysis confirms that the obtained expression data represent meaningful values and sample clustering produces gene lists that are biologically meaningful (Figure 4.B).
We next used the gene expression information as a guide to assist in the detection of relevant mutations. We found that the expression profile of PTK2B, a candidate driver identified above by ENDEAVOUR, significantly correlated with the JAK3 expression profile (PTM, with p-value threshold at 1E-05, see Materials and Methods) (Figure 4.C). Indeed, PTK2B was previously implicated in IL-2 mediated signaling and JAK/STAT signaling, and was shown to physically interact with JAK3 [42]. These data warrant further investigation of PTK2B as an important tyrosine kinase in T-ALL case with activated JAK/STAT signaling.
T-ALL presents robust transcript isoform usage
To our knowledge, only very few cancer specific alternative transcript events (ATE) have been described for any cancer type [43]–[45], and no ATE is reported for T-ALL. In contrast to SNVs, INDELS, copy number variations, and fusions, which are all curated and present in large numbers in public cancer mutation databases (e.g., COSMIC [36], CENSUS [46]), we could not find driver ATEs in those databases (although splice sites represent an important class of cancer mutations). If ATEs represent an important, yet underestimated, type of somatic variation in cancer, we would expect at least some of the known cancer driver genes to present a significant ATE. We thus asked whether novel variations could be found in these genes in the form of ATEs. To this end, we applied cufflinks and cuffdiff (see Materials and Methods) and found significant ATEs in 12 of the 47 known driver genes (BCL11B, FLT3, IL7R, LCK, MYB, NKX2-1, SFTA3, RPL10, RUNX1, SETD2, SUZ12, and TAL1) (Table S8). However, when we manually inspected these events in IGV, we found only two interesting cases. One case represents an unambiguous skipping of exon 7 in SUZ12, occurring in several patient samples, but most significant (cuffdiff p-value = 5.10E-05) in the R5 patient sample, and absent in the Thymus (Figure 4.E), and a potential, but less clear, skipping of exon 8 in LCK in three samples (Figure S8). Exon 7 of SUZ12 is a canonical exon (present in all known isoforms) according to RefSeq, Ensembl, and UCSC annotation. The ATE we observe is a heterozygous event with the wild-type junction supported by 90 reads and the novel junction supported by 71 reads. RT-PCR clearly confirmed the exon-skipping event in R5 and to a minor extent in other samples, while being absent in the thymus (Figure 4.F). The functional consequences of these splice variants remain to be determined, but the fact that these variants are both in-frame suggests that these proteins could be functional protein isoforms (Figure S8 and S9). Overall, relatively few significant ATEs are detected, and no obvious ATEs are found with consequences on the protein structure, therefore T-ALL presents robust isoform usage at the current resolution of sequencing and analysis.
Detection and validation of known and novel fusion transcripts
Most of the T-ALL cases harbor chromosomal rearrangements that lead to the generation of fusion genes or ectopic expression of genes due to juxtaposition to strong promoters or regulatory sequences. Chromosomal translocations involving the TCR genes are largely underestimated by karyotyping and the TCR partner genes remained unidentified in several cases [4], [47]. On the other hand, a multitude of mechanisms other than translocations could cause ectopic expression of oncogenes [48]. To detect fusion transcripts, we used the defuse algorithm on our entire dataset [49]. Briefly, this method identifies candidate gene fusions by discordant alignments produced by spanning reads (each read in the read pair aligns to a different gene) and by split reads (reads that harbor a fusion boundary). The total number of predicted fusions initially was 1,160 and 1,265 in patient and cell line samples, respectively. Also in normal thymus RNA, 60 fusion transcripts were detected. Next, we implemented additional filters, considering only predictions supported by 8 or more spanning reads and 5 or more split reads. Furthermore, we removed fusions involving ribosomal genes, mitochondrial genes and fusions between adjacent genes, as these could be caused by read-through or trans-splicing [50], [51] (Figure 1.C).
After applying these filters, we obtained an average of 5.5 fusion events per patient sample and 11.1 per cell line (Table S1.C). In total, 397 candidate genes are involved as potential partner in a gene fusion (Table S9). Details on the fusion breakpoints and validation of the novel candidate fusion transcripts are reported in Tables S9 and S12 (see also Materials and Methods: RT-PCR and Sanger Sequencing).
First, to determine the relevance of these predicted fusion transcripts we looked at functional enrichment of these genes. 278 of 397 genes correspond to functionally annotated protein-coding genes according to DAVID functional enrichment [52], [53]. Furthermore, this set is strongly enriched for cancer-related genes, and more specifically for genes involved in Acute Myeloid Leukemia (p-value = 4.48E-10) and T-ALL (p-value = 4.47E-05), including TP53, STAT5B, NOTCH1, IL7R, IKZF1, CDKN2A, MLLT10, ETV6, and ABL1.
Second, we specifically analyzed the 27 in-frame fusions, predicted to encode chimeric proteins (Table S10). This list contained known oncogenic fusion genes, including NUP214-ABL1 (n = 2), MLL-FOXO4 (n = 1), PICALM-MLLT10 (n = 1), ETV6-NCOA2 (n = 1) and SET-NUP214 (n = 1). In addition, we identified 3 novel chimeric transcripts in T-ALL, namely NUP98-PSIP1 (n = 1), TPM3-JAK2 (n = 1) and SSBP2-FER (n = 1) and a novel DDX3X-MLLT10 fusion transcript (n = 1) recently described in a pediatric T-ALL patient [54]. Conventional cytogenetic analysis confirmed the presence of a t(X;10) in the case with the DDX3X-MLLT10 fusion, whereas it failed to detect the chromosomal rearrangements for the TPM3-JAK2, NUP98-PSIP1 and SSBP2-FER fusions, demonstrating the power of RNA-seq to identify cryptic fusion genes and to provide genetic information even in patients with uninformative cytogenetics. Reassuringly, RT-PCR and Sanger sequencing confirmed the presence of these fusion transcripts (Table S12).
The TPM3-JAK2 and SSBP2-FER fusions encode typical tyrosine-kinase fusions that join the tyrosine-kinase domain of JAK2 or FER to the dimerization units of TPM3 or SSBP2, respectively (Figure 5.A). To assess whether the TPM3-JAK2 and SSBP2-FER fusions encode oncogenic proteins, we tested their transforming properties in the IL-3–dependent Ba/F3 cell line [55]. Both TPM3-JAK2 and SSBP2-FER transformed Ba/F3 cells to IL-3–independent growth, with even faster kinetics than the JAK1 A634D mutant, which is a known transforming kinase [18] (Figure 5.B). Western blot analysis confirmed the constitutive auto-phosphorylation of the JAK2 and FER fusion proteins, as well as the downstream STAT proteins (Figure 5.C). Ba/F3 cells transformed by the TPM3-JAK2 fusion were sensitive to a JAK kinase inhibitor, documenting the potential application of JAK2 kinase inhibitors for the treatment of T-ALL cases with JAK2 fusion genes. No specific FER inhibitors were available to test their activity. Both TPM3-JAK2 and SSBP2-FER fusion were screened in 50 additional T-ALL samples, but no additional case with these fusions was found.
