-
Články
- Časopisy
- Kurzy
- Témy
- Kongresy
- Videa
- Podcasty
An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
The genetic determination of eggshell coloration has not been determined in birds. Here we report that the blue eggshell is caused by an EAV-HP insertion that promotes the expression of SLCO1B3 gene in the uterus (shell gland) of the oviduct in chicken. In this study, the genetic map location of the blue eggshell gene was refined by linkage analysis in an F2 chicken population, and four candidate genes within the refined interval were subsequently tested for their expression levels in the shell gland of the uterus from blue-shelled and non-blue-shelled hens. SLCO1B3 gene was found to be the only one expressed in the uterus of blue-shelled hens but not in that of non-blue-shelled hens. Results from a pyrosequencing analysis showed that only the allele of SLCO1B3 from blue-shelled chickens was expressed in the uterus of heterozygous hens (O*LC/O*N). SLCO1B3 gene belongs to the organic anion transporting polypeptide (OATP) family; and the OATPs, functioning as membrane transporters, have been reported for the transportation of amphipathic organic compounds, including bile salt in mammals. We subsequently resequenced the whole genomic region of SLCO1B3 and discovered an EAV-HP insertion in the 5′ flanking region of SLCO1B3. The EAV-HP insertion was found closely associated with blue eggshell phenotype following complete Mendelian segregation. In situ hybridization also demonstrated that the blue eggshell is associated with ectopic expression of SLCO1B3 in shell glands of uterus. Our finding strongly suggests that the EAV-HP insertion is the causative mutation for the blue eggshell phenotype. The insertion was also found in another Chinese blue-shelled breed and an American blue-shelled breed. In addition, we found that the insertion site in the blue-shelled chickens from Araucana is different from that in Chinese breeds, which implied independent integration events in the blue-shelled chickens from the two continents, providing a parallel evolutionary example at the molecular level.
Published in the journal: An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken. PLoS Genet 9(1): e32767. doi:10.1371/journal.pgen.1003183
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1003183Summary
The genetic determination of eggshell coloration has not been determined in birds. Here we report that the blue eggshell is caused by an EAV-HP insertion that promotes the expression of SLCO1B3 gene in the uterus (shell gland) of the oviduct in chicken. In this study, the genetic map location of the blue eggshell gene was refined by linkage analysis in an F2 chicken population, and four candidate genes within the refined interval were subsequently tested for their expression levels in the shell gland of the uterus from blue-shelled and non-blue-shelled hens. SLCO1B3 gene was found to be the only one expressed in the uterus of blue-shelled hens but not in that of non-blue-shelled hens. Results from a pyrosequencing analysis showed that only the allele of SLCO1B3 from blue-shelled chickens was expressed in the uterus of heterozygous hens (O*LC/O*N). SLCO1B3 gene belongs to the organic anion transporting polypeptide (OATP) family; and the OATPs, functioning as membrane transporters, have been reported for the transportation of amphipathic organic compounds, including bile salt in mammals. We subsequently resequenced the whole genomic region of SLCO1B3 and discovered an EAV-HP insertion in the 5′ flanking region of SLCO1B3. The EAV-HP insertion was found closely associated with blue eggshell phenotype following complete Mendelian segregation. In situ hybridization also demonstrated that the blue eggshell is associated with ectopic expression of SLCO1B3 in shell glands of uterus. Our finding strongly suggests that the EAV-HP insertion is the causative mutation for the blue eggshell phenotype. The insertion was also found in another Chinese blue-shelled breed and an American blue-shelled breed. In addition, we found that the insertion site in the blue-shelled chickens from Araucana is different from that in Chinese breeds, which implied independent integration events in the blue-shelled chickens from the two continents, providing a parallel evolutionary example at the molecular level.
Introduction
Avian eggshell coloration is the result of crypsis or mimetism and plays important roles in filtering solar radiation and strengthening the eggshell [1]. Blue eggshell color has been proposed as post-mating signals of female phenotypic quality to their mates and is related to fitness of the offspring due to the antioxidant of biliverdin, a predominant pigment for blue eggs [2], [3]. Blue eggshells can be found not only in some wild birds, e.g. eastern bluebird [4], blue-footed booby [5], and pied flycatcher [6], but also in domestic birds such as Japanese quail [7], chickens [8] and ducks [9].
Brown and white are the two major eggshell colors in chickens. Protoporphyrin-IX, biliverdin, and biliverdin zinc chelate are the main pigments of the eggshell [10] and several blue egg laying breeds have been reported worldwide [11], [12]. The Araucana, an indigenous breed from Chile, was the first chicken breed described to lay blue eggs [8], and has been frequently used in genetic studies of the blue eggshell phenotype. In China, Dongxiang and Lushi chickens are representative breeds laying blue eggs and show dominant inheritance as that in Araucana. However, the blue eggshell phenotype has not been fixed in these three breeds which still produce brown eggs at low frequency.
Blue eggshell color exhibits an autosomal dominant inheritance and eggs laid by homozygotes are a darker blue than those from heterozygotes (Figure 1A). In 1933, Punnett firstly reported that blue or green shell appearance of the Araucana was determined by a single genetic factor, traditionally denoted as oocyan (O) [8]. A series of linkage analysis involving O have been performed with O affirmatively mapped to the short arm of chromosome 1 [12]–[16], and closely linked to ev1 and P which was identified as SRY (sex determining region Y)-box 5 (SOX5) [12], [13], [16], [17]. In the region around ev1, two single nucleotide polymorphisms (SNPs) (rs15297163 and rs15297165) were found to be highly associated with the blue eggshell phenotype [18]. A 1.8 Mb genomic interval harboring the O gene was defined in an F2 resource population [19]. The localization of the O was further refined to the vicinity of ss244244378 by linkage and association analysis [20]. The ss244244378 is very close to the two SNPs reported by Zhao et al. [18] with a physical distance of 0.12 Mb implies that the region around the three SNPs is mostly like to harbor the blue eggshell gene. Combined mapping information from traditional breeds and Chilean village chickens allowed the O to be fine mapped to two small regions (Gga 1 : 67.25–67.28 Mb, Gga 1 : 67.28–67.32 Mb) [21].
