-
Články
- Časopisy
- Kurzy
- Témy
- Kongresy
- Videa
- Podcasty
and Control Optic Cup Regeneration in a Prototypic Eye
Optic cups are a structural feature of diverse eyes, from simple pit eyes to camera eyes of vertebrates and cephalopods. We used the planarian prototypic eye as a model to study the genetic control of optic cup formation and regeneration. We identified two genes encoding transcription factors, sp6-9 and dlx, that were expressed in the eye specifically in the optic cup and not the photoreceptor neurons. RNAi of these genes prevented formation of visible optic cups during regeneration. Planarian regeneration requires an adult proliferative cell population with stem cell-like properties called the neoblasts. We found that optic cup formation occurred only after migration of progressively differentiating progenitor cells from the neoblast population. The eye regeneration defect caused by dlx and sp6-9 RNAi can be explained by a failure to generate these early optic cup progenitors. Dlx and Sp6-9 genes function as a module during the development of diverse animal appendages, including vertebrate and insect limbs. Our work reveals a novel function for this gene pair in the development of a fundamental eye component, and it utilizes these genes to demonstrate a mechanism for total organ regeneration in which extensive cell movement separates new cell specification from organ morphogenesis.
Published in the journal: and Control Optic Cup Regeneration in a Prototypic Eye. PLoS Genet 7(8): e32767. doi:10.1371/journal.pgen.1002226
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002226Summary
Optic cups are a structural feature of diverse eyes, from simple pit eyes to camera eyes of vertebrates and cephalopods. We used the planarian prototypic eye as a model to study the genetic control of optic cup formation and regeneration. We identified two genes encoding transcription factors, sp6-9 and dlx, that were expressed in the eye specifically in the optic cup and not the photoreceptor neurons. RNAi of these genes prevented formation of visible optic cups during regeneration. Planarian regeneration requires an adult proliferative cell population with stem cell-like properties called the neoblasts. We found that optic cup formation occurred only after migration of progressively differentiating progenitor cells from the neoblast population. The eye regeneration defect caused by dlx and sp6-9 RNAi can be explained by a failure to generate these early optic cup progenitors. Dlx and Sp6-9 genes function as a module during the development of diverse animal appendages, including vertebrate and insect limbs. Our work reveals a novel function for this gene pair in the development of a fundamental eye component, and it utilizes these genes to demonstrate a mechanism for total organ regeneration in which extensive cell movement separates new cell specification from organ morphogenesis.
Introduction
Animal retinas are susceptible to damage and degeneration from injury and because of sensitivity to light. Multiple vertebrates have evolved the ability to regenerate ocular tissue following damage or degeneration. In zebrafish, proliferating marginal zone cells, specialized rod progenitors, and Müller glia reside within the retina and are sources of regenerative tissue [1]. In urodele amphibians, cells of the retinal pigment epithelium can act as a source of new retinal neurons in the adult [2]. Some invertebrates, such as planarians, are also capable of eye regeneration. Unlike vertebrates, planarians can regenerate eyes completely de novo, using a population of cells that resides entirely outside of the eye.
The eyes of planarians are substantially simpler than vertebrate camera eyes, but there are important similarities between the two structures nonetheless. Eye formation in planarians, vertebrates, and other animals involves common genes such as sine oculis and eyes absent [3]–[4]. Furthermore, in both vertebrates and planarians, specialized pigment cells are organized such that they directly abut photoreceptive organelles in an optic cup. In vertebrates, cells of the retinal pigment epithelium (RPE) contact the outer segments of photoreceptor neurons from an adjacent layer of the optic cup. In planarians, the optic cup is entirely formed of pigment cells (it is commonly termed the “pigment cup”) and photoreceptor neurons project rhabdomeres into the cup [5] (Figure 1A and Video S1). A primary function of pigmented optic cups in simple eyes is to absorb incoming light prior to detection by photoreceptors [6], as this creates shade that allows the eye and brain to resolve the direction of incoming light. Light absorption is also an important function of the vertebrate RPE [7], although vertebrate eyes use sophisticated image-forming mechanisms for vision with spatial resolution.
Fig. 1. The planarian optic pigment cup expresses dlx and sp6-9. 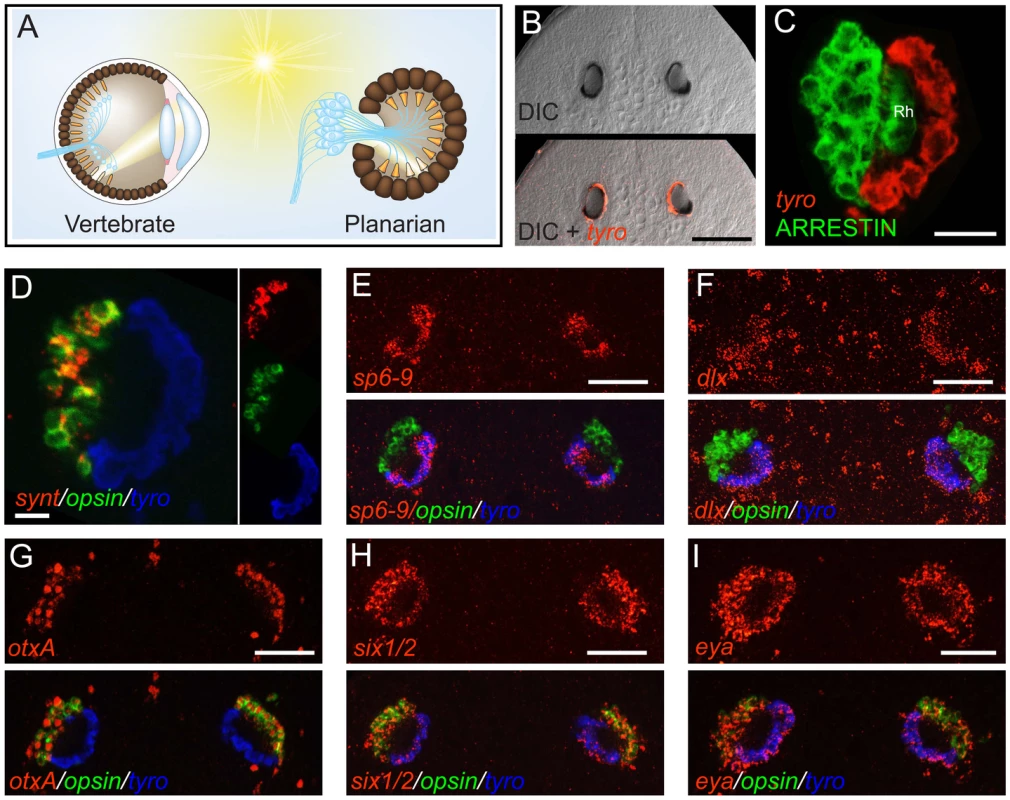
Anterior is up in all images, and all eyes are shown at day 6 of regeneration except in (B). Fluorescent images are fluorescent in situ hybridizations (FISH) unless otherwise noted. (A) Schematic highlighting similarities between the vertebrate (left) and planarian (right) optic cup. Light-sensing organelles (orange), neuronal cell bodies and processes (blue), and pigmented cells of the optic cup (brown) are depicted. Neural circuitry of the vertebrate retina is highly simplified. (B) Smed-tyrosinase (tyro) is expressed in the optic (pigment) cup of an intact planarian. (C) Planarian photoreceptor neurons, labeled by an anti-ARRESTIN antibody [60] are adjacent to the pigment cup and extend rhabdomeres (Rh) into the cup lumen. (D) The optic cup does not express opsin or synaptotagmin (synt). (E–I) Expression of transcription factors in the planarian eye during head regeneration. sp6-9 (E) and dlx (F) are expressed in the optic cup. (G) otxA is expressed in the photoreceptor neurons, similar to the case for D. japonica [20]. six1/2 (H) and eya (I) are expressed in both the optic cup and photoreceptor neurons, similar to the case for other planarian species [3]–[4]. Scale bars, 100 µm (B, D), 50 µm (C, E–I). Planarian eyes can regenerate even after decapitation, from tissue that originally resided far from the head. Planarian regeneration is possible because of a population of proliferative cells, the neoblasts, that includes pluripotent stem cells [8] as well as all cycling somatic cells of the adult. The neoblasts are distributed throughout the body in the parenchyma of the adult [9], and new tissue is produced at wounds by localized increase in neoblast proliferation, followed by cell cycle exit and differentiation [10]. Wounding in areas devoid of neoblasts ultimately results in localization of cycling cells at the wound site, indicating that cell migration can be important for repair of at least some injuries [10]. Furthermore, immediate neoblast descendants are more peripherally located than the neoblasts, suggesting that cell movements occur during differentiation [11]–[12]. In prior studies, neoblasts and their descendants were examined in vivo as large populations of cells with unidentified lineage and fate. Therefore, very little is currently known about the cellular and genetic events that occur between the pluripotent state and the terminally differentiated state during regeneration of specific organs such as the eye.
Here we identify the conserved transcription factors dlx and sp6-9 as novel regulators of planarian eye regeneration. These genes are expressed at early stages of pigment cup progenitor specification and are required for regeneration of the cup. We find that progenitors of pigment cup cells and photoreceptor neurons form distinct mesenchymal populations substantially before differentiation and morphogenesis. Our genetic characterization of the pigment cup allows us to identify lineage-specified pigment cup cells within the neoblast population, at surprisingly large distances from the final position of the regenerating eye, and we demonstrate that sp6-9+/dlx+ eye precursors differentiate in a spatially graded manner through the blastema prior to reaching the eye. Therefore, in contrast to epithelium-based modes of eye development, planarian eye regeneration relies on a dramatic spatial decoupling of progenitor specification and morphogenesis.