Fig. 5. SSBP2-FER and TPM3-JAK2 fusions transform lymphoid cells and show constitutive activity. 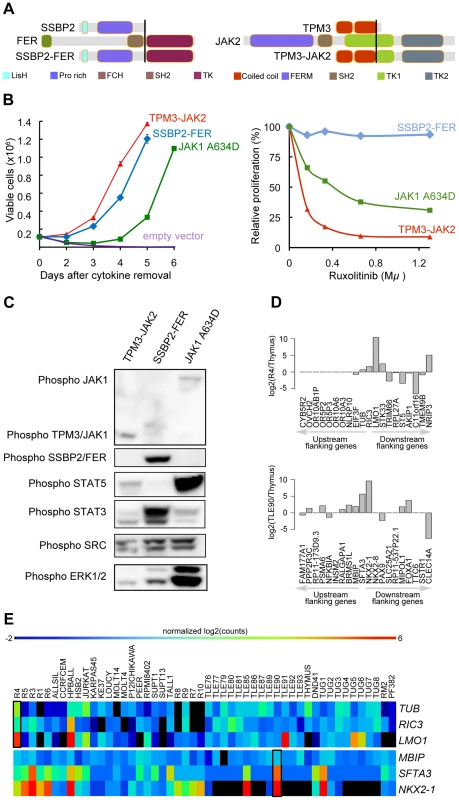
(A) Schematic representations of the predicted SSBP2-FER and TPM3-JAK2 fusion joining the dimerization units of SSBP2 (LisH domain) or TPM3 (coiled-coil domains) to the TK domain of FER or JAK2, respectively. (B) Proliferation curve of mouse Ba/F3 cells in the absence of the cytokine interleukin 3 (IL3) (upper graph) and in the presence of ruxolitinib (lower graph). In the absence of IL3, cells expressing empty vector died whereas cells expressing the SSBP2-FER or TPM3-JAK2 fusion protein were transformed and could proliferate. Ba/F3 cells expressing the oncogenic JAK1 A634D mutant were used as positive control for transformation [18]. The graph shows mean +/− st. dev. The lower graph illustrates the effects of the JAK kinase inhibitor ruxolitinib on Ba/F3 cell proliferation after 24 hours of treatment. The graph represents mean +/− st. dev. of triplicate measurements. (C) Western blot analysis of Ba/F3 cells transformed by the indicated kinases. The 2 upper panels show phosphorylation of the JAK and FER kinases, the panels below illustrate phosphorylation of downstream targets STAT5, STAT3, SRC and ERK1/2. (D) TCR gene fusions result in overexpression of a flanking gene in RIC3-TRBC2 and SFTA3-TRDC fusions. The barplot is drawn for relative (to Thymus) expression values for the upstream and downstream flanking genes around RIC3 and SFTA3 for R4 and TLE90 samples, respectively. In both cases, the nearest downstream neighbor shows increased expression. (E) The heatmap illustrates the expression patterns of RIC3 and SFTA3, together with their immediately upstream and downstream flanking genes in the genome, showing strong over-expression (red) of LMO1 near the RIC3 fusion, and of NKX2-1 near the SFTA3 fusion. Third, we also analyzed the identified fusions that did not seem to encode chimeric proteins (out-of-frame fusions), and which were the majority of fusions detected in T-ALL. These fusion events can be used as surrogate markers for the identification of chromosomal rearrangements, providing accurate information on the precise chromosomal breakpoints. In combination with the gene expression data obtained by RNA-seq, these data can identify genes that are located close to such potential breakpoints and for which the expression is significantly up - or down-regulated. As expected, we identified the STIL-TAL1 fusion in several T-ALL cases (n = 8). We also identified and validated 6 fusion events involving TCR genes. In 4 of these cases, the TCR gene was found to be fused to the potential oncogene (NOTCH1, IL7R, PLAG1, and TLX1). In the two other cases (R4, TLE90), the TCR gene was fused to RIC3 or SFTA3, resulting in the ectopic expression of LMO1 and NKX2-1, respectively, as indicated by RNA-seq gene expression data (Figure 5.D and E). Similarly, we could better characterize the t(10;14) in ALL-SIL cell line that expresses TLX1 at high level.
In addition to the TCR gene rearrangements, also other fusions were associated with overexpression. We detected out-of-frame fusion transcripts that joined exon 4 of CDK6 to exon 2 of HOXA11-AS and exon 5 of CDK6 to sequences downstream of EVX1. In the same patient we also detected a fusion joining DPY19L1 on chromosome 7p14 to HOXA11 on chromosome 7p15. The gene expression analysis documented high expression of genes of the HOXA cluster (i.e. HOXA9, -A5, -A13, -A10, -A11). Moreover, other fusions identified in this study, such as CLINT1-MEF2C, HNRP-ZNF219 (n = 2), ZEB1-BMI1 and AHI1-MYB (n = 2) were also associated with transcriptional activation of MEF2C, ZNF219, BMI1 and MYB as confirmed by the expression data (Table S9 and S12, and Figure S10). Increased MYB expression in T-ALL was previously observed as a consequence of MYB duplication (including in the BE-13 cell line), which may also explain the detected AHI1-MYB fusion [8], [56].
Finally, we also found out-of-frame fusion transcripts leading to the potential inactivation of tumor suppressor genes, such as TP53-TBC1D3F (ALLSIL cell line), PTEN-RNLS (LOUCY cell line), IKZF1-ABCA13 and CDKN2A-miR31HG (R6 case), indicating a third class of fusion events (Figure S10). FISH analysis performed in the R6 case confirmed the p15/p16 deletion. As the genes are in close proximity, the IKZF1-ABCA13 was presumably generated by deletion although no material was available to confirm this hypothesis.
Discussion
The landscape of genomic variation underlying T-ALL has recently been investigated by sequencing candidate genes [14], [21], whole exomes [17] and whole genomes [13]. The results of these studies, combined with a large body of gene-by-gene evidence collected over the last decade, provide a growing comprehension of the T-ALL genome. The T-ALL genome is mainly characterized by the over-expression of TF, such as TLX1/3 and TAL1, in combination with gain-of-function NOTCH1 mutations, and with additional mutations in chromatin modifiers, cellular signaling factors such as those involved in the JAK-STAT signaling pathway [57], tumor suppressor genes (TP53, PTEN, WT1), or in other genes such as ribosomal genes [17]. Since the majority of observed mutations are point mutations and gene fusions (much more than copy number variations [13]) we reasoned that RNA-seq would be effective to identify many of these mutations, certainly those associated with (over-)expressed oncogenes. Indeed, exome sequencing allows identifying point mutations but not gene fusions; and low coverage whole-genome sequencing allows identifying structural variation (gene fusions) but not point mutations. In this study we present RNA-seq analyses on a heterogeneous group of 31 T-ALL samples and 18 T-ALL cell-lines and demonstrate that RNA-Seq is indeed a very powerful approach to detect gene mutations and fusions as well as expression perturbations.
Our first challenge with regards to the accurate identification of point mutations was finding the optimal analysis pipeline – from read mapping to SNV calling and filtering – to avoid too many false positive SNVs. By exploiting whole-exome sequencing data for a subset of our samples we obtained a recovery ratio of 32% when compared to the exome derived SNVs; a ratio that is comparable with previous RNA-seq studies [30], [31]. However, this concordance could only be achieved by using the optimal read mapping methods and parameters: (1) use of a recent version of TopHat2 (v. 2.0.5. or higher) and (2) forcing this aligner to map all reads twice to the genome (once directly and once using split reads) and once to the transcriptome. Indeed, the computational task of sequence read mapping is more challenging for RNA-seq data because a large fraction of the obtained reads need to be split to allow reads that overlap exon-exon boundaries in the cDNA to be mapped to the genome. In this way, RNA-seq is more prone to the identification of false SNVs due to the erroneous mapping of reads, for example to highly similar non-spliced pseudogenes. For example, in the RPMI8402 cell line, 603 RNA-seq exclusive SNVs were found with the genome mapping strategy, while only 35 when using combined mapping strategy.
Among the previously published large scale RNA-seq cancer studies, only a handful performed variant calling on the RNA-seq data [30], [31], [58], [59]. A combined mapping strategy was followed in all cases either by mapping the reads to a customized genome reference file (by the addition of exon junction segments) or mapping the reads twice (once to the genome and once to the transcriptome). Variant calling pipelines also showed diversity: Morin et al and Shah et al used SNVMix [60] for variant calling, while Seo et al and Berger et al implemented filters based on alignment on the non-reference bases. To our knowledge there is no extensive benchmarking study evaluating aligners and variant callers for RNA-seq data, but a review paper by Quinn et al compared the performance of two variant callers (GATK [23] and SAMTools [27]) with the optional duplicate removal step (pre and post alignment), and concluded that post-alignment duplicate removal and variant calling with SAMTools achieved the best performance in terms of sensitivity and specificity [61]. We have also followed the same strategy in our study and we could achieve a comparable recovery ratio of 32% when compared to Exome-seq calls.