Fig. 1. The eggshell color and the expression of SLCO1B3 in the uterus of blue-shelled and non-blue-shelled chickens. 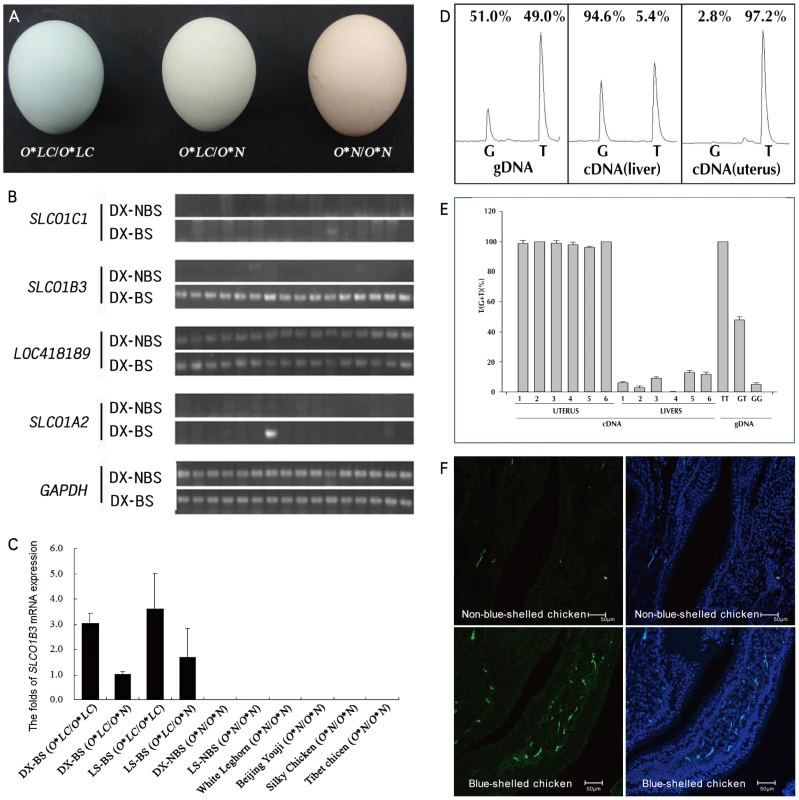
(A) Eggshell colors of homozygous blue-shelled (O*LC/O*LC), heterozygous blue-shelled (O*LC/O*N) and brown-shelled (O*N/O*N) of Dongxiang chickens. (B) The expression analysis of the four genes which are located in the refined region between markers L4 and L5 in the uterus of Dongxiang blue-shelled hens (n = 16) and Dongxiang non-blue-shelled hens (n = 16) by RT-PCRs. DX-BS: blue-shelled Dongxiang, DX-NBS: non-blue-shelled Dongxiang. (C) Analysis of SLCO1B3 expression in the uterus. Expression data were presented as fold relative to heterozygous blue-shelled Dongxiang chickens (O*LC/O*N) by the comparative Ct method (2−ΔΔCt). SLCO1B3 is exclusively expressed in blue-shelled chickens, and the amount of SLCO1B3 transcripts in homozygous blue-shelled chicken (O*LC/O*LC) is approximately two to three folds of that in heterozygous individuals (O*LC/O*N). DX-BS: blue-shelled Dongxiang, LS-BS: blue-shelled Lushi, DX-NBS: non-blue-shelled Dongxiang, LS-NBS: non-blue-shelled Lushi. (D) Micrographs of cDNA in situ hybridization for SLCO1B3 mRNA in the uterus from blue-shelled and non-blue-shelled chickens. (E) Differential expression of SLCO1B3 transcript in the uterus from blue-shelled heterozygotes using genomic DNA (gDNA) as control. The polymorphic position g. 67334934 G>T was used to monitor differential expression using pyrosequencing. T and G at this position correspond to blue-shell and non-blue-shell alleles, respectively. Due to two Ts next to the SNP at 3′end, the peaks of T in the schema contain three Ts including one T from blue-shell allele and two Ts from non-blue-shell allele. The percent expression on the peaks for T and G are the T or G at g. 67334934 G>T. (F) Summary of the detection of differential expression in uterus and liver from six heterozygotous blue-shelled (O*LC/O*N) birds. In the present study we found that the blue eggshell phenotype in chickens is caused by a retrovirus insertion in the 5′ flanking region of SLCO1B3 coding a membrane transporter OATP1B3 which is responsible for transporting amphipathic organic compounds including bile salt.
Results
Linkage analysis of Chicken blue eggshell gene
A linkage analysis was performed in an F2 resource population segregating for the O gene to refine the location of chicken blue shell gene in the present study. Eight molecular markers in the candidate region were used for linkage analysis (Table S1). By two-point analysis, the O gene was mapped in the region between marker L4 and L5 which were the closest flanking markers to O with recombination rate being both 0.02 (LOD = 15.84) (Figure 2). Fifteen SNP markers between L4 and L5 were further genotyped in the F2 resource population to narrow the mapping region and the O was finally located in a ∼120 kb region from 67296991 bp to 67416784 bp on chromosome 1 on the UCSC chicken genome (May 2006 assembly) (Table S1) and no recombination was found between the blue eggshell phenotype and the markers within the region.