Results
The optic (pigment) cup is defined by expression of tyrosinase and the transcription factors sp6-9 and dlx
As in vertebrates, but unlike most protostomes, the planarian optic shading pigment is primarily composed of melanin [13]. Consistent with this, we found that the Schmidtea mediterranea gene Smed-tyrosinase (Figure S1), homologs of which are required for melanin synthesis [14], was expressed exclusively in the pigment cup region of the planarian eye (Figure 1B and 1C and Figure S2). Whereas photosensing neurons in planarians express Smed-opsin [15], pigment cells did not have detectable expression of Smed-opsin, nor the pan-neuronal markers Smed-synapsin and Smed-synaptotagmin (Figure 1D and Figure S3), indicating that pigment cup cells do not function directly in light detection and phototransduction.
Eye development is controlled by similar transcription factors in diverse animals. We sought factors that might control formation of the pigmented optic cup in planarians during regeneration by broadly screening expression patterns of conserved transcription factor-encoding genes. We identified two genes, Smed-sp6-9 and Smed-dlx, that were both expressed in the regenerating eye, specifically in the optic cup of the regenerating eye (Figure 1E and 1F). These genes were also expressed in cells outside of the eye (Figure S2 and S4), including in neurons of the brain and cells of the head rim and pharynx.
Smed-sp6-9 (sp6-9) clustered within the Sp6-9 zinc finger gene family (Figure S1), which diverged from the Sp1-4 and Sp5 families prior to the evolution of the Bilateria [16], suggesting Smed-sp6-9 is equally related to vertebrate Sp6, Sp7, Sp8, and Sp9 genes. Smed-dlx (dlx) is homologous to the Distal-less family of homeobox genes (Figure S1), which have broadly important roles in development [17]. otxA, sine oculis, and eyes absent genes, homologs of which encode transcriptional regulatory proteins required for the development of diverse animal eyes [18]–[19], are expressed in the eyes of planarians [3]–[4], [20]. We cloned S. mediterranea orthologs of these genes. In contrast to sp6-9 and dlx, Smed-otxA (otxA) was expressed specifically in the photoreceptor neurons of the eye, but not the pigment cup (Figure 1G and Figure S2) [20]. Smed-eyes absent (eya) and Smed-sine oculis1/2 (six1/2) were expressed in both photoreceptor neurons and pigment cups (Figure 1H, 1I and Figure S2) [3]–[4]. We also identified an ortholog of dachshund (Figure S5A), a gene with important regulatory functions in Drosophila eye development [21], but did not detect Smed-dachshund expression in the regenerating eye (Figure S5B).
Expression of transcription factors was only weakly detected in the pigment cups of intact (non-regenerating) animals (Figure S2). As the pigment of the optic cup could obscure signal in intact eyes, we used RNAi of tyrosinase (see below) to reduce cup pigmentation prior to fixation of animals for FISH. Even with reduced pigmentation, however, expression of dlx, sp6-9, six-1/2 and eya was faintly detected in the intact optic cup (Figure S6). In summary, we have characterized eye-expressed transcription factors in planarians on the basis of expression in photoreceptor neurons and/or the optic cup during regeneration, and have identified two genes that are expressed specifically in the optic cup during regeneration.
dlx and sp6-9 are required for regeneration of pigment cups
Dlx and Sp6-9 genes have essential roles during development of several animal tissues. We therefore examined whether dlx and sp6-9 loss of function could affect formation of optic cups during regeneration. RNAi of dlx or sp6-9 followed by decapitation resulted in planarians that did not regenerate visible optic cups, but did make photoreceptor neurons that contacted the brain (Figure 2A–2C). Abnormal targeting of photoreceptor neuron processes in dlx(RNAi) and sp6-9(RNAi) animals was observed, possibly due to the physical absence of a cup, which typically encloses many processes (Figure 1C). tyrosinase(RNAi) animals had weakly pigmented optic cups, and displayed normal photoreceptor neuron morphology (Figure 2D). In contrast, RNAi of otxA resulted in animals that successfully made optic cups, but lacked photoreceptor neurons (Figure 2E). Consistent with six-1/2 expression in both pigment cups and photoreceptor neurons, RNAi of this gene strongly affected pigment cup regeneration and photoreceptor neuron formation (Figure 2F), as previously observed [3]. We were unable to observe an eye defect following RNAi of dachshund (n = 8/8 animals regenerated pigment cups).
Fig. 2. dlx and sp6-9 are required for optic cup regeneration. 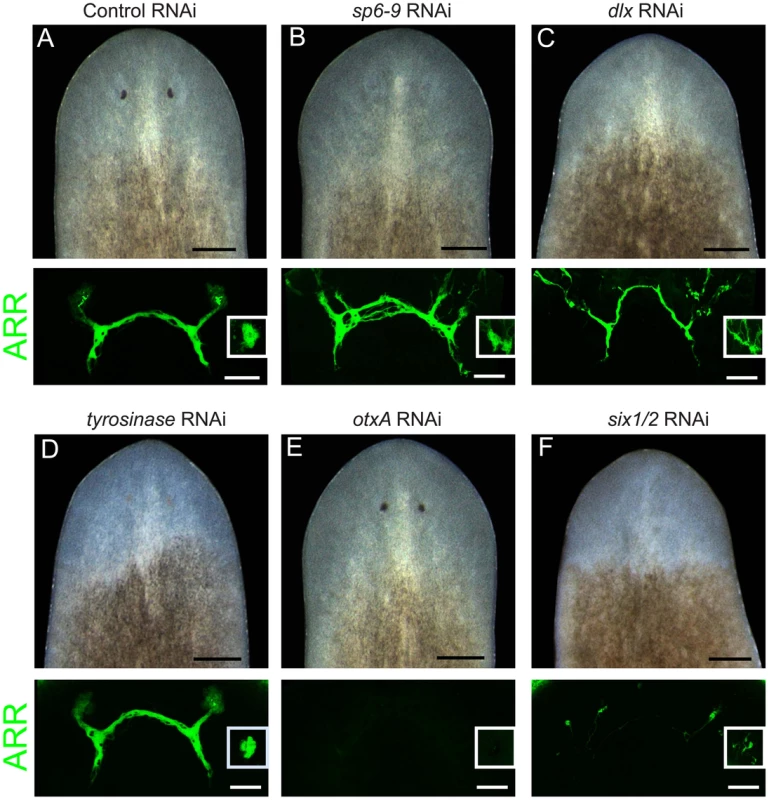
Pigment cups are visualized in live animals on day 7 of regeneration after decapitation. Photoreceptor neurons are visualized with anti-ARRESTIN (ARR) antibody. Insets in ARR panels show a photoreceptor from the dorsal side. Penetrance in (A) is n = 10/10 (presence of pigment cups) and n = 6/6 (photoreceptor neurons that contact brain); in (B) n = 0/10 and n = 9/9; (C) n = 0/10 and 9/9; (D) n = 0/7 and n = 7/7; (E) n = 10/10 and n = 0/8; (F) n = 0/10 and n = 3/11. Scale bars, 100 µm for fluorescent images, 200 µm for live worm images. dlx and sp6-9 were also required for eye regeneration following targeted removal of the eye without decapitation. Control animals regenerated a small pigment cup within 7 days of surgical excision of the eye. However, dlx(RNAi) and sp6-9(RNAi) animals did not regenerate pigment cups following this type of injury (Figure S7A). In order to examine whether dlx and sp6-9 are required for homeostatic maintenance of the eye in uninjured animals, dsRNA was delivered by feeding over 3–7 weeks without injury (Figure S7B). dlx(RNAi) animals developed lesions in the region of the eye (n = 5/9). Lesions also occurred in other areas of the body, including the tail and pre-pharyngeal region, and animals ultimately died from lysis. Homeostatic sp6-9(RNAi) animals exhibited gradual loss in size and pigmentation of the optic cup (n = 20/20), culminating in loss of visible optic cups in some animals (n = 4/20) after 7 weeks of RNAi (Figure S7B). Our expression and functional analyses indicate that dlx and sp6-9 have essential roles in pigment cup maintenance as well as regeneration following diverse injuries to the eye.
A population of cells directly posterior to the optic cup expresses markers of the optic cup
In order to understand the mechanism by which dlx and sp6-9 act to promote eye regeneration, we sought to identify the source of pigment cup cells. smedwi-1 mRNA is a marker for neoblasts, whereas SMEDWI-1 protein is a marker for neoblasts and immediate neoblast descendants, because of the fact that SMEDWI-1 protein perdures beyond expression of smedwi-1 mRNA [10], [22]. In intact animals we were able to identify isolated cells in the dorsal anterior and in the brain that were positive for both SMEDWI-1 protein and sp6-9 mRNA or dlx mRNA (Figure S8), indicating that these genes are expressed in diverse neoblast-descendant progenitor populations during homeostasis. During regeneration following decapitation, we detected a dense trail of SMEDWI-1+/sp6-9+ cells behind the eye primordium (Figure 3A). These cells were present in a band proximal and posterior to the eye, and at the same plane of the eye on the dorso-ventral axis. This is the region imaged and analyzed in all experiments referring to a cell “trail.”