A second challenge in identifying point mutations was the prioritization of candidate driver mutations versus passenger mutations. Due to the lack of matched germline RNA for each patient as control, we used a large cohort of local normal exome datasets, in combination with the commonly used variants from dbSNP and 1000genomes, to distinguish SNPs from candidate somatic mutations. This strategy has been successfully used before on transcriptome sequencing studies [62]. Identifying candidate cancer genes by gene mutation frequency is a frequently used approach [13], [30], [58]. Remarkably, by simply selecting all genes having a candidate somatic mutation in at least two samples (213 genes in total), we already achieved a highly significant enrichment for T-ALL related genes, such as NOTCH1, BCL11B, FBXW7, DNM2, JAK3, JAK1, and IL7R. Among the remaining candidates we searched for additional evidence and we propose seven additional candidate drivers because they are either “functionally similar” to the previously known drivers, or because they were mutated somatically at least once in another T-ALL cohort [17], or both. Six of these genes, namely CIC, H3F3A, PTK2B, STAT5B, ANKRD1 and HADHA have already been implicated in other cancers [63]–[70] while DOCK2 has no association with cancer yet.
We found a remarkable clustering of molecular functions among the identified T-ALL driver genes, with enrichment for functions related to the regulation of gene expression. TFs and their co-factors play a central role in transcriptional regulation and these proteins are often mutated in T-ALL. Also, many of these play important roles in the normal T-cell developmental gene regulatory network [71], such as NOTCH1, TLX1, TLX3, TAL1, BCL11B, CTCF, FOXO4, MYB, and others. Upstream of these activated TFs, multiple kinases and other signaling factors control their activity, and these regulators are also often mutated in T-ALL (for example, JAK1, JAK3, and IL7R). Finally, chromatin modifiers and methylation factors are recurrently mutated and these can have both generally pervasive but also specific effects on the expression of oncogenes, such as MYC [72]. When multiple driver mutations are serially acquired, their combined effect will result in oncogenic expression profiles, whereby genes supporting a growth advantage increase and genes negatively affecting growth advantage (e.g., apoptosis, senescence) decrease in expression. It will be an interesting future challenge to draw the connections between the observed DNA mutations, the oncogenic program, and the final gene expression changes that we and others observe in T-ALL samples. Finally, it is likely that non-coding mutations, such as those in promoters, enhancers, microRNAs, and lncRNAs, add to the cancer-related gene regulatory network changes underlying leukemogenesis.
As mentioned above, only mutations in genes that are actively transcribed are detected, and this likely adds to the specificity of driver gene detection. On the other hand, this could also present a limitation of RNA-seq, because loss-of-function mutations in tumor suppressor genes may lead to nonsense-mediated decay, and as consequence low sequence coverage to call mutations. Based on our data however, this is not the case because we could detect PHF6 mutations in up to 4/31 patient cases (13%), where exome sequencing identified PHF6 mutations in 9/67 cases (13%) [17] and Zhang et al identified PHF6 mutations in 24/106 cases by means of whole genome sequencing and capillary sequencing [13].
Interestingly, the gene expression information used above (i.e., read coverage to identify point mutations) can be further exploited at the quantitative level, similar to gene expression studies performed with microarray technology over the last 15 years. As many leukemia driver genes are characterized by changes in gene expression, this level of information is invaluable, both in research and diagnostic settings. We investigated how accurate gene expression levels can be achieved and we found that multiple normalization steps are required, both within-sample (gene length and gene GC content) and across samples (library size), and that batch effects can be effectively removed using a previously published Generalized Linear Model (GLM) [73]. The gene expression levels of the known drivers (e.g., TLX1/3, TAL1, NOTCH1) are highly representative as driving events and as subtype identifiers. However, to discover driver genes de novo, using only gene expression values, is to our opinion not feasible (data not shown). Alternatively, we attempted to select candidate drivers based on the expression similarity (i.e., co-expression across the cohort) with known drivers. This led to the identification of PTK2B, whose expression strongly correlated with JAK3 and which is known to be implicated in JAK-STAT signaling. The next level of gene expression analysis would preferably be a network-level analysis [74], but this requires a larger sample cohort.
Another kind of information that can be extracted from RNA-seq data, besides point mutations and gene expression changes, are alternative transcript events (ATE) and gene fusions [75]. We found only few significant ATEs but could confirm two exon-skipping events in the known T-ALL oncogenes SUZ12 and LCK. More importantly, we identified (i) known and novel in-frame fusions encoding chimeric proteins, (ii) TCR gene arrangements resulting in over-expression of oncogenes, and (iii) fusions not involving TCR genes but also resulting in over-expression of oncogenic transcription factors. The most recurrent fusion event, observed in 8/31 samples, was the STIL-TAL1 fusion resulting in the ectopic over-expression of the TAL1 gene. We also identified novel gene fusions, including two in-frame fusions, TPM3-JAK2 and SSBP2-FER, producing chimeric oncoproteins; and other fusions resulting in the ectopic expression of transcription factors such as PLAG1, MEF2C, ZNF219, and BMI1. The ectopic expression of these genes is associated with a fusion event and with changed expression, which can both be detected by RNA-seq, making this technology extremely powerful to accurately detect such oncogenic events. Each of these novel events appears to be rare in T-ALL, as we identified at most 2 cases of each fusion. However the evidence of transcriptional activation of the partner genes suggests that further studies are required to establish the recurrence of these lesions and their functional meaning. It is notable that the normal thymus sample also shows four fusion events. However, as these genes are located in close proximity to each other, they may represent unannotated isoforms in the human transcriptome. Despite RNA-seq has offered a deeper insight into the complexity of the transcriptome, several studies have highlighted that the catalogue of all expressed transcripts is still far from complete and it is increasing the number of novel splice junctions connecting novel exon, non-exon regions, or linking independent transcripts [76].
Today, high-quality catalogues of driver genes across cancer types are available, and this influences how and why cancer genomes need to be sequenced. For T-ALL, and for many common cancer types, the objectives of sequencing are shifting from the discovery of cancer genes, to a diagnostic setting in which a list of driver events are a priori known. Targeted re-sequencing provides an interesting route, although this poses technical challenges of amplification or capturing, and perhaps more importantly, is focused on a limited number of genes and on one particular mutation type, namely point mutations and small insertions/deletions. We have shown in this study that, with a list of interesting cancer drivers at hand, and with other datasets being available (e.g., rare variants from local exome studies, 1000 genomes, TCGA data, etc), RNA-sequencing of only the cancer sample provides a technically straightforward approach and delivers at once the point mutations, gene fusions and gene expression changes across the entire transcriptome. And as a corollary, the data analysis strategies provided here would be beneficial for any cancer type as long as a body of knowledge is available for selecting and prioritizing candidate events.
Materials and Methods
Patient samples and cell lines
Diagnostic total RNAs from 31 T-ALL patients (20 adults and 11 children) were collected at various institutions. All patients have given their informed consent and all samples were obtained according to the guidelines of the local ethical committees. This study was approved by the ethical committee of the University Hospital Leuven. Diagnosis of T-ALL was based on morphology, cytochemistry and immunophenotyping according to the World Health Organization and European Group for the Immunological Characterization of Leukemia criteria [77]. The clinical and hematologic features of the 31 patients at the diagnosis are summarized in Table S11 Total RNAs from 18 T-ALL cell lines (DSMZ, Braunschweig, Germany) were extracted using QIAGEN RNeasy Mini Kit. A pool of total RNAs from 5 normal human thymuses was purchased from Capital Biosciences.
All the RNA samples showed a high quality RNA Integrative Number (RIN>/ = 7) score on the Bioanalyzer (Agilent Technologies).
Fifty additional RNA samples were used for TPM3-JAK2 and SSBP2-FER analysis.
Genomic DNA from of 71 adult T-ALL patients were used for H3F3A K28 screening.
RNA-seq
Next generation sequencing libraries were constructed from 500 ng of total RNA using the Truseq RNA sample prep kit (Illumina). RNA-seq libraries were subjected to 2×100 bp paired-end sequencing on a HiSeq2000 instrument (Illumina). Sequence reads were processed to identify gene fusion transcripts, single nucleotide variants (SNVs) and gene expression levels. For the read mapping, variant calling and transcriptome assembly, we used the infrastructure of the VSC - Flemish Supercomputer Center, funded by the Hercules foundation and the Flemish Government - department EWI.