Fig. 2. Refined localization of O on chicken chromosome 1. 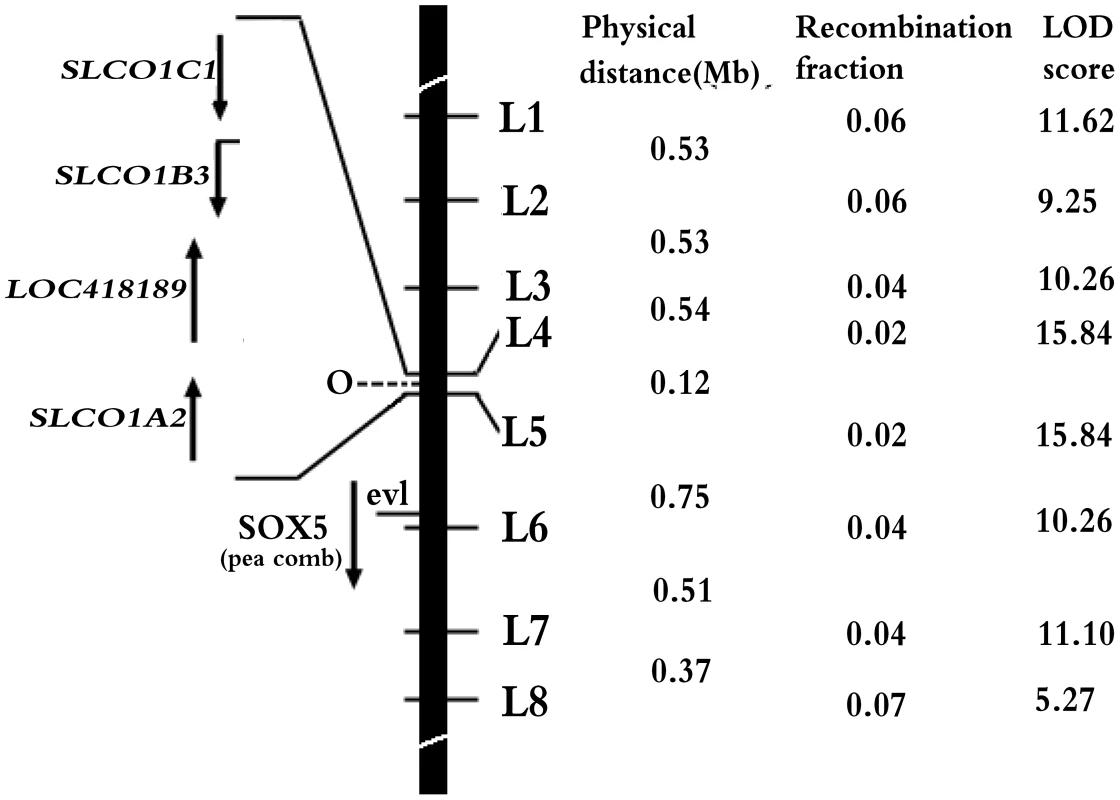
The black bar represents the short arm of chicken chromosome 1. Numbers in middle column are recombination fractions between markers and O, the corresponding LOD scores are shown in the last column. Here, O is assigned to the interval of L4 and L5 at Chr1: 67296991–67419904. Four candidate genes in the interval, EAV-HP insertion with recombination fraction and LOD in the parentheses, SRY (sex determining region Y)-box 5 (SOX5) for pea comb phenotype [17], ev1 marker [16] are indicated at the left of the black bar. Specific expression of SLCO1B3 in uterus of blue-shelled chicken
Totally, four genes (SLCO1C1, SLCO1B3, LOC418189 and SLCO1A2) were found in the ∼120 kb interval by a MapView search (http://www.ncbi.nlm.nih.gov/mapview/) (Table S2). The uterus is where pigment is secreted to eggshell. We performed expression analysis for the four candidate genes in the uterus of blue-shelled (n = 16) and brown-shelled (n = 16) Dongxiang hens by RT-PCR. We found that SLCO1B3 was the only gene expressed specifically in the uterus of blue-shelled Dongxiang chickens (Figure 1B), thus we measured its expression in the uterus in 8 blue-shelled (4 O*LC/O*LC and 4 O*LC/O*N. O*LC is for the blue-shell allele in the two Chinese breeds, Dongxiang and Lushi Chicken and O*N denotes the non-blue-shell allele or wild-type allele) and 4 brown-shelled (O*N/O*N) Dongxiang chickens, 6 blue-shelled (3 O*LC/O*LC and 3 O*LC/O*N) and 6 brown-shelled (O*N/O*N) Lushi chickens, and 24 chickens from three brown-shelled and one white-shelled breeds (O*N/O*N, 6 chickens per breed) by real-time PCR. All blue-shelled chickens expressed the gene in the uterus while non-blue-shelled chickens did not (Figure 1C). In addition, expression of SCLO1B3 was 2 to 3 fold higher in homozygous blue-shelled chickens than in heterozygous blue-shelled Dongxiang and Lushi individuals (Figure 1C). Fluoscence labeled cDNA in situ hybridization demonstrated that the transcripts of SLCO1B3 were only expressed in the uterus of blue-shelled but not brown-shelled hens (Figure 1D). These results suggest that SLCO1B3 is the causative gene for blue eggshell in the chicken.
Allele-specific expression of SLCO1B3
We found a SNP (g.67334934 G>T) in exon 5 of SLCO1B3 gene by sequencing the coding region and the SNP presented complete association with the blue eggshell phenotype in the Dongxiang chicken by genotyping it in Dongxiang blue-shelled and brown-shelled chickens. With six heterozygous individuals produced by mating a homozygous Dongxiang blue-shelled male with a White Leghorn female, the allelic expression of SLCO1B3 gene was demonstrated by RT-PCR analysis and pyrosequencing. More than 95% of the transcripts expressed in the uterus originated from the T allele corresponding to the blue-shell allele (Figure 1E). This means the expression of the gene is regulated by a cis-acting element. Surprisingly, its expression in liver is also allele specific, and ∼95% of the transcripts in liver come from the G allele which is non-blue-shell allele (Figure 1F).
An EAV-HP insertion is completely associated with blue eggshell phenotype
We sequenced the genomic region of SLCO1B3 in order to reveal the potential causative mutation of the gene with 5 blue-shelled and 5 brown-shelled Dongxiang chickens. Twenty-one SNPs evenly covering the whole genomic region (∼24 kb) of SLCO1B3 were taken for genotyping in 353 chickens from 3 blue-shelled breeds (Araucana, Dongxiang and Lushi) and 9 non-blue-shelled breeds. However, none of the SNPs was found to be in complete linkage disequilibrium with blue eggshell (Table 1).