Fig. 3. A population of mesenchymal cells posterior to the eye during regeneration expresses optic cup transcription factors. 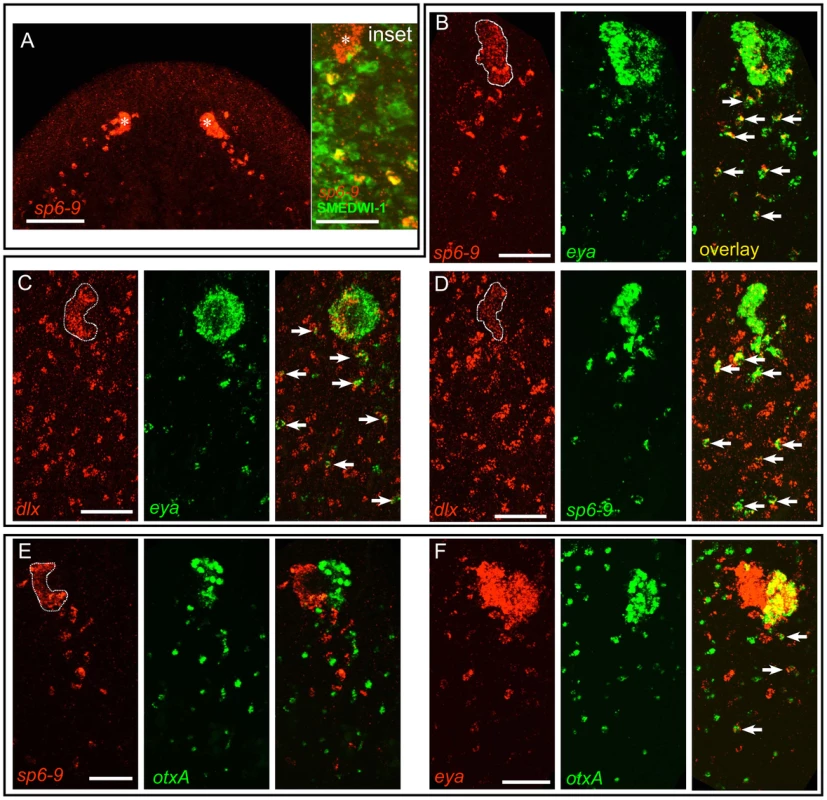
Anterior is up in all images, and all eyes and trails are in blastemas at day 6 of regeneration following decapitation. In all high magnification images of trails only one of two eyes is shown, and images are rotated slightly so that the trail fits into a vertical frame. Fluorescent images are FISH. (A) During regeneration sp6-9 is expressed in the pigment cups (Pcup) and in trails of cells directly behind the pigment cups on the dorsal side of the animal. Inset: sp6-9+ trail cells, but not the eye primordium, are positive for SMEDWI-1 protein. Asterisk indicates eye primordium. (B–D) Double FISH for transcription factors showing that trail cells express combinations of genes also expressed in the cup primordium. sp6-9-expressing (B) and dlx-expressing (C) cells in the Pcup and trail also express eya. (D) sp6-9 and dlx are co-expressed in the Pcup and in many trail cells. (E) sp6-9-expressing cells did not express the photoreceptor neuron marker otxA. (F) Numerous eya-expressing cells did express otxA. Arrows show double positive cells. Pigment cup is outlined in (B–E). Scale bars, 100 µm (A), 50 µm in (B–F). The close proximity of trail cells to the optic cup, the expression of the optic cup gene sp6-9, and the presence of a marker for immediate neoblast progeny cells raised the possibility that these trail cells are progenitors of optic cup cells. To explore this possibility we first sought to determine the extent of molecular similarity between trail cells and optic cup cells. Using combinatorial FISH, we found that sp6-9+ trail cells co-expressed other markers of the optic cup, including eya, six1/2, and dlx (Figure 3B–3D and Figure S9A–S9B). The identified trail cells therefore possess an expression profile that is uniquely similar to the optic cup; outside of the eye, no other differentiated cell types, including the neighboring sp6-9+ head rim cells, were found to be positive for this combination of markers (Figure S10). To test whether sp6-9+ trail cells shared molecular similarity with photoreceptor neurons as well as the optic cup cells, we looked for overlap between expression of sp6-9 and otxA, which is specifically expressed in the photoreceptor neurons in the eye. Cells double-positive for these markers or for dlx/otxA were observed only very rarely (Figure 3E and Figure S9C), indicating that sp6-9+ trail cells share molecular identity with the pigment cup, but not with photoreceptor neurons. Numerous otxA+/eya+ cells were, however, detected in mesenchymal cells posterior to the eye (Figure 3F), consistent with the fact that eya is also expressed in photoreceptor neurons. In summary, we identified a population of sp6-9+ cells posterior to the eye during head regeneration that shares a highly similar expression profile with optic cup cells (sp6-9+/dlx+/eya+/six1-2+/otxA−), but not with any other cell type of the animal. Furthermore, we identified an adjacent population of mesenchymal blastema cells that share a similar expression profile with the photoreceptor neurons of the eye (sp6-9−/dlx−/otxA+/eya+).
We hypothesized that the mesenchymal trails of sp6-9+/eya+/dlx+/six1-2+/otxA− cells are progenitor cells that are the source of new pigment cup cells during eye regeneration after decapitation. Some sp6-9+/eya+ cells in the trail expressed tyrosinase (Figure 4A). tyrosinase expression serves as a marker for the differentiation of optic cup cells, and its expression in the trail indicates that these cells are indeed in the process of differentiation. sp6-9+/eya+/tyrosinase+ cells of the trail were typically closer to the optic cup (Figure 4A and 4B) than were sp6-9+/eya+/tyrosinase− cells. Using single rather than triple probe in situ hybridization conditions, some tyrosinase+ cells could be detected 150 µm or more away from the pigment cup (Figure 4C). Expression of tyrosinase in these distant cells was consistently much weaker, further indicating that proximity to the cup primordium correlates with extent of differentiation. tyrosinase expression in intact animals was not detected in any cells of the body outside of the optic cup, confirming that the expression profile of the sp6-9+ trail cells identified here is uniquely similar to that of optic cup cells. We refer to these cells as “optic cup trail cells,” and use detection of sp6-9 and eya co-expression to identify these cells in subsequent experiments.
Fig. 4. Optic cup trail cells exhibit a gradient of differentiation that correlates with distance from the cup primordium. 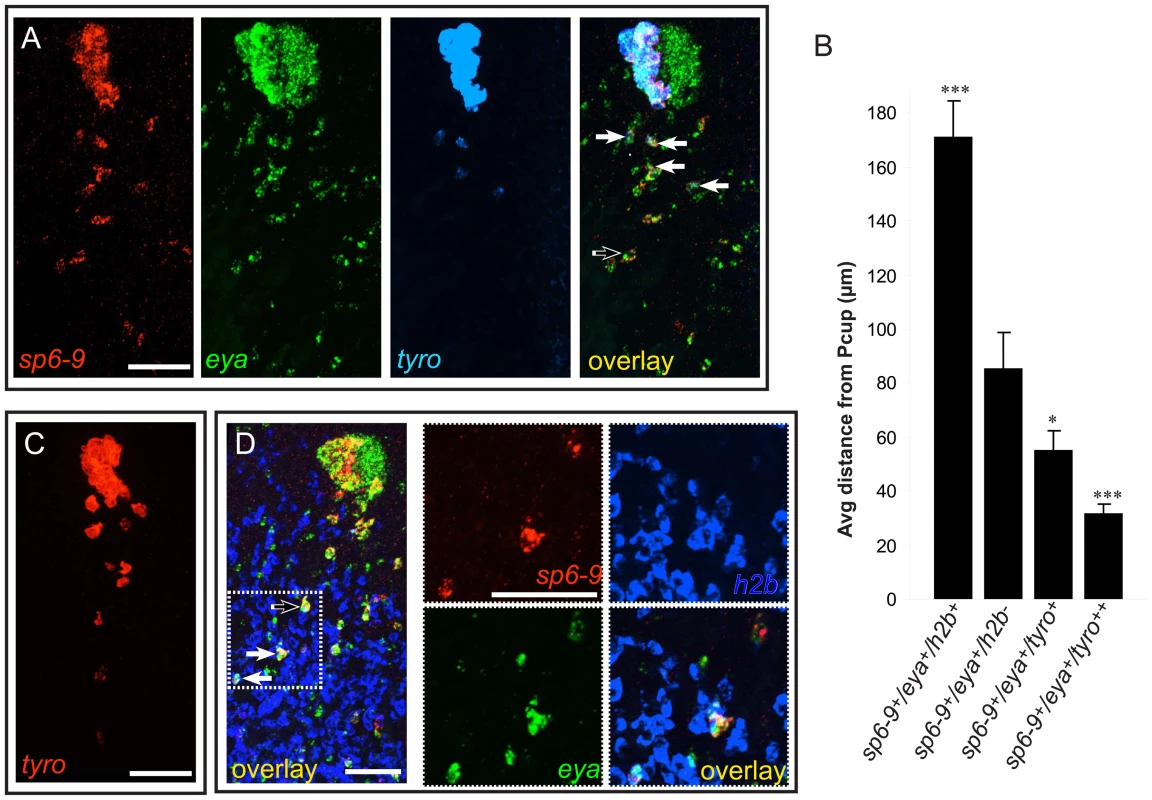
Anterior is up in all images, and all eyes and trails are in blastemas at day 6 of regeneration following decapitation. (A) sp6-9+/eya+ trail cells strongly expressing tyrosinase are close to the optic cup. (B) Quantification of distance of trail cells from the Pcup (mean ± s.e.m; n = 4 eyes for each category; significance by two-tailed t-test is shown relative to second bar (sp6-9+/eya+) ***P<.001, *P<.05). tyro++ indicates strong, non-granular signal. (C) Weak tyrosinase expression can be detected in cells far from the optic cup. (D) Some sp6-9+/eya+ cells express the proliferation marker histone h2b. Solid arrows show triple-positive cells and the open arrow shows double-positive cells. Scale bars, 50 µm. Optic cup trail cells originate in the neoblast population
Differentiated cells in planarians are non-mitotic, and neoblasts are the only cycling somatic cells of the animal [9]. Surprisingly, at positions relatively far from the pigment cup, sp6-9+/eya+ trail cells were observed that also expressed the proliferative marker [23] histone h2b (Figure 4D) and the neoblast marker [24] smedwi-1 mRNA (Figure S11). RT-PCR analysis of FACS sorted “X1” neoblasts has indicated that the neoblast population contains heterogeneity of gene expression and possible commitment to specific lineages [25]. smedwi-1 and h2b positive optic cup trail cells, described here, are the first lineage-committed neoblast subpopulation observed in vivo. It will be interesting to investigate whether these sp6-9+/eya+ cycling cells function as stem cells, or whether they are a transient, differentiating cell type. Neither h2b nor smedwi-1 expression was observed within the optic cup at any point during regeneration. Indeed, sp6-9+/eya+/h2b+ cells were typically present in a region of the trail that was relatively distant from the regenerating eye primordium, in contrast to the sp6-9+/eya+/tyrosinase+ cells that were closer to the eye (Figure 4B and 4D). These data indicate that there exists a distal-to-proximal distribution of progenitor trail cells with respect to the eye that begin in the dividing neoblast population (distal), exit the cell cycle and begin differentiation (intermediate), and finally differentiate fully and aggregate (proximal).