Fusion transcript discovery
Fusion transcript discovery was performed using defuse v.0.5.0 [49] with default parameters. The resulting list was filtered as described in [78]. Briefly, fusion transcripts with less than 8 spanning reads and less than 5 split reads were filtered out. In addition, we removed fusion events observed in adjacent genes and fusion events involving ribosomal genes (ribosomal genes were downloaded from Biomart on 24-05-2011 using GO:0005840) and the genes located on chrM. Fusion events were annotated using Pegasus (http://sourceforge.net/projects/pegasus-fus/).
Gene expression analysis
For Gene Expression Profiling analysis, reads were mapped to the human reference genome (assembly GRCh37.68) using TopHat v.2.0.5 [26] with the following parameters: transcriptome-only. Read counts per gene were obtained with the HTSeq package (htseq-count) (http://www-huber.embl.de/users/anders/HTSeq). The aggregated read counts were normalized with EDASeq v1.4.0 [79] and generalized linear model was fitted with edgeR v3.0.4 [73] to remove batch effect originating from the sample collection center. The pathways, and upstream regulators were generated through the use of IPA (Ingenuity Systems, www.ingenuity.com). Expression neighbors were detected with Pavlidis Template Matching (PTM) analysis [80]. Transcript based gene expression values were obtained using cufflinks suite [81], [82]. Transcript assembly was performed with cufflinks v2.1.1 with –g option using assembly GRCh37.68.
Gene set enrichment analysis (GSEA) was performed for TAL1, TLX and LYL1 clusters [83]. We have obtained whole genome rankings for TAL1, TLX (TLX1 and TLX3), and LYL1 simply by calculating the log fold changes between samples expressing the respective gene versus the remaining samples. The gene signatures from Soulier et al were obtained from Table S2 [41].
Alternative transcript event discovery
Tumor patient samples and Thymus RNA-Seq samples were mapped to the Ensembl GRCh37.68 reference genome by Tophat2 [26]. Mapped reads were realigned, and transcript abundance were estimated using cufflinks v2.1.1 [81], [82]. Transcript assembly was reconstructed using the cuffmerge program of the cufflinks package from the realigned transfrags for each of patient RNA-seq samples, merged with the Thymus sample (control), followed by differential expression analysis performed using cuffdiff program. The significant events were extracted from the list of differentially expressed genes, isoforms, primary transcripts and coding sequence and assessed manually with IGV [84]. The mRNA sequences for novel SUZ12 and LCK transcripts were extracted using gffread command of cufflinks, and these sequences were translated using the translate tool of the ExPASy Bioinformatics Resource Portal [85]. The longest ORF sequence was used to verify the domain architecture of the resulting proteins using SMART [86], [87].
Prediction of single nucleotide variation
The sequence reads were mapped to the human reference genome (assembly GRCh37.68) using TopHat2 setting the option “read-realign-edit-dist” to zero [26]. Duplicate removal process was performed on the aligned reads using Picard v1.74 (http://picard.sourceforge.net). Then SAMTools package v0.1.19+ (pulled from the git repository on 29-07-2013) [27] was used for single nucleotide variant (SNV) and small insertion and deletion (INDEL) detection with minimum mapping quality threshold of 1 and minimum base quality threshold of 13 (-q 1 -Q 13) [27]. The variant calling was done on the coding regions of the genome only (extracted from the transcript definitions in the assembly GRCh37.68). The variant predictions that were supported exclusively by variants located in the beginning or the end of the read were filtered out. Then the SNVs were further filtered with depth of coverage threshold of 20 and minimum variant allele frequency threshold of 0.20. INDELs predictions were filtered with the SAMTools recommended parameters (varFilter -10 -20 -30 -40 -a4 -G90 -S30) and additionally INDELs located in homopolymer stretches longer than 5 bps were filtered. The high quality list of variants was filtered for common population variants using the calls from 1000 genomes, dbSNP, HapMap, and Complete Genomics. Note that, the list of common population variants was cleaned from oncogenic variants using COSMIC listed variants (v66) [36]. Moreover, the variants located in the repeat regions (simple repeat and RepeatMasker) were filtered out. Finally, the variants that are observed in the exomes of remission (i.e. healthy) samples (including the previously published 39 exome remissions [17] and the 6 additional exome remission sequenced) and the variants that are observed in Thymus were also filtered out. The final filtered list of variants was annotated with the Variant Effect Predictor version 2.7 [25] and the protein-altering mutations were selected. The following terms were used for selecting protein-altering SNVs: splice-donor-variant, splice-acceptor-variant, stop-gained, initiator-codon-variant, missense-variant, splice-region-variant. The same terms were used for filtering the INDELs with the addition of the following terms: inframe-insertion, inframe-deletion, frameshift-variant.
The list of candidate genes was created by intersecting the genes with recurrent mutations (SNVs and INDELs) in RNA-seq patient cohort with the somatic mutations in Exome-seq patient cohort [17]. The list of genes that have recurrent mutations in the RNA-seq patient cohort was filtered for mutations observed in chrM.
The list of T-ALL driver genes were curated using the Census database [46] and T-ALL literature and includes the following genes: TLX1, TLX3, PHF6, MYC, BCL11B, HOXA1, SET, MLL, MLLT1, PICALM, MLLT10, WT1, MYB, LEF1, LMO2, LMO1, TAL1, NUP98, NOTCH1, FBXW7, CCND2, PTEN, PTPN2, NF1, FLT3, JAK1, NRAS, LCK, NUP214, ABL1, EZH2, SETD2, SUZ12, JAK3, MEF2C, NKX2-1, NKX2-2, CDKN2A, CDKN2B, RUNX1, KRAS, EED, ETV6, RPL10, DNM2, IL7R, CNOT3.
Exome-seq analysis
Somatic mutations from the exome pairs were obtained as described previously [17]. Briefly, the alignment was performed with BWA [22] and post-alignment modifications (duplicate removal, realignment around INDELs and calibration of the quality scores) were done with the Genome Analysis Toolkit (GATK) [23]. Variant calling was performed with GATK using Variant Quality Score Recalibration (VQSR) method. Putative somatic variants were identified by subtracting the mutations observed in the primary samples from the mutations observed in the corresponding remission samples. SomaticSniper score above 70 was used to identify the final list of somatic events [24].
Variant allele frequency (VAF) plots were drawn for the positions that are novel SNVs in either of the RNA-seq or Exome-seq data and covered by at least 20 reads in both datasets.
RT-PCR and Sanger sequencing
Novel candidate fusion transcripts were validated by Reverse-Transcription Polymerase-Chain-Reaction (RT-PCR) and Sanger sequencing. In all cases Thymus was used as negative control. cDNA synthesis and PCR amplification were performed using standard protocols that come with Superscript III Reverse Transcriptase (Invitrogen) and GoTaq (Promega). PCR primers were designed to amplify 200–400 bp fragments containing the fusion boundary detected by RNA-seq. The PCR products were analyzed using a QIAxcel automated multicapillary electrophoresis system (QIAGEN). The results were processed and visualized using the BioCalculator Software. PCR products were analyzed by Sanger Sequencing. In cases where multiple PCR products were detected, we performed conventional agarose gel electrophoresis and extraction of specific bands using the gel DNA Recovery Kit (Zymo). Analysis of Sanger chromatograms was performed using CLC Main Workbench 6 (CLC Bio, Aarhus, Denmark). Fusion detection was performed using NCBI Blast alignment. Analysis of the breakpoint was done on the longest isoform reported on the Ensembl genome browser. The tested fusions predictions and the primers used for validations are reported in Table S12.
Validation of SUZ12 exon skipping was performed by RT-PCR, gel extraction and sequencing of the two PCR products (Figure 4.I). The following primers were used for RT-PCR and Sanger sequencing: SUZ12_EX1F (CTGACCACGAGCTTTTCCTC) and SUZ12_EX9R (CCATTTCCTGCATGGCTACT).