Tab. 1. Distribution of allelic frequencies of SNPs and EAV-HP in SLCO1B3 in several blue-shelled and non-blue-shelled breeds. 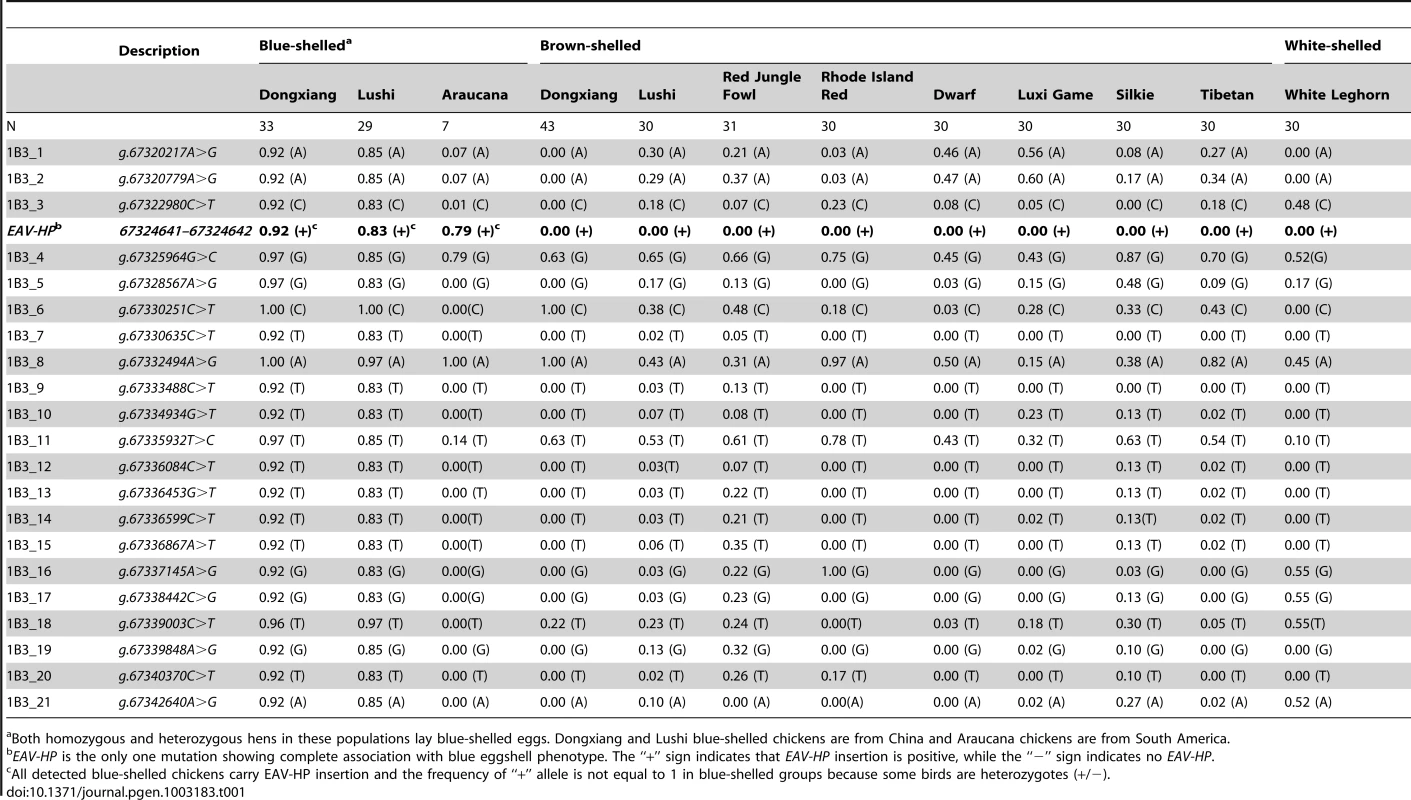
Both homozygous and heterozygous hens in these populations lay blue-shelled eggs. Dongxiang and Lushi blue-shelled chickens are from China and Araucana chickens are from South America. We subsequently cloned the 5′UTR (GenBank accession number: JN381032) of SLCO1B3 by 5′ RACE in a blue-shelled (O*LC/O*LC) and a brown-shelled (O*N/O*N) Dongxiang chicken and an extra 24 bps were found at the beginning of 5′UTR end in blue-shelled Dongxiang chicken (Figure S1). We further sequenced 5 kb upstream of the promoter using 5 blue-shelled (O*LC/O*LC) and 5 brown-shelled (O*N/O*N) Dongxiang chickens. A ∼4.2 kb insertion adjacent to 5′UTR containing the extra 24 bps was found in the blue-shelled but not in the brown-shelled chickens. The sequence of the ∼4.2 kb insertion (GenBank accession number: JF837512) represents an incomplete retrovirus and shows 95.8% identity with the sequence of the avian EAV-HP retrovirus (EMBL accession number: AJ238124) [22]. A typical proviral structure consists of gag, pol and env flanked by long terminal repeat (LTR), which are arranged in the order of 5′LTR-gag-pol-env-LTR3′ [22]. Here, the inserted retrovirus is absent of the whole pol gene and part of gag and env (Figure 3A). The retrovirus was integrated into the blue-shelled chicken genome in an inverted orientation (Figure 3B) at Chr1 : 67324641–67324642. We also found that the EAV encompassed some promoter elements by sequence analysis, indicating its expression promotion activities (Figure 3A).