Optic cup trail cells are a source of new eye tissue during regeneration
To further test the possibility that optic cup trail cells are the source of new optic cup tissue in regeneration, animals were irradiated with 6,000 Rads on the third day of regeneration following decapitation, a point at which aggregates of eye cells were first apparent. Irradiation permanently blocks all new cell division in planarians [9]. Nonetheless, optic cups underwent significant growth, measured in number of cells, following irradiation on day 3 of regeneration (Figure 5A and 5B). Simultaneously, the number of sp6-9+/eya+ trail cells gradually decreased following irradiation. The extinction of trail cells 24–48 hours after irradiation correlated with the end of cup growth. Furthermore, the number of trail cells present shortly following irradiation approximately matched the number of cells gained by the optic cup with time following irradiation. When BrdU was injected at day three of regeneration, incorporation into sp6-9+/eya+ cells was first detected distal to the cup in the trail, at 24 hours after injection (Figure 5C and 5D). Incorporation of BrdU first into distal cells of the trail provides further evidence that these cells have recently undergone S-phase or are actively in S-phase. At later time points following the BrdU pulse, however, a majority of sp6-9+/eya+/BrdU+ cells were located in the cup aggregate (Figure 5C and 5D). Because most cycling sp6-9+/eya+ cells are located at sites distal to the eye, and no cycling cells are found in any part of the eye primordium (Figure 4D), the presence of BrdU signal in the eye three days after the BrdU pulse indicates that a net movement of sp6-9+/eya+/BrdU+ cells from distal to proximal to the eye occurs during regeneration. Together, these data suggest a model in which optic cup progenitors arise after decapitation and migrate toward the eye while undergoing differentiation.
Fig. 5. Optic cup trail cells are a source of new optic cup tissue. 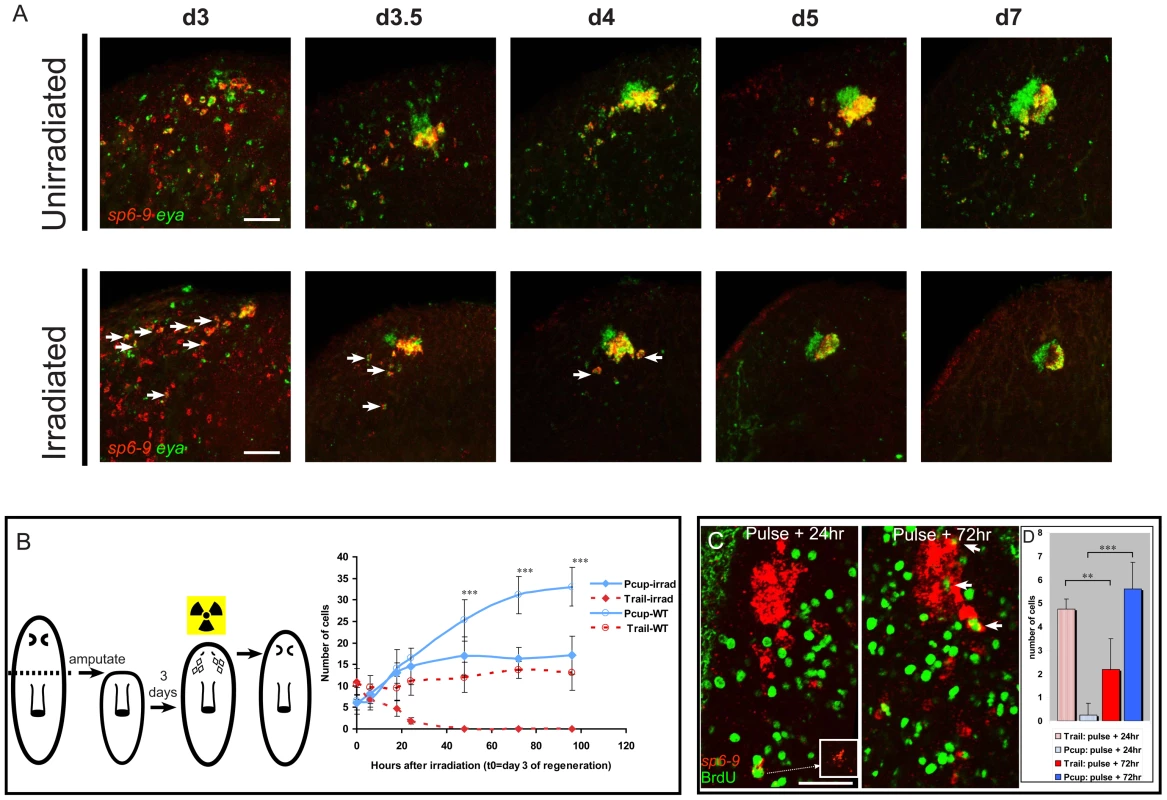
All fluorescent images are FISH, except for fluorescent detection of BrdU by antibody. (A) Time course of eye regeneration in unirradiated animals and animals irradiated at day 3 of regeneration following decapitation. In irradiated panels arrows indicate double positive cells. (B) Experimental design diagram and quantification of cell numbers in pigment cups (Pcups) and trails of irradiated and unirradiated animals over time (mean ± s.d; n≥9 eyes/trails; significance by two-tailed t-test is shown for the Pcup cell numbers in irradiated vs. un-irradiated ***P<.05). (C) BrdU injection at day three of regeneration after decapitation only labels trail cells after one day, but labels Pcup cells after a longer time delay. Inset: magnification of sp6-9 signal in BrdU/sp6-9-positive cell. Arrowheads indicate sp6-9+/eya+/BrdU+ Pcup cells. (D) Quantification of BrdU+/sp6-9+/eya+ cell numbers in the trail and Pcup 24 h and 72 h after BrdU injection (mean ± s.d; n = 5 eyes; significance by two-tailed t-test ***P<.001, **P<.005). Scale bars, 50 µm. dlx and sp6-9 are required for eye progenitor formation after decapitation
Because sp6-9 and dlx are expressed in the undifferentiated cells of the optic cup trail, we sought to determine whether they are required for generation of the optic cup progenitors or for later stages of cup formation. RNAi of dlx to eliminate gene function resulted in animals that completely lacked sp6-9+/eya+ optic cup trail cells during regeneration as well as sp6-9+/eya+ double-positive signal in the optic cup primordium (Figure 6A, 6C and 6H; higher magnification images of RNAi phenotypes are in Figure S12). We obtained a similar result for sp6-9(RNAi) animals, which largely lacked dlx+/eya+ progenitor cells (Figure 6B and 6D). eya+ signal remaining in these animals reflects photoreceptor neuron expression of this gene. tyrosinase(RNAi) animals lacked visible eye pigment but displayed normal sp6-9+/eya+ progenitor patterns (Figure 2D, Figure 6E and 6H), and eye markers were normally positioned, indicating that, unlike sp6-9(RNAi) and dlx(RNAi) animals, tyrosinase(RNAi) animals are able to regenerate optic cup tissue. otxA(RNAi) animals also had normal and abundant sp6-9+/eya+ progenitors, as expected given that these animals regenerate visible pigment cups (Figure 6F and 6H). RNAi of six1/2 led to severe reduction in numbers of both pigment cup progenitors and photoreceptor neurons following regeneration (Figure 6G and 6H). We verified the presence or absence of pigment cups in the above RNAi conditions using DIC light microscopy (Figure S13). Pigment cups (epithelial cells arranged in a crescent around a lumen) were observed in control, tyrosinase(RNAi), and otxA(RNAi) animals; no such cellular formations were observed in sp6-9(RNAi), dlx(RNAi), or six1/2(RNAi) animals.