Cloning
The plasmid TPM3-JAK2 pMSCV-GFP was obtained as follows: a DNA fragment containing TPM3 coding region till exon 7 was PCR amplified from thymus cDNA using Phusion High Fidelity DNA Polymerase (Finzyme) and primers containing BglII and XhoI restriction sites. Primers containing XhoI and EcoRI restriction sites were used to amplify JAK2 coding exons 17–25. PCR products were cloned into the BglII and EcoRI sites of the pMSCV-GFP vector after subcloning into the pJET1.2 CloneJET vector (Fermentas). As a final control, plasmid DNA was sequenced by Sanger sequencing.
SSBP2-FER fusion was synthesized by Genscript (Piscataway, NJ, USA) and cloned into pMSCV-GFP by using the unique restriction sites XhoI and EcoRI. The plasmid contained the full length SSBP2-FER fusion including the first 16 coding exons of SSBP2 and the coding exons 14–20 of FER.
Cell culture
Viral supernatants were produced in HEK293T cells using an EcoPack packaging plasmid and TurboFect transfection reagent (Fermentas). Viruses were harvested 48 hours after transfection followed by transduction of the Ba/F3 murine pro-B cells (DSMZ, Braunschweig, Germany) as described previously [88].
Transformation experiments
Ba/F3 cells were washed twice in PBS to remove all traces of cytokines and were seeded in triplicate in 24-well dishes at 100 000 cells/mL. GFP expression and cell number were measure on a Guava flow cytometer (Millipore). All experiments were terminated at day 8 after cytokine removal and cell lines showing no sign of cell proliferation at that timepoint were declared to be non-transforming.
Western blotting
Total cell lysates were analyzed by standard electrophoresis and western blotting procedures using the following antibodies: anti-phospho-JAK1 (Tyr1022/1023), anti-phospho-STAT1, anti–phospho-STAT5 (Tyr694), anti–phospho-STAT3 (Tyr705), anti-phosphoERK1-2, anti-phospho-SRC families (Tyr416) (from Cell Signaling Technology).
Inhibitor experiments
TPM3-JAK2 and SSBP2-FER IL3-independent Ba/F3 cells were seeded in triplicate in 96-well plates at a density of 0.03×106 cells in the presence of JAK inhibitor Ruxolitinib (INCB018424, Azon Medchem). Cell proliferation and viability were assessed on a Guava flow cytometer after 24 hours to determine the IC50, the concentration of inhibitor that gave a 50% inhibition.
Accession numbers
Genome data has been deposited at the European Genome-phenome Archive (EGA, http://www.ebi.ac.uk/ega/) which is hosted at the EBI, under accession number EGAS00001000536.
Supporting Information
Zdroje
1. PietersR, CarrollWL (2008) Biology and treatment of acute lymphoblastic leukemia. Pediatr Clin North Am 55 : 1–20–ix doi:10.1016/j.pcl.2007.11.002
2. van VlierbergheP, FerrandoA (2012) The molecular basis of T cell acute lymphoblastic leukemia. J Clin Invest 122 : 3398–3406 doi:10.1172/JCI61269
3. GrauxC, CoolsJ, MichauxL, VandenbergheP, HagemeijerA (2006) Cytogenetics and molecular genetics of T-cell acute lymphoblastic leukemia: from thymocyte to lymphoblast. Leukemia 20 : 1496–1510 doi:10.1038/sj.leu.2404302
4. Le NoirS, Ben AbdelaliR, LelorchM, BergeronJ, SungaleeS, et al. (2012) Extensive molecular mapping of TCRα/δ - and TCRβ-involved chromosomal translocations reveals distinct mechanisms of oncogene activation in T-ALL. Blood 120 : 3298–3309 doi:10.1182/blood-2012-04-425488
5. Van VlierbergheP, HommingaI, ZuurbierL, Gladdines-BuijsJ, van WeringER, et al. (2008) Cooperative genetic defects in TLX3 rearranged pediatric T-ALL. Leukemia 22 : 762–770 doi:10.1038/sj.leu.2405082
6. FerrandoAA, NeubergDS, StauntonJ, LohML, HuardC, et al. (2002) Gene expression signatures define novel oncogenic pathways in T cell acute lymphoblastic leukemia. Cancer Cell 1 : 75–87.
7. SulongS, MoormanAV, IrvingJAE, StreffordJC, KonnZJ, et al. (2009) A comprehensive analysis of the CDKN2A gene in childhood acute lymphoblastic leukemia reveals genomic deletion, copy number neutral loss of heterozygosity, and association with specific cytogenetic subgroups. Blood 113 : 100–107 doi:10.1182/blood-2008-07-166801
8. LahortigaI, de KeersmaeckerK, van VlierbergheP, GrauxC, CauwelierB, et al. (2007) Duplication of the MYB oncogene in T cell acute lymphoblastic leukemia. Nat Genet 39 : 593–595 doi:10.1038/ng2025
9. GrauxC, CoolsJ, MelotteC, QuentmeierH, FerrandoA, et al. (2004) Fusion of NUP214 to ABL1 on amplified episomes in T-cell acute lymphoblastic leukemia. Nat Genet 36 : 1084–1089 doi:10.1038/ng1425
10. WengAP, FerrandoAA, LeeW, MorrisJP, SilvermanLB, et al. (2004) Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306 : 269–271 doi:10.1126/science.1102160
11. ShochatC, TalN, BandapalliOR, PalmiC, GanmoreI, et al. (2011) Gain-of-function mutations in interleukin-7 receptor-α (IL7R) in childhood acute lymphoblastic leukemias. Journal of Experimental Medicine 208 : 901–908 doi:10.1084/jem.20110580
12. ZenattiPP, RibeiroD, LiW, ZuurbierL, SilvaMC, et al. (2011) Oncogenic IL7R gain-of-function mutations in childhood T-cell acute lymphoblastic leukemia. Nat Genet 43 : 932–939 doi:10.1038/ng.924
13. ZhangJ, DingL, HolmfeldtL, WuG, HeatleySL, et al. (2012) The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481 : 157–163 doi:10.1038/nature10725
14. Kalender AtakZ, de KeersmaeckerK, GianfeliciV, GeerdensE, VandepoelR, et al. (2012) High accuracy mutation detection in leukemia on a selected panel of cancer genes. PLoS ONE 7: e38463 doi:10.1371/journal.pone.0038463
15. BainsT, HeinrichMC, LoriauxMM, BeadlingC, NelsonD, et al. (2012) Newly described activating JAK3 mutations in T-cell acute lymphoblastic leukemia. Leukemia 26 : 2144–2146 doi:10.1038/leu.2012.74
16. ElliottNE, ClevelandSM, GrannV, JanikJ, WaldmannTA, et al. (2011) FERM domain mutations induce gain of function in JAK3 in adult T-cell leukemia/lymphoma. Blood 118 : 3911–3921 doi:10.1182/blood-2010-12-319467
17. de KeersmaeckerK, AtakZK, LiN, VicenteC, PatchettS, et al. (2013) Exome sequencing identifies mutation in CNOT3 and ribosomal genes RPL5 and RPL10 in T-cell acute lymphoblastic leukemia. Nat Genet 45 : 186–190 doi:10.1038/ng.2508
18. FlexE, PetrangeliV, StellaL, ChiarettiS, HornakovaT, et al. (2008) Somatically acquired JAK1 mutations in adult acute lymphoblastic leukemia. Journal of Experimental Medicine 205 : 751–758 doi:10.1158/1078-0432.CCR-05-2832
19. PorcuM, KleppeM, GianfeliciV, GeerdensE, de KeersmaeckerK, et al. (2012) Mutation of the receptor tyrosine phosphatase PTPRC (CD45) in T-cell acute lymphoblastic leukemia. Blood 119 : 4476–4479 doi:10.1182/blood-2011-09-379958
20. MeyersonM, GabrielS, GetzG (2010) Advances in understanding cancer genomes through second-generation sequencing. Nat Rev Genet 11 : 685–696 doi:10.1038/nrg2841
21. van VlierbergheP, PalomeroT, KhiabanianH, van der MeulenJ, CastilloM, et al. (2010) PHF6 mutations in T-cell acute lymphoblastic leukemia. Nat Genet 42 : 338–342 doi:10.1038/ng.542
22. LiH, DurbinR (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25 : 1754–1760 doi:10.1093/bioinformatics/btp324
23. DePristoMA, BanksE, PoplinR, GarimellaKV, MaguireJR, et al. (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43 : 491–8 doi:10.1038/ng.806
24. LarsonDE, HarrisCC, ChenK, KoboldtDC, AbbottTE, et al. (2011) SomaticSniper: Identification of Somatic Point Mutations in Whole Genome Sequencing Data. Bioinformatics 28 : 311–7 doi:10.1093/bioinformatics/btr665
25. McLarenW, PritchardB, RiosD, ChenY, FlicekP, et al. (2010) Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26 : 2069–2070 doi:10.1093/bioinformatics/btq330
26. KimD, PerteaG, TrapnellC, PimentelH, KelleyR, et al. (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14: R36 doi:10.1186/gb-2013-14-4-r36
27. LiH, HandsakerB, WysokerA, FennellT, RuanJ, et al. (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25 : 2078–2079 doi:10.1093/bioinformatics/btp352
28. BassB, HundleyH, LiJB, PengZ, PickrellJ, et al. (2012) The difficult calls in RNA editing. Interviewed by H Craig Mak. Nature Biotechnology 30 : 1207–1209 doi:10.1038/nbt.2452
29. GarberM, GrabherrMG, GuttmanM, TrapnellC (2011) Computational methods for transcriptome annotation and quantification using RNA-seq. Nature Methods 8 : 469–477 doi:10.1038/nmeth.1613
30. ShahSP, RothA, GoyaR, OloumiA, HaG, et al. (2012) The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486 : 395–399 doi:10.1038/nature10933
31. MorinRD, Mendez-LagoM, MungallAJ, GoyaR, MungallKL, et al. (2011) Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476 : 298–303 doi:10.1038/nature10351
32. KentWJ (2002) BLAT—The BLAST-Like Alignment Tool. Genome Res 12 : 656–664 doi:10.1101/gr.229202
33. SherryST, WardMH, KholodovM, BakerJ, PhanL, et al. (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29 : 308–311.