Fig. 3. The EAV-HP insertion in blue-shelled chickens. 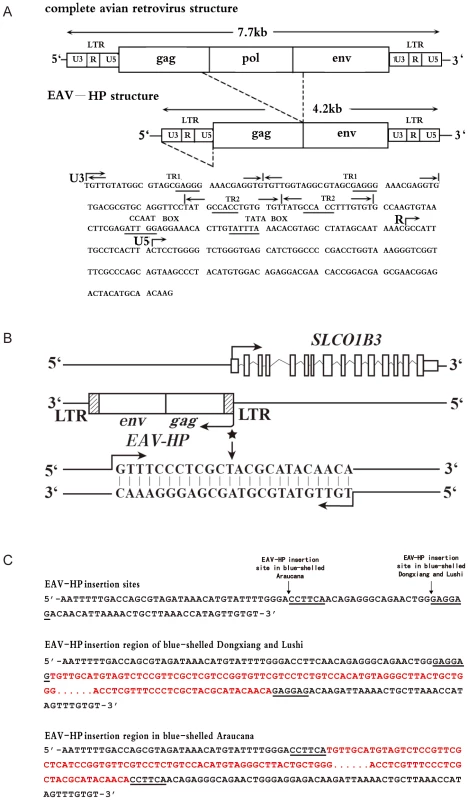
(A) Schematic diagram of the relationship of a complete avian leukosis virus and the EAV-HP retrovirus. Some putative cis-regulatory elements in long terminal repeat (LTR) are underlined. The diagram was redrawn by referencing Sacco et al. [22]. (B) Arrangement of EAV-HP retrovirus and SLCO1B3 in chicken genome. EAV-HP integrates into the 5′ end of SLCO1B3 in inverted orientation. When SLCO1B3 is transcribed, an extra 24-bp sequence from the EAV-HP is also compiled into SLCO1B3 transcript. The asterisk indicates the site of 24-bp sequence from EAV-HP, and arrows indicate the direction of transcription. (C) EAV-HP junction sites in blue-shelled chickens of Dongxiang, Lushi and Araucana breeds. The arrows show the EAV-HP inserted sites and the underlined sequences are the EAV-HP integration specific sequences. The sequences in red denote the inserted EAV-HP. A wide-range survey of the EAV-HP insertion was performed in 705 chickens from 12 worldwide breeds and the F2 resource population (Table 2) using diagnostic PCR test. The results that the EAV-HP insertion is completely associated with the blue eggshell phenotype provide strong evidence that the mutation is causative.
Tab. 2. EAV-HP insertion distributions in various populations. 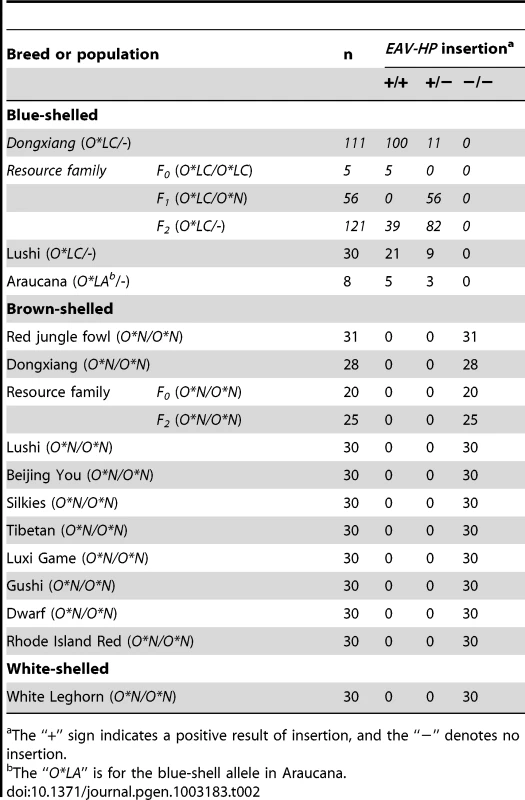
The “+” sign indicates a positive result of insertion, and the “−” denotes no insertion. Independent EAV-HP insertion events in blue-shelled chickens from China and Chile
In order to elucidate whether the blue-shelled chickens from China and Chile have the same origin for the genotypic mutation, we further sequenced the EAV-HP insertion regions in a homozygous Araucana and a homozygous blue-shelled Lushi chicken. The EAV-HP insertion was found in both samples and the alignments of Araucana and Lushi to Dongxiang blue-shelled chicken showed the identity of the inserted EAV-HP being around 97%. Interestingly, the insertion sites in Araucana are different from that in the two Chinese blue-shelled chickens. The break point for EAV-HP insertion in blue-shelled Araucana is located at 23 bp upstream to that in the two Chinese breeds (Figure 3C). We sequenced the junction sites in homozygous blue-shelled chickens of Araucana (n = 5), Lushi (n = 5) and Dongxiang (n = 5) and confirmed the insertion sites in Dongxiang and Lushi are the same but different from that in Araucana.
We also typed 21 SNPs in genomic region of SLCO1B3 in multiple breeds of blue-shelled and non-blue-shelled chicken breeds. It is obvious to see that the EAV-HP insertions in blue-shelled chickens from the two continents were embedded in two distinguished different haplotypes (Table S3), which supports independent integrations accounting for the blue shelled phenotypes.
Discussion
In birds, eggshell color is a variable Mendelian trait. Colored eggshell could function as avoiding predation through either crypsis or aposematism, distinguishing from brood parasitism, reinforcing eggshell strength, regulating egg temperature, combating harmful solar radiation and sending sexually selected signal to males [2], [23]. However, molecular mechanism of all kinds of eggshell color formation is poorly understood to date. Here, we demonstrate that a ∼4.2 kb EAV-HP insertion at upstream of SLCO1B3 is responsible for blue eggshell phenotype in the chicken.
By linkage analysis, we fine mapped the O locus to a 120 kb region, where four candidate genes of SLCO1C1, SLCO1B3, LOC418189 and SLCO1A2 are located. These genes are all members of organic anion transporting polypeptides (OATP, gene symbol SLCO). Functionally, the OATPs serve as membrane transporters that mediate a wide range of sodium-independent transport of amphipathic organic compounds, such as some endobiotic compounds of bile salts, eicosanoids, sterioids, thyroid hormones and some xenobiotic compounds of anionic oligopeptides, organic dyes, toxins, and drugs [24]. SLCO1B3 codes a membrane transporter OATP1B3 which is considered a liver-specific transporter and is highly expressed in liver where it transports a wide range of substrates including bile salts [24], [25]. A genome-wide association study (GWAS) for serum bilirubin levels also showed SLCO1B3 is a plausible candidate gene responsible for changes in bilirubin levels in humans [26]. As blue egg is colored mainly by deposition of biliverdin on the eggshell and biliverdin is just one component of the bile salts, the expression of the SLCO1B3 in uterus could enhance transportation of biliverdin to eggshell. In this study, we found that SLCO1B3 is exclusively expressed in shell gland of uterus of blue-shelled chickens rather than in that of brown - or white-shelled chickens, which supports that the gene plays a pivotal role for coloration of blue eggs.