Fig. 6. dlx and sp6-9 are required for regenerative optic cup cells. 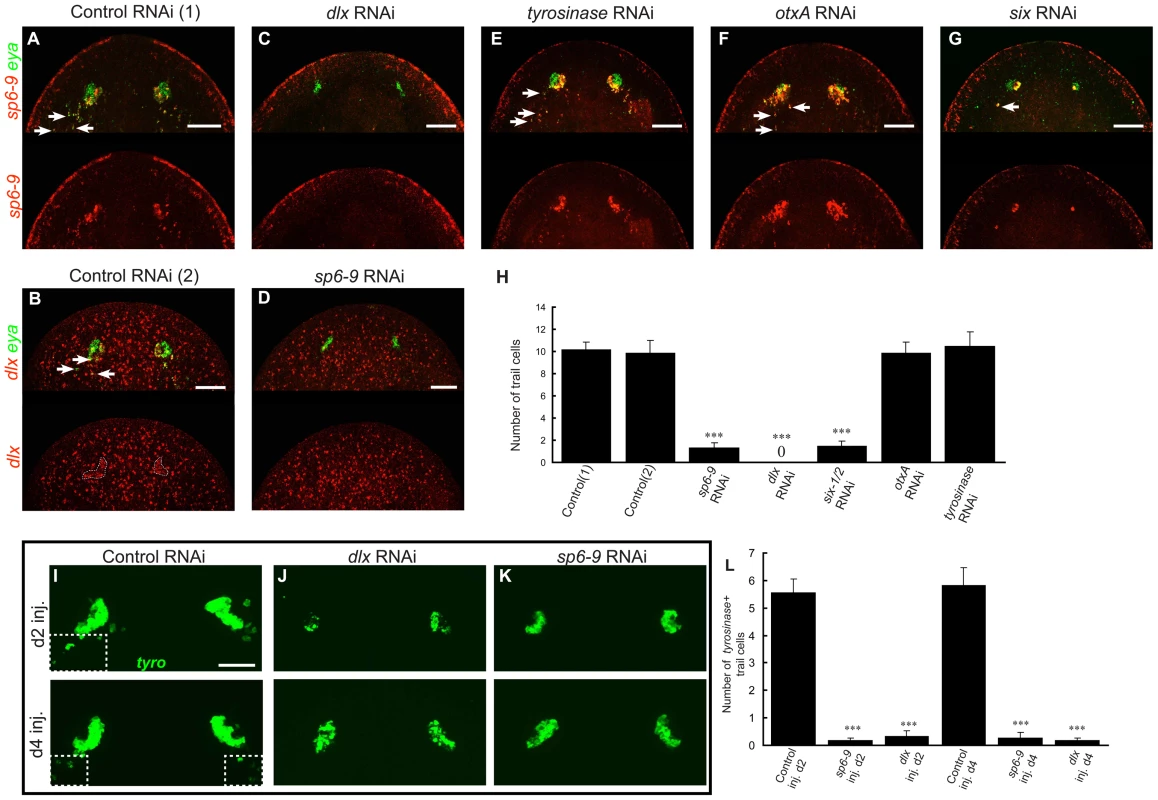
All fluorescence is FISH. Day 7 regeneration blastemas are shown, anterior is up. (A–G) Photoreceptor defects following RNAi of eye-expressed genes. Control RNAi animals have co-localization of (A) sp6-9 with eya, or (B) dlx with eya in the eyes and progenitor trail, n = 10/10 animals for each. (C), dlx(RNAi) animals do not regenerate sp6-9+/eya+ cells, n = 10/10. (D) sp6-9(RNAi) animals do not regenerate dlx+/eya+ cells, n = 10/10. (E–F) tyrosinase(RNAi) and otxA(RNAi) animals are able to regenerate sp6-9+/eya+ cells. (G) six-1/2(RNAi) regenerate greatly reduced numbers of sp6-9+/eya+ cells. (H) Quantification of the number of optic cup trail cells for RNAi conditions in (A–G). For control(2) and sp6-9 RNAi, dlx and eya coexpression is used to count trail cells. For all other conditions sp6-9 and eya coexpression is used. (I–K) RNAi of sp6-9 and dlx beginning on either day 2 or day 4 of regeneration results in smaller pigment cups and fewer terminal trail cells, assessed by tyrosinase expression. Boxes with dashed outline in (I) enclose terminal trail cells. (L) Quantification of number of tyrosinase+ trail cells in the indicated RNAi conditions. Arrows indicate double positive cells. Scale bars, 50 µm. The RNAi experiments described above were performed by feeding dsRNA to animals for one week prior to cutting. In order to confirm that sp6-9 and dlx gene functions are required during regeneration, we applied dsRNA only after amputation by using an injection strategy. dsRNA was applied beginning at either day 2 or day 4 of regeneration following decapitation. In both cases, pigment cups were typically present at day 7 of regeneration in dlx(RNAi) and sp6-9(RNAi) animals, as expected because some progenitors are formed before the onset of RNAi (Figure 6I–6L). However, RNAi animals lacked the tyrosinase+ terminal trail cells posterior to the eye (>97% reduction for dlx and sp6-9 RNAi at d2) and had pigment cups that were smaller than in control animal (Figure 6I–6L and Figure S14). Therefore, inhibition of sp6-9 and dlx only during regeneration successfully impaired ongoing pigment cup cell specification. Despite a reduction in total numbers, the pigment cup cells that were generated successfully aggregated into concave cups properly positioned in the head. We conclude, based on these data, that dlx and sp6-9 are required for progenitor formation during regeneration of the optic cup (Figure 7).
Fig. 7. A model for cell state changes in pigment cup regeneration. 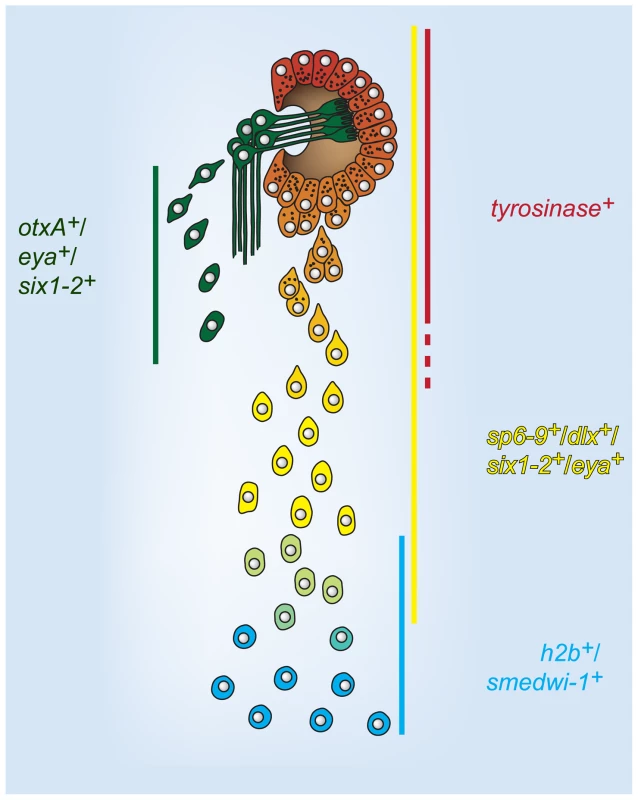
Lines indicate domain of expression of the adjacent gene labels in the optic cup trail during regeneration. Regenerative photoreceptor neuron cells are shown as spatially separated for clarity. This model proposes that optic cup progenitors are specified at distance from the aggregated cells of the optic cup primordium, within the neoblast population (h2b+, smedwi-1+). Progenitors undergo changes in gene expression, including loss of neoblast markers and activation of differentiation markers, as migration towards the eye primordium proceeds. Ultimately, terminally differentiating progenitors incorporate into the eye and undergo a mesenchymal-to-epithelial transition. Discussion
A cellular model for regeneration of the eye
All regions of the planarian body that can regenerate contain somatic dividing cells (neoblasts), which are typically identified by expression of cell cycle genes and smedwi-1, a piwi homolog. Recent work has shown that some of the adult dividing cells are pluripotent stem cells, called cNeoblasts, which are capable of generating all essential cells of the body [8]. Here we show that some neoblasts have gene expression and distribution that is indicative of restriction to a specific cell type, the optic cup cells of the eye. Optic cup progenitors are induced in abundance within the parenchyma of the animal after decapitation, at a distance from the (now absent) site of the lost eye as well as the aggregated primordium of the newly regenerating eye. Differentiation and migration toward the eye primordium are temporally correlated, as expression of neoblast markers is downregulated and expression of the differentiation marker tyrosinase is upregulated in progenitor cells approaching the eye (Figure 7). Therefore, we spatially and molecularly identify progenitors between the pluripotent state and the terminally differentiated state during regeneration of a specific tissue. The eye is an ideal system with which to address questions of how the identity of lost tissue impacts the generation of regenerative cells, as its pigmentation and restricted localization greatly simplify targeting for surgery. Therefore, many questions can now be asked about how specification, migration, and differentiation of a specific progenitor population are regulated following diverse injuries to a specific organ. For example, what is the minimal injury to the eye required for induction of eye progenitors?
Previous work has suggested that optic cup cells and photoreceptor neurons in planarians derive from common progenitors cell that expresses terminal differentiation markers of both the optic cup cells (tryptophan hydroxylase) and photoreceptor neurons (arrestin) [26]. By contrast, our work indicates that progenitors for photoreceptor neurons and cup cells exist as distinct progenitor populations prior to terminal differentiation and aggregation in the eye primordium, based on distinct expression of dlx/sp6-9 and otxA in mesenchymal six+/eya+ cells present posterior to the eye. With these results we cannot exclude that all eye cells derive from a common upstream, eye-specific progenitor. However, it is apparent that they exist as spatially distinct populations with distinct gene expression for a substantial period before differentiation and aggregation into an eye primordium. In most developing eyes, including in Drosophila and vertebrates, pigment cells and photoreceptor neurons are formed from a common epithelial eye field [27]. We find no epithelial eye field that contains proliferating or undifferentiated eye progenitors in the regenerating or intact planarian. Therefore, it will be interesting to determine whether the pigment cells and photoreceptor neurons likewise share a common eye-specific precursor population in this organism.
A significant consequence of development within an epithelial field is that position and numerical proportion of various cell types can be influenced by direct signaling from one cell type to another. For instance, in Drosophila eye development many cell fate decisions depend on direct cell-cell signaling via Notch/Delta, including the induction of primary pigment cells by adjacent cone cells [28]. Signaling between mesenchymal eye progenitors in planarians is less likely to depend on cell contact. Longer-range signaling molecules in planarians could be responsible for coordinating progenitor numbers and position. Perturbation with RNAi of Wnt [29]–[33], Bmp [34]–[36], or Fgf [37] signaling in planarians can result in ectopic eyes. It is unknown, however, whether these pathways act directly in eye progenitor specification or whether ectopic photoreceptors are the indirect result of major regional tissue changes in these RNAi animals, such as expansion of the brain. Investigation of the signaling molecules that directly act on eye progenitors might promote understanding of how progenitor numbers are coordinated during de novo organ regeneration.