34. Genomes Project Consortium (2012) AbecasisGR, AutonA, BrooksLD, DePristoMA, et al. (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491 : 56–65 doi:10.1038/nature11632
35. DrmanacR, SparksAB, CallowMJ, HalpernAL, BurnsNL, et al. (2010) Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327 : 78–81 doi:10.1126/science.1181498
36. ForbesSA, BhamraG, BamfordS, DawsonE, KokC, et al. (2008) The Catalogue of Somatic Mutations in Cancer (COSMIC). Curr Protoc Hum Genet Chapter 10: Unit10.11 doi:10.1002/0471142905.hg1011s57
37. VogelsteinB, PapadopoulosN, VelculescuVE, ZhouS, DiazLA, et al. (2013) Cancer genome landscapes. Science 339 : 1546–1558 doi:10.1126/science.1235122
38. SturmD, WittH, HovestadtV, Khuong-QuangD-A, JonesDTW, et al. (2012) Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22 : 425–437 doi:10.1016/j.ccr.2012.08.024
39. AertsS, LambrechtsD, MaityS, Van LooP, CoessensB, et al. (2006) Gene prioritization through genomic data fusion. Nature Biotechnology 24 : 537–544 doi:10.1038/nbt1203
40. BashRO, HallS, TimmonsCF, CristWM, AmylonM, et al. (1995) Does activation of the TAL1 gene occur in a majority of patients with T-cell acute lymphoblastic leukemia? A pediatric oncology group study. Blood 86 : 666–676.
41. SoulierJ, ClappierE, CayuelaJ-M, RegnaultA, García-PeydróM, et al. (2005) HOXA genes are included in genetic and biologic networks defining human acute T-cell leukemia (T-ALL). Blood 106 : 274–286 doi:10.1182/blood-2004-10-3900
42. MiyazakiT, TakaokaA, NogueiraL, DikicI, FujiiH, et al. (1998) Pyk2 is a downstream mediator of the IL-2 receptor-coupled Jak signaling pathway. Genes Dev 12 : 770–775.
43. GardinaPJ, ClarkTA, ShimadaB, StaplesMK, YangQ, et al. (2006) Alternative splicing and differential gene expression in colon cancer detected by a whole genome exon array. BMC Genomics 7 : 325 doi:10.1186/1471-2164-7-325
44. ThorsenK, SørensenKD, Brems-EskildsenAS, ModinC, GaustadnesM, et al. (2008) Alternative splicing in colon, bladder, and prostate cancer identified by exon array analysis. Mol Cell Proteomics 7 : 1214–1224 doi:10.1074/mcp.M700590-MCP200
45. GutteryDS, ShawJA, LloydK, PringleJH, WalkerRA (2010) Expression of tenascin-C and its isoforms in the breast. Cancer Metastasis Rev 29 : 595–606 doi:10.1007/s10555-010-9249-9
46. FutrealPA, CoinL, MarshallM, DownT, HubbardT, et al. (2004) A census of human cancer genes. Nat Rev Cancer 4 : 177–183 doi:10.1038/nrc1299
47. CauwelierB, DastugueN, CoolsJ, PoppeB, HerensC, et al. (2006) Molecular cytogenetic study of 126 unselected T-ALL cases reveals high incidence of TCRbeta locus rearrangements and putative new T-cell oncogenes. Leukemia 20 : 1238–1244 doi:10.1038/sj.leu.2404243
48. OramSH, ThomsJ, SiveJI, Calero-NietoFJ, KinstonSJ, et al. (2013) Bivalent promoter marks and a latent enhancer may prime the leukaemia oncogene LMO1 for ectopic expression in T-cell leukaemia. Leukemia 27 : 1348–57 doi:10.1038/leu.2013.2
49. McPhersonA, HormozdiariF, ZayedA, GiulianyR, HaG, et al. (2011) deFuse: an algorithm for gene fusion discovery in tumor RNA-Seq data. PLoS Comput Biol 7: e1001138 doi:10.1371/journal.pcbi.1001138
50. NacuS, YuanW, KanZ, BhattD, RiversCS, et al. (2011) Deep RNA sequencing analysis of readthrough gene fusions in human prostate adenocarcinoma and reference samples. BMC Med Genomics 4 : 11 doi:10.1186/1755-8794-4-11
51. ZhouJ, LiaoJ, ZhengX, ShenH (2012) Chimeric RNAs as potential biomarkers for tumor diagnosis. BMB Rep 45 : 133–140.
52. HuangDW, ShermanBT, LempickiRA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37 : 1–13 doi:10.1093/nar/gkn923
53. HuangDW, ShermanBT, LempickiRA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4 : 44–57 doi:10.1038/nprot.2008.211
54. BrandimarteL, PieriniV, Di GiacomoD, BorgaC, NozzaF, et al. (2013) New MLLT10 gene recombinations in pediatric T-acute lymphoblastic leukemia. Blood 121 : 5064–7 doi:10.1182/blood-2013-02-487256
55. WarmuthM, KimS, GuX-J, XiaG, AdriánF (2007) Ba/F3 cells and their use in kinase drug discovery. Curr Opin Oncol 19 : 55–60 doi:10.1097/CCO.0b013e328011a25f
56. O'NeilJ, TchindaJ, GutierrezA, MoreauL, MaserRS, et al. (2007) Alu elements mediate MYB gene tandem duplication in human T-ALL. Journal of Experimental Medicine 204 : 3059–3066 doi:10.1084/jem.20071637
57. VainchenkerW, ConstantinescuSN (2013) JAK/STAT signaling in hematological malignancies. Oncogene 32 : 2601–2613 doi:10.1038/onc.2012.347
58. SeoJ-S, JuYS, LeeW-C, ShinJ-Y, LeeJK, et al. (2012) The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res 22 : 2109–2119 doi:10.1101/gr.145144.112
59. BergerMF, LevinJZ, VijayendranK, SivachenkoA, AdiconisX, et al. (2010) Integrative analysis of the melanoma transcriptome. Genome 20 : 413–27.