Regulatory mutations demonstrate an important role for phenotypic diversity which may be explained by cis-acting elements [17], [27]–[34]. The effect of endogenous retrovirus (ERV) on hosts is extensive. It can unfavorably influence certain production traits, i.e. egg production, egg weight and body weight [35], induce lymphoid or erythroid leukosis and a variety of tumors [35], and cause some phenotype variants, i.e. dilute coat color mutation [36] and hairless mutation in mice [37], recessive white [38], henny-feathering mutation [39], and the sex-linked late-feathering mutation [40] in chickens and outheld wing mutation in Drosophila melanogaster [41]. ERV could alter splicing patterns of transcript to produce variants such as the recessive white mutation in the chickens [38]. ERV could also promote expression of genes in alternative tissues, which is associated with activity of LTR which contains promoter and/or enhancer sequences responsible for transcription of virus genes [22] and may induce expression of flanking genes. Avian lymphoid leukosis and the henny-feathering mutation are respectively related to activation of c-myc in B cell and aromatase in the extrogonadal tissues by LTR [35], [39]. We found an EAV-HP inserted in 5′ flanking region of SLCO1B3 in reverse orientation. The LTR of EAV-HP could induce expression of the downstream gene (SLCO1B3) by its bidirectional promoter activity [22]. Moreover, 5′UTR of SLCO1B3 transcripts from the blue-shell allele containing the 24 bp EAV-HP partial sequence implies that the expression of SLCO1B3 in blue-shelled chickens is closely related to the insertion.
Blue eggshells are also seen in other avian species, such as domestic duck, Japanese quail and wild birds [4]–[7], [9]. The genetic pattern of duck blue egg is similar to that of chicken, displaying a dominant phenotype determined by a single gene [42]. However, SLCO1B3 is not expressed in the uterus of blue-shelled and non-blue-shelled ducks, and the EAV-HP insertion was not found in the homologous region in duck (Figure S2, Primers in Table S4). Thus, the causative gene for blue eggshell in ducks may be different from that in chickens. Moreover, the genetic pattern in the chicken is also different from that for Japanese blue-eggshell quail which arise from a recessive mutation ce [7]. Because there is no record showing that the two ancestral species of domestic chickens, red jungle fowl and grey jungle fowl [33], lay blue eggs, we may conclude that the causative EAV-HP insertion for blue eggshell is a derived mutation in the domestic chicken.
China and Chile are two countries reported for having indigenous blue-shelled chicken breeds. Araucana from Chile, Dongxiang and Lushi from China got the blue eggshell phenotype and were all bred for several hundred years. Analysis with mtDNA showed both Indo-European and Asian origins of Chilean and Pacific chickens and blue/green-shell trait in the Araucana did not originated from ancient pacific/pre-Columbian chickens [43]. It is noted in the present study that though all these blue-shelled chickens had the EAV-HP insertion, the EAV-HPs inserted into two different genomic sites in the 5′ flanking region of SLCO1B3 in the blue-shelled chickens from the two countries (Figure 3C) and the EAV-HP insertion in blue-shelled Araucana embedded in a haplotype which is distinctly different from the corresponding haplotype from blue-shelled Lushi and Dongxiang chickens (Table S3). Here, we provide unambiguous evidences that the genetic basis of blue shell phenotype in Araucana is different from that in Chinese blue-shelled breeds, indicating independent originations of the trait in different continents. Due to the blue eggshell mutation having been artificially selected for consumption and variable eggshell color types for human requirements, the separated insertion events present us another parallel evolution case at the molecular level under adaptive selection by humans.
Materials and Methods
Animals
Two Chinese indigenous blue-shelled chicken breeds, Dongxiang and Lushi, and an American blue-shelled breed, Araucana, were used in the present study. Dongxiang chicken is from Dongxiang town, Jiangxi province of China. It is characterized by blue eggshell, single comb and black feather. Historically Dongxiang chicken is selected for blue eggshell, however, the trait has not been fixed to date. Lushi chicken is another local breed laying blue-shelled egg from Lushi town, Henan province of China. Because Lushi chicken has not been systematically bred, some appearance traits, eggshell color, as well as feather color does not show homogeneity. Araucana is an indigenous breed from Chile of South America. Besides blue-shelled egg, two distinguishing characteristics of Araucana breed are rumpless and tufts of feathers which protrude from each side of neck.
In the present study, Dongxiang chicken and Lushi chicken were collected from Jiangxi Hualv breed poultry conservation farm and Henan Sanmenxia Lushi chicken farm, respectively. The blood samples of Araucana were obtained from members of the Araucana Club of America. We also collected 9 non-blue-shelled chicken breeds including Red Jungle Fowl, White Leghorn, Rhode Island Red, Beijing You, Silkie, Tibetan, Luxi Game, Gushi and Dwarf (a commercial layer line in China).
A three-generation F2 resource family was constructed by crossing homozygous blue-shelled Dongxiang (O*LC/O*LC) males, which has been verified by a test cross, and brown-shelled Dongxiang (O*N/O*N) females. All F1 hens laid blue shelled eggs and individual egg color phenotypes were recorded for all 146 F2 hens.
Six heterozygous (O*LC/O*N) hens were produced by mating one homozygous Dongxiang blue-shelled (O*LC/O*LC) male and one White Leghorn (O*N/O*N) female, and the progeny were used for pyrosequencing analysis.
All animal research was approved by Beijing Administration Committee of Laboratory Animals under the leadership of the Beijing Association for Science and Technology, the approve ID is SYXK (Beijing) 2007–0023.