Transcription regulatory genes and eye regeneration
The planarian eye has important similarities to many animal eyes, including expression of regulatory transcription factors such sine oculis [3] and eyes absent [4], downstream genes such as opsin, arrestin, and tyrosinase, as well as the overall morphology of a pigmented cup that encapsulates photoreceptive organelles. We identify planarian dlx and sp6-9 as novel regulators of optic cup progenitor formation during planarian eye regeneration. Dlx and Sp6-9 transcription factors are well conserved throughout the Eumetazoa and can function as an evolutionarily conserved module. Together, Dlx and Sp6-9 genes regulate limb outgrowth in both insects and vertebrates [38]–[42] and are important for development of the vertebrate tooth [40], [43]–[44], another appendage-like structure. Dlx and Sp6-9 family genes are also both expressed in the anterior of the body axis in Drosophila embryos and the basal deuterostome Saccoglossus [45], although a coordinated role for these two genes in the anterior of the body axis has not yet been supported by functional data. Multiple Dlx genes are expressed in the vertebrate eye from early stages of retina formation [46]–[47], and a Dlx gene is expressed in the adult Platynereis cup-shaped eye [48].
Planarian eye regeneration differs from eye development in a number of other animals in that it does not appear to rely on orthologs of the pax6(eyeless) [49] or dachshund genes, two important components of the Drosophila retinal determination gene network (RDGN). Pax6 genes are important regulators of eye development across many animal lineages [50]. dachshund is required for Drosophila eye development [51], and dachshund orthologs are expressed, but not functionally required, during mouse eye development [52]. Therefore, potentially ancestral eye specification genes [21] might not be involved in planarian eye regeneration. One possible explanation for this is that differences in specification mechanisms may exist between embryonic development and adult regeneration. The roles for eye regeneration genes are currently unknown in planarian embryonic eye development. Another possibility, if these genes indeed had roles in eye specification in the ancestor of Bilateria, is that planarians have diverged over evolutionary time with respect to their reliance on pax6 and dachshund.
Planarians also differ from vertebrates, but are similar to Drosophila, in the role of Orthodenticle homologs in pigment cell specification in the eye. Vertebrate RPE development depends on otx2 [53], and regeneration of the newt RPE is associated with expression of otx [54]. By contrast, in planarians the only known eye-expressed orthodenticle homolog, otxA, is expressed specifically in the photoreceptor neurons, and not the pigmented cells (Figure 1H) [20] and is primarily required for photoreceptor neuron specification (Figure 1N). Similarly, in Drosophila development, orthodenticle is required for photoreceptor neuron development, and a role in pigment cells of the eye has not been described [55]. Therefore, the function of Orthodenticle homologs in maintenance and specification of pigmented eye cells may differ between protostomes and deuterostomes.
Despite these differences, the demonstrated Eumetazoa-wide conservation of numerous other eye genes, as well as the demonstrated capacity of Dlx and Sp6-9 to execute deeply conserved developmental programs, indicates that investigation of this gene pair in diverse animals will be an important direction for the study of optic cup formation and regeneration. Furthermore, the identification of a highly localized eye progenitor population based on expression of dlx and sp6-9 will facilitate the exploration of progenitor dynamics during regeneration.
Materials and Methods
Gene cloning
Genes were cloned from cDNA using gene-specific primers designed from EST databases and gene predictions (Genscan and Maker) [56]. For cDNA library generation, RNA from mixed stage regenerating animals was isolated in Trizol, and used as template for first-strand reverse transcription with Superscript III (Invitrogen). PCR using cDNA template and gene-specific primers was typically followed by a secondary, interval PCR to improve specificity and yield. The following primer sequences were used with gateway adapters or addition of T7 promoter sequence (see below) for cloning genes.
dachshund
5′ primer: GTGGGCTTCACACGGTTTAT
nested 5′ primer: TTGAAGAGACTAGAAATCGTTCCA
3′ primer: TTGCACAAACTTTGCAGGAG
dlx
5′ primer: AATGAACCTCCCACTGCA
nested 5′ primer: CAGGATCAGAAACCCAATCC
3′ primer: CGGTTATTCGAAAAATTAACTGG
eyes absent
5′ primer: GGCCTTTCAAAAGACGACTC
3′ primer: AAGACTCAATGCGTGGTGAA
opsin
5′ primer: TGGTTTCATCGGTGGTCTTT
nested 5′ primer: TGGGTTTATATCCATCAACACAAT
3′ primer: TTTTTGCACCCGTTTTCAT
otxA
5′ primer: CCACAAATCCCTCTCTACGG
nested 5′ primer: ACGTAGCTGGGATCAACACC
3′ primer: TGGACCTGACAAATTGTTCC
six
5′ primer: ATCGATATCCACGAGCCAAG
3′ primer: CCAGATTCGCATTCGTTACTT
nested 3′ primer: ACAGGACTCCGAACAAATCA
sp6-9
5′ primer: TTCAATAAATAACGTTGAGAGCAA
nested 5′ primer: ATCAATCTTGGCTATTGGAACG
3′ primer: TTCACAATTGTTTGTTAACGACTC
tyrosinase
5′ primer: TGCTCGTAATCACAATAGGCATAG
nested 5′ primer: TTTGCATCTTTCTTACCTTTGAGT
3′ primer: TTTCTTAATAGCCAAATTTCAAAGA
For gateway cloning, the following adapters were appended to the above primers
5′ primer adapter: AAGCTGGAGCTCCACCGCGG
3′ primer adapter: GGGCGAATTGGGTACCGGG
cDNA amplified by PCR was cloned into pGEM (pGEM T-Easy, Promega) for use in riboprobe and dsRNA reactions (see below). For RNAi experiments by feeding, gateway recombination was used to clone genes in pPR244, a dsRNA-expression vector, as described [24]. For determination of complete gene sequences, 5′ and 3′ RACE libaries were generated from mixed stage planarian RNA (FirstChoice RLM-RACE, Ambion). Nested primers were designed to either the 5′ or 3′ prime end of the known gene sequence, and candidate bands following the second (nested) PCR were excised, purified, cloned into pGEM, and sequenced.
RNA synthesis
To generate template for riboprobe synthesis, amplified cDNAs were first cloned into pGEM (as above). Template was generated with same primers as were used for cloning, except that gateway adapters were not present on primers, and T7 promoter sequence was appended to the 3′ primer. Transcription reactions were performed with T7 (Promega) and either DIG-, FITC - (Roche), or DNP - (Perkin Elmer) labeled ribonucleotides. RNA was purified using ethanol precipitation with 7.5 M ammonium actetae (1∶2). Pellets were resuspended in formamide (50 µl per 25 µl synthesis reaction), and stored at −20°.
To generate template for dsRNA synthesis in vitro, generic primers recognizing gateway adapter sequences were used with T7 appended to either the forward or reverse primer, and PCR was performed using amplified cDNA (with gateway adapters) cloned into pGEM. T7 transcription reactions for sense and antisense RNA were performed separately, and pooled prior to phenol-chloroform extraction followed by ethanol precipitation with sodium acetate (1∶10). RNA was annealed using the following program in a thermocycler: 70°C for 10′, 37°C for 30′, 4°C for 10′.
RNAi
RNAi was performed by feeding intact animals with E. coli expressing dsRNA under an inducible promoter. HT115 competent cells were transformed with pPR244 vector [57] containing the gene of interest. Negative control experiments used a 1 kb region of unc-22, a Caenorhabditis elegans gene with no significant nucleotide sequence similarity to a planarian gene. Cultures were grown in 2×YT media to an OD600 of 0.35–0.45, and then induced for two hours with 1 mM IPTG. Cultures were pelleted and resuspended in a volume of 70% liver/30% water equal to 1/300th of the original volume of the culture, and stored at −80°C. Worms were fed three times prior to amputation. Amputation was performed on the day following the final feeding, and worms were fixed following one round of regeneration. For targeted eye surgery, a microsurgical blade (MSP, 15°, 3 mm depth) was inserted into the eye and the optic cup was excised upon removal of the blade. Surgery was performed one day after the third RNAi feeding, and one additional feeding was administered 4 days after surgery. For homeostasis RNAi experiments, animals were fed every four days.
For RNAi by injection, dsRNA was diluted to 4 µg/µl in water. For standard RNAi application (dachshund RNAi experiment), animals were amputated and injected after 30 min, and again after 24 hours. After 3 days of regeneration, animals were amputated again and injected once more after 30 min. For RNAi injection in Figure 6I–6K, animals were injected once, either on day 2 or day 4 of regeneration after decapitation. Injection was performed with a Drummond Nanoject II and 3.5″ Drummond capillaries (3-000-203-G/X). Animals were immobilized for injection with use of a peltier cooling block. Several injections of 32 nl were applied as needed to observe swelling of animals with liquid.
Histology and imaging
Whole-mount fluorescent in situ hybridization (FISH) and antibody staining was performed as described [58] except that FITC-tyramide was used at 1∶500, and HRP enzyme was inactivated using 4% formaldehyde. Tyramide was generated by conjugation of succinimidyl esters of rhodamine, FITC, Cy5 and AMCA with tyramide-HCL (Roche) [59]. Riboprobes (see below) were used at 1∶800 dilution in hyb solution, except for opsin and tyrosinase, which were used at 1∶1200. Anti-ARRESTIN (VC-1) antibody (1∶5000 dilution) was kindly provided by Kiyo Agata, and SMEDWI-1 antibody (1∶1000 dilution) was obtained as described [10], [22].