60. GoyaR, SunMGF, MorinRD, LeungG, HaG, et al. (2010) SNVMix: predicting single nucleotide variants from next-generation sequencing of tumors. Bioinformatics 26 : 730–736 doi:10.1093/bioinformatics/btq040
61. QuinnEM, CormicanP, KennyEM, HillM, AnneyR, et al. (2013) Development of strategies for SNP detection in RNA-seq data: application to lymphoblastoid cell lines and evaluation using 1000 Genomes data. PLoS ONE 8: e58815 doi:10.1371/journal.pone.0058815
62. LiuJ, LeeW, JiangZ, ChenZ, JhunjhunwalaS, et al. (2012) Genome and transcriptome sequencing of lung cancers reveal diverse mutational and splicing events. Genome Res 22 : 2315–2327 doi:10.1101/gr.140988.112
63. BettegowdaC, AgrawalN, JiaoY, SausenM, WoodLD (2011) Mutations in CIC and FUBP1 contribute to human oligodendroglioma. Science 333 : 1453–1455.
64. SchwartzentruberJ, KorshunovA, LiuX-Y, JonesDTW, PfaffE, et al. (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482 : 226–231 doi:10.1038/nature10833
65. WuG, BroniscerA, McEachronTA, LuC, PaughBS, et al. (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44 : 251–253 doi:10.1038/ng.1102
66. SunCK, ManK, NgKT, HoJW, LimZX, et al. (2008) Proline-rich tyrosine kinase 2 (Pyk2) promotes proliferation and invasiveness of hepatocellular carcinoma cells through c-Src/ERK activation. Carcinogenesis 29 : 2096–2105 doi:10.1093/carcin/bgn203
67. SunCK, NgKT, LimZX, ChengQ, LoC-M, et al. (2011) Proline-rich tyrosine kinase 2 (Pyk2) promotes cell motility of hepatocellular carcinoma through induction of epithelial to mesenchymal transition. PLoS ONE 6: e18878 doi:10.1371/journal.pone.0018878
68. RajalaHLM, EldforsS, KuusanmäkiH, van AdrichemAJ, OlsonT, et al. (2013) Discovery of somatic STAT5b mutations in large granular lymphocytic leukemia. Blood 121 : 4541–50 doi:10.1182/blood-2012-12-474577
69. NollJE, JefferyJ, Al-EjehF, KumarR, KhannaKK, et al. (2012) Mutant p53 drives multinucleation and invasion through a process that is suppressed by ANKRD11. Oncogene 31 : 2836–2848 doi:10.1038/onc.2011.456
70. MamtaniM, KulkarniH (2012) Association of HADHA expression with the risk of breast cancer: targeted subset analysis and meta-analysis of microarray data. BMC Res Notes 5 : 25 doi:10.1186/1756-0500-5-25
71. KuehHY, RothenbergEV (2012) Regulatory gene network circuits underlying T cell development from multipotent progenitors. Wiley Interdiscip Rev Syst Biol Med 4 : 79–102 doi:10.1002/wsbm.162
72. UribesalgoI, BenitahSA, Di CroceL (2012) From oncogene to tumor suppressor: the dual role of Myc in leukemia. Cell Cycle 11 : 1757–64.
73. RobinsonMD, McCarthyDJ, SmythGK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26 : 139–140 doi:10.1093/bioinformatics/btp616
74. CarroMS, LimWK, AlvarezMJ, BolloRJ, ZhaoX, et al. (2010) The transcriptional network for mesenchymal transformation of brain tumours. Nature 463 : 318–325 doi:10.1038/nature08712
75. MaherCA, Kumar-SinhaC, CaoX, Kalyana-SundaramS, HanB, et al. (2009) Transcriptome sequencing to detect gene fusions in cancer. Nature 458 : 97–101 doi:doi:10.1038/nature07638
76. HalvardsonJ, ZaghloolA, FeukL (2013) Exome RNA sequencing reveals rare and novel alternative transcripts. Nucleic Acids Res 41: e6 doi:10.1093/nar/gks816
77. BeneMC, CastoldiG, KnappW, LudwigWD, MatutesE, et al. (1995) Proposals for the immunological classification of acute leukemias. European Group for the Immunological Characterization of Leukemias (EGIL) Vol. 9 : 1783–1786.
78. SteidlC, ShahSP, WoolcockBW, RuiL, KawaharaM, et al. (2011) MHC class II transactivator CIITA is a recurrent gene fusion partner in lymphoid cancers. Nature 471 : 377–381 doi:10.1038/nature09754
79. RissoD, SchwartzK, SherlockG, DudoitS (2011) GC-content normalization for RNA-Seq data. BMC Bioinformatics 12 : 480 doi:10.1186/1471-2105-12-480
80. GillisJ, PavlidisP (2013) Assessing identity, redundancy and confounds in Gene Ontology annotations over time. Bioinformatics 29 : 476–482 doi:10.1093/bioinformatics/bts727
81. TrapnellC, WilliamsBA, PerteaG, MortazaviA, KwanG, et al. (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature Biotechnology 28 : 511–515 doi:10.1038/nbt.1621
82. RobertsA, PimentelH, TrapnellC, PachterL (2011) Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27 : 2325–2329 doi:10.1093/bioinformatics/btr355
83. SubramanianA, TamayoP, MoothaVK, MukherjeeS, EbertBL, et al. (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102 : 15545–15550 doi:10.1073/pnas.0506580102
84. RobinsonJT, ThorvaldsdóttirH, WincklerW, GuttmanM, LanderES, et al. (2011) Integrative genomics viewer. Nature Biotechnology 29 : 24–26 doi:10.1038/nbt.1754
85. ArtimoP, JonnalageddaM, ArnoldK, BaratinD, CsardiG, et al. (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40: W597–W603 doi:10.1093/nar/gks400
86. SchultzJ, MilpetzF, BorkP, PontingCP (1998) SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci USA 95 : 5857–5864.
87. LetunicI, DoerksT, BorkP (2012) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res 40: D302–D305 doi:10.1093/nar/gkr931
88. de KeersmaeckerK, GrauxC, OderoMD, MentensN, SomersR, et al. (2005) Fusion of EML1 to ABL1 in T-cell acute lymphoblastic leukemia with cryptic t(9;14)(q34;q32). Blood 105 : 4849–4852 doi:10.1182/blood-2004-12-4897
Štítky
Genetika Reprodukčná medicína
Článek Interaction between and during Mammalian Jaw Patterning and in the Pathogenesis of SyngnathiaČlánek Clustering of Tissue-Specific Sub-TADs Accompanies the Regulation of Genes in Developing LimbsČlánek Transcription Factor Occupancy Can Mediate Active Turnover of DNA Methylation at Regulatory RegionsČlánek Tay Bridge Is a Negative Regulator of EGFR Signalling and Interacts with Erk and Mkp3 in the Wing
Článok vyšiel v časopisePLOS Genetics
Najčítanejšie tento týždeň
2013 Číslo 12- Gynekologové a odborníci na reprodukční medicínu se sejdou na prvním virtuálním summitu
- Je „freeze-all“ pro všechny? Odborníci na fertilitu diskutovali na virtuálním summitu
-
Všetky články tohto čísla
- Stressing the Importance of CHOP in Liver Cancer
- The AmAZI1ng Roles of Centriolar Satellites during Development
- Flies Get a Head Start on Meiosis
- Recommendations from Jane Gitschier's Bookshelf
- And Baby Makes Three: Genomic Imprinting in Plant Embryos
- Bugs in Transition: The Dynamic World of in Insects
- Defining the Role of ATP Hydrolysis in Mitotic Segregation of Bacterial Plasmids
- Synaptonemal Complex Components Promote Centromere Pairing in Pre-meiotic Germ Cells
- Cohesinopathies of a Feather Flock Together
- Genetic Recombination Is Targeted towards Gene Promoter Regions in Dogs
- Parathyroid-Specific Deletion of Unravels a Novel Calcineurin-Dependent FGF23 Signaling Pathway That Regulates PTH Secretion
- MAN1B1 Deficiency: An Unexpected CDG-II
- Phosphate Flow between Hybrid Histidine Kinases CheA and CheS Controls Cyst Formation
- Basolateral Mg Extrusion via CNNM4 Mediates Transcellular Mg Transport across Epithelia: A Mouse Model
- Truncation of Unsilences Paternal and Ameliorates Behavioral Defects in the Angelman Syndrome Mouse Model
- Autozygome Sequencing Expands the Horizon of Human Knockout Research and Provides Novel Insights into Human Phenotypic Variation