DNA was extracted from blood using standard phenol/chloroform method. RNA was extracted from the liver and uterus. All the tissue samples for RNA isolation were collected at 3 to 5 hours before the next expected oviposition.
Linkage analysis
The F2 resource family was used for linkage analysis. A set of 8 markers covering the region anchored by GWAS and SOX5, ev1 were used in the linkage analysis (Figure 2). Marker L1 and L4–L7 were adopted from previous reports [18], [20] and L2, L3 and L8 were mined from the chicken genome assemble (Build 2.1) at http://genome.ucsc.edu/cgi-bin/hgGateway. Fifteen SNP markers (L9–L23) between L4 and L5 were added to narrow the mapping region. Primers and genotyping methods for all markers were present in Table S1. CRI-MAP 2.4 was used for linkage analysis [44]. The TWO-POINT option was used to calculate the recombination fractions between loci as well as corresponding LOD-scores. The CHROMPIC option was used to find unlikely double recombinants.
Expression analysis
Total RNA was extracted from the uterus using Trizol reagent (TianGen, Dalian, China), followed by synthesis of cDNA from 2 µg of RNA using M-MLV reverse transcriptase (Promega, CA, USA). Five pairs of primer were designed for the four candidate genes (SLCO1C1, SLCO1B3, LOC418189 and SLCO1A2) and housekeeping gene GAPDH using Primer 5.0 for RT-PCR (Table S4) and all primer pairs were designed to span an intron at least. Expression analysis of the four candidate genes was performed in the uterus of blue-shelled (n = 16) and brown-shelled (n = 16) Dongxiang chickens. RT-PCR amplification conditions were as follows: 94°C for 5 min, followed by 36 cycles of amplification (94°C for 30 s, 58°C for 30 s, 72°C for 20 s) and one cycle of 72°C for 5 min.
Expression of SLCO1B3 in the uterus was subsequently detected in a set of samples including homozygous blue-shelled (O*LC/O*LC, n = 4), heterozygous blue-shelled (O*LC/O*N, n = 4), and brown-shelled (O*N/O*N, n = 4) Dongxiang chickens, homozygous blue-shelled (O*LC/O*LC, n = 3), heterozygous blue-shelled (O*LC/O*N, n = 3), and brown-shelled (O*N/O*N, n = 6) Lushi chickens, and 4 non-blue-shelled breeds including White Leghorn O*N/O*N, n = 6), Beijing You (O*N/O*N, n = 6), Silkies (O*N/O*N, n = 6), Tibetan chicken (O*N/O*N, n = 6) by real-time PCR with a Bio-Rad CFX96 instrument (Bio-Rad, CA, USA). Samples were run in triplicates using RealMasterMix (SYBR Green I) (Tiangen, Dalian, China). PCR amplifications were carried out in a 20 µL reaction volume containing 1.2 pmol of each primer, 9 µL of 2.5×working concentration RealMasterMix and 1 µL of cDNA in following cycling conditions: 95°C for 2 min, followed by 40 cycles of 95°C for 10 s, 58°C for 10 s, 68°C for 10 s. GAPDH is used as endogenous reference gene to normalizing amounts of input cDNA, heterozygous blue-shelled Dongxiang (O*LC/O*N) group was designed as a calibrator. Fold change of every group related to the calibrator was calculated as described in Livak et al. [45].
Pyrosequencing
Tissues (the uterus and liver) were collected from the six blue-eggshell heterozygotes. Total RNA was extracted from the uterus and liver with trizol (Tiangen, Dalian, China). The RNA quality was controlled using NanoVue plus spectrophotometer (GE Healthcare, USA). The first-strand cDNA synthesis used M-MLV (Promega, CA, USA) with the 18 hexamers. A fragment containing the SNP (g. 67334934 G>T) in exon 5 was amplified with forward (CATGTTGCGAGGAATTGGTG) and reverse (TTCCTTAGCAAAATCGTCAAGATA) primers. The relative expression of the two allele (O*LC or O*N) transcripts in heterzygotes was scored by analyzing the SNP (g. 67334934 G>T) by pyrosequencing. A pyro-seq primer (CGTCAAGATAAGAGATGCC) was used as the sequencing primer and all steps were performed according to manufacturer's protocol. All samples were analyzed in triplicates.
Fluorescent in situ hybridization
The uterus from a 60-week-old egg laying blue-shelled hen and a non-blue-shelled hen were collected and fixed in 4% paraformaldehyde in phosphate buffered saline (PBS) for 24 hours at room temperature. Fixed uterus was embedded in irrigation solution PBS for six hours to eliminate 4% paraformaldehyde. Then slides were dehydrated in increasing concentrations of ethanol (50%, 70%, 80%, 90%, 95% for 1.5 hours each and 100% for 2 hours) followed by transparentizing in two clearing agents respectively of xylene for 15 minutes each. After transparentizing, the slides were pretreated by the mixture of xylene and low melting paraffin for 30 minutes then were directly transferred into pure melting paraffin (58°C) twice for 3 hours each.
The cDNA probe 5′-AACTCTGGCTGAACGCATCT-3′ were labeled by 6-FAM and were synthesized from mRNA of SLCO1B3 (XM416418.2) by Boxing Bio-engineering Limited Company (Boxing, Guangdong, China). The in situ hybridization was then carried out according to the instruction of the FISH Detection Kit (Boxing, Guangdong, China). Imaging was performed using a fluorescence microscope equipped with vision software.
Resequencing of SLCO1B3
Twenty-four kilobases fragment (GenBank accession No. JN020139) covering the whole SLCO1B3 was resequenced using a panel of ten birds from 5 blue-shelled (O*LC/O*LC) and 5 brown-shelled (O*N/O*N) Dongxiang chickens. Seventeen primer pairs used to generate overlapping PCR amplicons ranging from approximately 800 bp to 2000 bp in size were listed in Table S5. The PCR amplifications were performed in a total volume of 50 µL containing 5 µL of 10×Taq polymerase buffer, 10 mmol of each deoxynucleotide triphosphate (dNTP), 20 pmol of each primer, 2.5 U Taq DNA polymerase (HT-biotech, Beijing, China), and 50 ng genomic DNA. All purified PCR products were directly sequenced in both directions using the same primers. The sequences were assembled and analyzed for polymorphisms using the ChromasPro 1.5 or BLAST program in UCSC (http://genome.ucsc.edu/cgi-bin/hgBlat?command=start).