For BrdU experiments, animals were injected with a solution in 5 mg/ml of BrdU (Fluka) in planarian water. Similar injection methods were used as for dsRNA injection (see above), and animals were only injected once. Animals were then fixed and labeled according to the regular FISH protocol, and following FISH development animals were treated with 2N HCl for 45 minutes at RT. BrdU was detected with rat-anti-BrdU (1∶100) antibody (Oxford Biotech) followed by incubation with anti-rat-HRP (1∶100) (Abcam) and development with commercial tyramide (Invitrogen).
Optical sectioning was performed using an Apotome, Axiocam digital camera, a Zeiss AxioImager, and Axiovision software. Images in Figure 5A, Figures S4 and S8 were generated with a Zeiss confocal microscope (LSM 700). Brightness, contrast, and gamma were adjusted as needed to improve visibility. In most images, optical sections are overlayed to show co-expression in cells across a depth of tissue. Stacks of sections were manually examined at the level of individual optical sections to determine true instances of co-expression, and instances of artifact overlapping signal created by digital overlaying were excluded from analyses.
Phylogenetic analyses
S. mediterranea amino acid sequence predictions were aligned (ClustalW) with sequences of putatively homologous proteins from other metazoans. Sequences were trimmed by gblocks under the lowest stringency settings. Phylogenetic trees for tyrosinase, sp6-9, dlx, and otxA were constructed using Bayesian inference (MrBayes) with >2,000,000 generations, and >1,500 burn-in trees discarded. Branch labels display posterior probabilities.
Supporting Information
Zdroje
1. StenkampDL 2007 Neurogenesis in the fish retina. Int Rev Cytol 259 173 224
2. OkadaTS 1980 Cellular metaplasia or transdifferentiation as a model for retinal cell differentiation. Curr Top Dev Biol 16 349 380
3. PinedaDGonzalezJCallaertsPIkeoKGehringWJ 2000 Searching for the prototypic eye genetic network: Sine oculis is essential for eye regeneration in planarians. Proc Natl Acad Sci U S A 97 4525 4529
4. ManniniLRossiLDeriPGremigniVSalvettiA 2004 Djeyes absent (Djeya) controls prototypic planarian eye regeneration by cooperating with the transcription factor Djsix-1. Dev Biol 269 346 359
5. SatoYKobayashiKMatsumotoMHoshiMNegishiS 2005 Comparative study of eye defective worm ‘menashi’ and regenerating wild-type in planarian, Dugesia ryukyuensis. Pigment Cell Res 18 86 91
6. NilssonDE 2009 The evolution of eyes and visually guided behaviour. Philos Trans R Soc Lond B Biol Sci 364 2833 2847
7. StraussO 2005 The retinal pigment epithelium in visual function. Physiol Rev 85 845 881
8. WagnerDEWangIEReddienPW 2011 Clonogenic neoblasts are pluripotent adult stem cells that underlie planarian regeneration. Science 332 811 816
9. ReddienPWSánchez AlvaradoA 2004 Fundamentals of planarian regeneration. Annu Rev Cell Dev Biol 20 725 757
10. WenemoserDReddienPW 2010 Planarian regeneration involves distinct stem cell responses to wounds and tissue absence. Dev Biol 344 979 991
11. NewmarkPASánchez AlvaradoA 2000 Bromodeoxyuridine specifically labels the regenerative stem cells of planarians. Dev Biol 220 142 153
12. EisenhofferGTKangHSánchez AlvaradoA 2008 Molecular analysis of stem cells and their descendants during cell turnover and regeneration in the planarian Schmidtea mediterranea. Cell Stem Cell 3 327 339
13. HaseSWakamatsuKFujimotoKInabaAKobayashiK 2006 Characterization of the pigment produced by the planarian, Dugesia ryukyuensis. Pigment Cell Res 19 248 249
14. MullerGRuppertSSchmidESchutzG 1988 Functional analysis of alternatively spliced tyrosinase gene transcripts. Embo J 7 2723 2730
15. Sánchez AlvaradoANewmarkPA 1999 Double-stranded RNA specifically disrupts gene expression during planarian regeneration. Proc Natl Acad Sci U S A 96 5049 5054
16. SchaeperNDPrpicNMWimmerEA 2010 A clustered set of three Sp-family genes is ancestral in the Metazoa: evidence from sequence analysis, protein domain structure, developmental expression patterns and chromosomal location. BMC Evol Biol 10 88
17. PanganibanGRubensteinJL 2002 Developmental functions of the Distal-less/Dlx homeobox genes. Development 129 4371 4386
18. RanadeSSYang-ZhouDKongSWMcDonaldECCookTA 2008 Analysis of the Otd-dependent transcriptome supports the evolutionary conservation of CRX/OTX/OTD functions in flies and vertebrates. Dev Biol 315 521 534
19. NilssonDE 2004 Eye evolution: a question of genetic promiscuity. Curr Opin Neurobiol 14 407 414
20. UmesonoYWatanabeKAgataK 1999 Distinct structural domains in the planarian brain defined by the expression of evolutionarily conserved homeobox genes. Dev Genes Evol 209 31 39
21. SilverSJRebayI 2005 Signaling circuitries in development: insights from the retinal determination gene network. Development 132 3 13
22. GuoTPetersAHNewmarkPA 2006 A Bruno-like gene is required for stem cell maintenance in planarians. Dev Cell 11 159 169
23. HewitsonTDKelynackKJDarbyIA 2006 Histochemical localization of cell proliferation using in situ hybridization for histone mRNA. Methods Mol Biol 326 219 226
24. ReddienPWOviedoNJJenningsJRJenkinJCSánchez AlvaradoA 2005 SMEDWI-2 is a PIWI-like protein that regulates planarian stem cells. Science 310 1327 1330
25. HayashiTShibataNOkumuraRKudomeTNishimuraO 2010 Single-cell gene profiling of planarian stem cells using fluorescent activated cell sorting and its “index sorting” function for stem cell research. Dev Growth Differ 52 131 144
26. TakedaHNishimuraKAgataK 2009 Planarians maintain a constant ratio of different cell types during changes in body size by using the stem cell system. Zoolog Sci 26 805 813
27. CaganR 2009 Principles of Drosophila eye differentiation. Curr Top Dev Biol 89 115 135
28. NagarajRBanerjeeU 2007 Combinatorial signaling in the specification of primary pigment cells in the Drosophila eye. Development 134 825 831
29. PetersenCPReddienPW 2009 A wound-induced Wnt expression program controls planarian regeneration polarity. Proc Natl Acad Sci U S A 106 17061 17066
30. GurleyKARinkJCSánchez AlvaradoA 2008 Beta-catenin defines head versus tail identity during planarian regeneration and homeostasis. Science 319 323 327
31. PetersenCPReddienPW 2008 Smed-beta-catenin-1 is required for anteroposterior blastema polarity in planarian regeneration. Science 319 327 330
32. IglesiasMGomez-SkarmetaJLSaloEAdellT 2008 Silencing of Smed-beta-catenin-1 generates radial-like hypercephalized planarians. Development 135 1215 1221
33. AdellTSaloEBoutrosMBartschererK 2009 Smed-Evi/Wntless is required for beta-catenin-dependent and -independent processes during planarian regeneration. Development 136 905 910
34. MolinaMDSaloECebriaF 2007 The BMP pathway is essential for re-specification and maintenance of the dorsoventral axis in regenerating and intact planarians. Dev Biol 311 79 94
35. ReddienPWBermangeALKiczaAMSánchez AlvaradoA 2007 BMP signaling regulates the dorsal planarian midline and is needed for asymmetric regeneration. Development 134 4043 4051
36. OriiHWatanabeK 2007 Bone morphogenetic protein is required for dorso-ventral patterning in the planarian Dugesia japonica. Dev Growth Differ 49 345 349
37. CebriaFKobayashiCUmesonoYNakazawaMMinetaK 2002 FGFR-related gene nou-darake restricts brain tissues to the head region of planarians. Nature 419 620 624
38. KawakamiYEstebanCRMatsuiTRodriguez-LeonJKatoS 2004 Sp8 and Sp9, two closely related buttonhead-like transcription factors, regulate Fgf8 expression and limb outgrowth in vertebrate embryos. Development 131 4763 4774
39. TalamilloADelgadoINakamuraTde-VegaSYoshitomiY 2010 Role of Epiprofin, a zinc-finger transcription factor, in limb development. Dev Biol 337 363 374
40. HertveldtVLouryanSvan ReethTDrezePvan VoorenP 2008 The development of several organs and appendages is impaired in mice lacking Sp6. Dev Dyn 237 883 892
41. SchaeperNDPrpicNMWimmerEA 2009 A conserved function of the zinc finger transcription factor Sp8/9 in allometric appendage growth in the milkweed bug Oncopeltus fasciatus. Dev Genes Evol 219 427 435
42. EstellaCMannRS 2010 Non-redundant selector and growth-promoting functions of two sister genes, buttonhead and Sp1, in Drosophila leg development. PLoS Genet 6 e1001001 doi:10.1371/journal.pgen.1001001
43. ThomasBLTuckerASQuiMFergusonCAHardcastleZ 1997 Role of Dlx-1 and Dlx-2 genes in patterning of the murine dentition. Development 124 4811 4818
44. NakamuraTUndaFde-VegaSVilaxaAFukumotoS 2004 The Kruppel-like factor epiprofin is expressed by epithelium of developing teeth, hair follicles, and limb buds and promotes cell proliferation. J Biol Chem 279 626 634