- Huntington's Disease Induced Cardiac Amyloidosis Is Reversed by Modulating Protein Folding and Oxidative Stress Pathways in the Heart
- Low Frequency Variants, Collapsed Based on Biological Knowledge, Uncover Complexity of Population Stratification in 1000 Genomes Project Data
- Targeted Ablation of and in Retinal Progenitor Cells Mimics Leber Congenital Amaurosis
- Genomic Imprinting in the Embryo Is Partly Regulated by PRC2
- Binary Cell Fate Decisions and Fate Transformation in the Larval Eye
- The Stress-Regulated Transcription Factor CHOP Promotes Hepatic Inflammatory Gene Expression, Fibrosis, and Oncogenesis
- A Global RNAi Screen Identifies a Key Role of Ceramide Phosphoethanolamine for Glial Ensheathment of Axons
- Functional Analysis of the Interdependence between DNA Uptake Sequence and Its Cognate ComP Receptor during Natural Transformation in Species
- Cross-Modulation of Homeostatic Responses to Temperature, Oxygen and Carbon Dioxide in
- Alcohol-Induced Histone Acetylation Reveals a Gene Network Involved in Alcohol Tolerance
- Molecular Characterization of Host-Specific Biofilm Formation in a Vertebrate Gut Symbiont
- CRIS—A Novel cAMP-Binding Protein Controlling Spermiogenesis and the Development of Flagellar Bending
- Dual Regulation of the Mitotic Exit Network (MEN) by PP2A-Cdc55 Phosphatase
- Expanding the Marine Virosphere Using Metagenomics
- Detection of Slipped-DNAs at the Trinucleotide Repeats of the Myotonic Dystrophy Type I Disease Locus in Patient Tissues
- Interaction between and during Mammalian Jaw Patterning and in the Pathogenesis of Syngnathia
- Mutations in the UQCC1-Interacting Protein, UQCC2, Cause Human Complex III Deficiency Associated with Perturbed Cytochrome Protein Expression
- Reactivation of Chromosomally Integrated Human Herpesvirus-6 by Telomeric Circle Formation
- Anoxia-Reoxygenation Regulates Mitochondrial Dynamics through the Hypoxia Response Pathway, SKN-1/Nrf, and Stomatin-Like Protein STL-1/SLP-2
- The Midline Protein Regulates Axon Guidance by Blocking the Reiteration of Neuroblast Rows within the Drosophila Ventral Nerve Cord
- Tomato Yield Heterosis Is Triggered by a Dosage Sensitivity of the Florigen Pathway That Fine-Tunes Shoot Architecture
- Selection on Plant Male Function Genes Identifies Candidates for Reproductive Isolation of Yellow Monkeyflowers
- Role of Tomato Lipoxygenase D in Wound-Induced Jasmonate Biosynthesis and Plant Immunity to Insect Herbivores
- Meiotic Cohesin SMC1β Provides Prophase I Centromeric Cohesion and Is Required for Multiple Synapsis-Associated Functions
- Identification of Sphingolipid Metabolites That Induce Obesity via Misregulation of Appetite, Caloric Intake and Fat Storage in
- Genome-Wide Screen Reveals Replication Pathway for Quasi-Palindrome Fragility Dependent on Homologous Recombination
- Histone Methylation Restrains the Expression of Subtype-Specific Genes during Terminal Neuronal Differentiation in
- A Novel Intergenic ETnII-β Insertion Mutation Causes Multiple Malformations in Mice
- The NuRD Chromatin-Remodeling Enzyme CHD4 Promotes Embryonic Vascular Integrity by Transcriptionally Regulating Extracellular Matrix Proteolysis
- A Domesticated Transposase Interacts with Heterochromatin and Catalyzes Reproducible DNA Elimination in
- Acute Versus Chronic Loss of Mammalian Results in Distinct Ciliary Phenotypes
- MBD3 Localizes at Promoters, Gene Bodies and Enhancers of Active Genes
- Positive and Negative Regulation of Gli Activity by Kif7 in the Zebrafish Embryo
- A Hereditary Spastic Paraplegia Mouse Model Supports a Role of ZFYVE26/SPASTIZIN for the Endolysosomal System
- The CCR4-NOT Complex Mediates Deadenylation and Degradation of Stem Cell mRNAs and Promotes Planarian Stem Cell Differentiation
- Reconstructing Native American Migrations from Whole-Genome and Whole-Exome Data
- Contributions of Protein-Coding and Regulatory Change to Adaptive Molecular Evolution in Murid Rodents
- Comprehensive Analysis of Transcriptome Variation Uncovers Known and Novel Driver Events in T-Cell Acute Lymphoblastic Leukemia
- A -Acting Protein Effect Causes Severe Eye Malformation in the Mouse
- Clustering of Tissue-Specific Sub-TADs Accompanies the Regulation of Genes in Developing Limbs
- Germline Progenitors Escape the Widespread Phenomenon of Homolog Pairing during Development
- Transcription Factor Occupancy Can Mediate Active Turnover of DNA Methylation at Regulatory Regions
- Somatic mtDNA Mutation Spectra in the Aging Human Putamen
- ESCRT-I Mediates FLS2 Endosomal Sorting and Plant Immunity
- Ethylene Promotes Hypocotyl Growth and HY5 Degradation by Enhancing the Movement of COP1 to the Nucleus in the Light
- The PAF Complex and Prf1/Rtf1 Delineate Distinct Cdk9-Dependent Pathways Regulating Transcription Elongation in Fission Yeast
- Dual Regulation of Gene Expression Mediated by Extended MAPK Activation and Salicylic Acid Contributes to Robust Innate Immunity in
- Quantifying Missing Heritability at Known GWAS Loci
- Smc5/6-Mms21 Prevents and Eliminates Inappropriate Recombination Intermediates in Meiosis
- Smc5/6 Coordinates Formation and Resolution of Joint Molecules with Chromosome Morphology to Ensure Meiotic Divisions
- Tay Bridge Is a Negative Regulator of EGFR Signalling and Interacts with Erk and Mkp3 in the Wing
- Meiotic Crossover Control by Concerted Action of Rad51-Dmc1 in Homolog Template Bias and Robust Homeostatic Regulation
- Active Transport and Diffusion Barriers Restrict Joubert Syndrome-Associated ARL13B/ARL-13 to an Inv-like Ciliary Membrane Subdomain
- An Regulatory Circuit Modulates /Wnt Signaling and Determines the Size of the Midbrain Dopaminergic Progenitor Pool
- Variants Induce Differential Protection to Viruses in : A Phenotypic and Phylogenomic Analysis
- Base Pairing Interaction between 5′- and 3′-UTRs Controls mRNA Translation in
- Evidence That Masking of Synapsis Imperfections Counterbalances Quality Control to Promote Efficient Meiosis
- Insulin/IGF-Regulated Size Scaling of Neuroendocrine Cells Expressing the bHLH Transcription Factor in
- Sumoylated NHR-25/NR5A Regulates Cell Fate during Vulval Development
- TATN-1 Mutations Reveal a Novel Role for Tyrosine as a Metabolic Signal That Influences Developmental Decisions and Longevity in
- PLOS Genetics
- Archív čísel
- Aktuálne číslo
- Informácie o časopise
Najčítanejšie v tomto čísle- The NuRD Chromatin-Remodeling Enzyme CHD4 Promotes Embryonic Vascular Integrity by Transcriptionally Regulating Extracellular Matrix Proteolysis
- Mutations in the UQCC1-Interacting Protein, UQCC2, Cause Human Complex III Deficiency Associated with Perturbed Cytochrome Protein Expression
- The Midline Protein Regulates Axon Guidance by Blocking the Reiteration of Neuroblast Rows within the Drosophila Ventral Nerve Cord
- Tomato Yield Heterosis Is Triggered by a Dosage Sensitivity of the Florigen Pathway That Fine-Tunes Shoot Architecture
Prihlásenie#ADS_BOTTOM_SCRIPTS#Zabudnuté hesloZadajte e-mailovú adresu, s ktorou ste vytvárali účet. Budú Vám na ňu zasielané informácie k nastaveniu nového hesla.
- Časopisy