45. LemonsDFritzenwankerJHGerhartJLoweCJMcGinnisW 2010 Co-option of an anteroposterior head axis patterning system for proximodistal patterning of appendages in early bilaterian evolution. Dev Biol 344 358 362
46. DhawanRRSchoenTJBeebeDC 1997 Isolation and expression of homeobox genes from the embryonic chicken eye. Mol Vis 3 7
47. de MeloJDuGFonsecaMGillespieLATurkWJ 2005 Dlx1 and Dlx2 function is necessary for terminal differentiation and survival of late-born retinal ganglion cells in the developing mouse retina. Development 132 311 322
48. ArendtDHausenHPurschkeG 2009 The ‘division of labour’ model of eye evolution. Philos Trans R Soc Lond B Biol Sci 364 2809 2817
49. PinedaDRossiLBatistoniRSalvettiAMarsalM 2002 The genetic network of prototypic planarian eye regeneration is Pax6 independent. Development 129 1423 1434
50. KozmikZ 2008 The role of Pax genes in eye evolution. Brain Res Bull 75 335 339
51. MardonGSolomonNMRubinGM 1994 dachshund encodes a nuclear protein required for normal eye and leg development in Drosophila. Development 120 3473 3486
52. DavisRJPesahYIHardingMPaylorRMardonG 2006 Mouse Dach2 mutants do not exhibit gross defects in eye development or brain function. Genesis 44 84 92
53. Martinez-MoralesJRDolezVRodrigoIZaccariniRLeconteL 2003 OTX2 activates the molecular network underlying retina pigment epithelium differentiation. J Biol Chem 278 21721 21731
54. SakamiSHisatomiOSakakibaraSLiuJRehTA 2005 Downregulation of Otx2 in the dedifferentiated RPE cells of regenerating newt retina. Brain Res Dev Brain Res 155 49 59
55. VandendriesERJohnsonDReinkeR 1996 orthodenticle is required for photoreceptor cell development in the Drosophila eye. Dev Biol 173 243 255
56. CantarelBLKorfIRobbSMParraGRossE 2008 MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res 18 188 196
57. ReddienPWBermangeALMurfittKJJenningsJRSánchez AlvaradoA 2005 Identification of genes needed for regeneration, stem cell function, and tissue homeostasis by systematic gene perturbation in planaria. Dev Cell 8 635 649
58. PearsonBJEisenhofferGTGurleyKARinkJCMillerDE 2009 Formaldehyde-based whole-mount in situ hybridization method for planarians. Dev Dyn 238 443 450
59. HopmanAHRamaekersFCSpeelEJ 1998 Rapid synthesis of biotin-, digoxigenin-, trinitrophenyl-, and fluorochrome-labeled tyramides and their application for In situ hybridization using CARD amplification. J Histochem Cytochem 46 771 777
60. SakaiFAgataKOriiHWatanabeK 2000 Organization and regeneration ability of spontaneous supernumerary eyes in planarians -eye regeneration field and pathway selection by optic nerves. Zoolog Sci 17 375 381
Štítky
Genetika Reprodukčná medicína
Článek The T-Box Factor MLS-1 Requires Groucho Co-Repressor Interaction for Uterine Muscle SpecificationČlánek B Chromosomes Have a Functional Effect on Female Sex Determination in Lake Victoria Cichlid FishesČlánek Distinct Cdk1 Requirements during Single-Strand Annealing, Noncrossover, and Crossover RecombinationČlánek Specification of Corpora Cardiaca Neuroendocrine Cells from Mesoderm Is Regulated by Notch SignalingČlánek Ongoing Phenotypic and Genomic Changes in Experimental Coevolution of RNA Bacteriophage Qβ and
Článok vyšiel v časopisePLOS Genetics
Najčítanejšie tento týždeň
2011 Číslo 8- Gynekologové a odborníci na reprodukční medicínu se sejdou na prvním virtuálním summitu
- Je „freeze-all“ pro všechny? Odborníci na fertilitu diskutovali na virtuálním summitu
-
Všetky články tohto čísla
- Polo, Greatwall, and Protein Phosphatase PP2A Jostle for Pole Position
- Genome-Wide Association Analysis of Incident Coronary Heart Disease (CHD) in African Americans: A Short Report
- The T-Box Factor MLS-1 Requires Groucho Co-Repressor Interaction for Uterine Muscle Specification
- B Chromosomes Have a Functional Effect on Female Sex Determination in Lake Victoria Cichlid Fishes
- Analysis of DNA Methylation in a Three-Generation Family Reveals Widespread Genetic Influence on Epigenetic Regulation
- PP2A-Twins Is Antagonized by Greatwall and Collaborates with Polo for Cell Cycle Progression and Centrosome Attachment to Nuclei in Drosophila Embryos
- Discovery of Sexual Dimorphisms in Metabolic and Genetic Biomarkers
- Pervasive Sharing of Genetic Effects in Autoimmune Disease
- DNA Methylation and Histone Modifications Regulate Shoot Regeneration in by Modulating Expression and Auxin Signaling
- Mutations in and Reveal That Cartilage Matrix Controls Timing of Endochondral Ossification by Inhibiting Chondrocyte Maturation
- Variance of Gene Expression Identifies Altered Network Constraints in Neurological Disease
- Frequent Beneficial Mutations during Single-Colony Serial Transfer of
- Increased Gene Dosage Affects Genomic Stability Potentially Contributing to 17p13.3 Duplication Syndrome
- Distinct Cdk1 Requirements during Single-Strand Annealing, Noncrossover, and Crossover Recombination
- Hunger Artists: Yeast Adapted to Carbon Limitation Show Trade-Offs under Carbon Sufficiency
- Suppression of Scant Identifies Endos as a Substrate of Greatwall Kinase and a Negative Regulator of Protein Phosphatase 2A in Mitosis
- Temporal Dynamics of Host Molecular Responses Differentiate Symptomatic and Asymptomatic Influenza A Infection
- MK2-Dependent p38b Signalling Protects Hindgut Enterocytes against JNK-Induced Apoptosis under Chronic Stress
- Specification of Corpora Cardiaca Neuroendocrine Cells from Mesoderm Is Regulated by Notch Signaling
- Genome-Wide Gene-Environment Study Identifies Glutamate Receptor Gene as a Parkinson's Disease Modifier Gene via Interaction with Coffee
- Identification of Functional Toxin/Immunity Genes Linked to Contact-Dependent Growth Inhibition (CDI) and Rearrangement Hotspot (Rhs) Systems
- Genomic Analysis of the Necrotrophic Fungal Pathogens and
- Celsr3 Is Required for Normal Development of GABA Circuits in the Inner Retina
- Genetic Architecture of Aluminum Tolerance in Rice () Determined through Genome-Wide Association Analysis and QTL Mapping
- Predisposition to Cancer Caused by Genetic and Functional Defects of Mammalian
- Regulation of p53/CEP-1–Dependent Germ Cell Apoptosis by Ras/MAPK Signaling
- and but Not Interact in Genetic Models of Amyotrophic Lateral Sclerosis
- Gamma-Tubulin Is Required for Bipolar Spindle Assembly and for Proper Kinetochore Microtubule Attachments during Prometaphase I in Oocytes
- Ongoing Phenotypic and Genomic Changes in Experimental Coevolution of RNA Bacteriophage Qβ and
- Genetic Architecture of a Reinforced, Postmating, Reproductive Isolation Barrier between Species Indicates Evolution via Natural Selection
- -eQTLs Reveal That Independent Genetic Variants Associated with a Complex Phenotype Converge on Intermediate Genes, with a Major Role for the HLA
- The GATA Factor ELT-1 Works through the Cell Proliferation Regulator BRO-1 and the Fusogen EFF-1 to Maintain the Seam Stem-Like Fate
- and Control Optic Cup Regeneration in a Prototypic Eye
- A Comprehensive Map of Mobile Element Insertion Polymorphisms in Humans
- An EMT–Driven Alternative Splicing Program Occurs in Human Breast Cancer and Modulates Cellular Phenotype
- Evidence for Hitchhiking of Deleterious Mutations within the Human Genome
- A Broad Brush, Global Overview of Bacterial Sexuality
- Global Chromosomal Structural Instability in a Subpopulation of Starving Cells
- A Pre-mRNA–Associating Factor Links Endogenous siRNAs to Chromatin Regulation
- Glutamine Synthetase Is a Genetic Determinant of Cell Type–Specific Glutamine Independence in Breast Epithelia
- The Repertoire of ICE in Prokaryotes Underscores the Unity, Diversity, and Ubiquity of Conjugation
- Genome-Wide Association Analysis of Autoantibody Positivity in Type 1 Diabetes Cases
- Natural Polymorphism in BUL2 Links Cellular Amino Acid Availability with Chronological Aging and Telomere Maintenance in Yeast
- Chromosome Painting Reveals Asynaptic Full Alignment of Homologs and HIM-8–Dependent Remodeling of Chromosome Territories during Meiosis
- Ku Must Load Directly onto the Chromosome End in Order to Mediate Its Telomeric Functions
- PLOS Genetics
- Archív čísel
- Aktuálne číslo
- Informácie o časopise
Najčítanejšie v tomto čísle- An EMT–Driven Alternative Splicing Program Occurs in Human Breast Cancer and Modulates Cellular Phenotype
- Chromosome Painting Reveals Asynaptic Full Alignment of Homologs and HIM-8–Dependent Remodeling of Chromosome Territories during Meiosis
- Discovery of Sexual Dimorphisms in Metabolic and Genetic Biomarkers
- Regulation of p53/CEP-1–Dependent Germ Cell Apoptosis by Ras/MAPK Signaling
Prihlásenie#ADS_BOTTOM_SCRIPTS#Zabudnuté hesloZadajte e-mailovú adresu, s ktorou ste vytvárali účet. Budú Vám na ňu zasielané informácie k nastaveniu nového hesla.
- Časopisy