Rapid amplification of cDNA end (RACE) of SLCO1B3
In order to analyze the 5′ and 3′ untranslated regions (UTR) of the SLCO1B3 gene, RACE experiments were performed on 2 µg total RNA extracted from the uterus of a homozygous blue-shelled (O*LC/O*LC) and a brown-shelled (O*N/O*N) Dongxiang chicken using 5′ and 3′-Full RACE Kit (Takara, Dalian, China), according to the manufacturer's instructions. 5′ and 3′ UTR of SLCO1B3 gene transcripts were amplified by nested PCR with gene specific (Table S4) and adaptor primers (Table S4) for the first and second amplifications of 5′ and 3′ UTR respectively. First and second PCR amplifications were carried out in a 50 µL reaction volume containing 20 pmol of each primer, 5 µL of 10× LA PCR buffer (Mg2+ plus), 2.5 U of LA Taq (Takara, Dalian, China), 20 mM of each dNTP and 1–2 µL of cDNA or 1st PCR product. RACE products were cloned to pMD-18 vector (Takara, Dalian, China), and then sequenced in both directions.
Long-range PCR
A long-range PCR amplification with 1B3_5F & 5R primer pair (Table S5) was performed in volumes of 50 µL containing 5 µL of 10× LA PCR buffer (Mg2+ plus), 2.5 U of LA Taq (Takara, Dalian, China), 20 mM of each dNTP, 20 pmol of each primer and 50 ng genomic DNA. The PCR condition was as follow: 94°C for 3 min followed by 33 cycles of 94°C for 30 s, 58°C for 30 s, 72°C for 5 min, and a final extension at 72°C for 10 min. The PCR product was completely sequenced using the other three pairs of bridging primers (Table S5) besides the 1B3_5F & 5R.
MassARRAY analysis
Twenty-one SNPs found in resequencing of SLCO1B3 were used to analyze the genetic variants of SLCO1B3 (Table S6). SNP markers were genotyped by iPLEX SEQUENOM MassARRAY platform (Sequenom, CA, USA). This genotyping system used single-base extension reactions to create allele-specific products that are separated automatically and scored in a matrix-assisted laser desorption ionization/time of flight mass spectrometer. Primer design was performed using MassARRAY Assay Design software (v3.1) according to Sequenom's instructions. Multiplex PCR amplification of amplicons containing SNPs of interest was performed using HotStart Taq Polymerase (Qiagen, CA, USA) with 12 ng genomic DNA. Assay data were analyzed using Sequenom TYPER software (v3.4).
Diagnostic genotyping test of EAV-HP insertion
The retrovirus insertion was genotyped with a mix of three primers: the primer “test-nor-up” 5′ - TTTGACCAGCGTAGATAA-3′ and “test-nor-down” 5′-ATGTTAGCAGTGTAGTTG-3′ were located in the wild type genomic sequence of SLCO1B3, the primer “test-eav” 5′-TAGGTTCCGAACGCGATGT-3′ was located in the gag region of the inserted retroviral sequence (Figure S3). The PCR amplifications were preformed in a total volume of 25 µL containing 2.5 µL of 10×Taq polymerase buffer, 5 mmol of each deoxynucleotide triphosphate (dNTP), 10 pmol of each primer, 1.25 U Taq DNA polymerase (HT-biotech, Beijing, China), and 50 ng genomic DNA in the following condition: 94°C for 5 min, followed by 36 cycles of 94°C for 30 s, 58°C for 30 s, 72°C for 20 s, and a final extension at 72°C for 5 min. The PCR products was separated by 2% agarose gel electrophoresis, and the length of target fragment was 340 bp for test-nor-up and test-nor-down, and 425 bp for test-nor-up and test-eav, respectively (Figure S3).
EAV-HP insertion site sequencing
Two pairs of PCR primers (EAVIS-1F, EAVIS-1R, EAVIS-2F and EAVIS-2R, Table S5) were designed for amplifying 5′ and 3′ end of EAV-HP junction regions. The PCR condition was as follow: 94°C for 3 min followed by 33 cycles of 94°C for 30 s, 57°C for 30 s and 54°C for 40 s, respectively, 72°C for 45 s, and a final extension at 72°C for 10 min. The PCR products were sequenced bidirectionally using the PCR primers.
Supporting Information
Zdroje
1. Underwood TJ, Sealy SG (2002) Adaptive significance of egg coloration. In: Deeming DC, editors. Avian Incubation: Behaviour, Environment and Evolution: Oxford University Press. pp. 280–298.
2. MorenoJ, OsornoJL (2003) Avian egg colour and sexual selection: Does eggshell pigmentation reflect female condition and genetic quality? Ecol Lett 6 : 803–806.
3. MoralesJ, SanzJJ, MorenoJ (2006) Egg colour reflects the amount of yolk maternal antibodies and fledging success in a songbird. Biol Lett 2 : 334–336.
4. SieffermanL, NavaraKJ, HillGE (2006) Egg coloration is correlated with female condition in eastern bluebirds (Sialia sialis). Behav Ecol Sociobiol 59 : 651–656.
5. MoralesJ, TorresR, VelandoA (2010) Parental conflict and blue egg coloration in a seabird. Naturwissenschaften 97 : 173–180.
6. MorenoJ, MoralesJ, LobatoE, MerinoS, TomásG, et al. (2005) Evidence for the signaling function of egg color in the pied flycatcher Ficedula hyoleuca. Behav Ecol 16 : 931–937.
7. ItoS, TsudzukiM, KomoriM, MizutaniM (1993) Celadon: An eggshell color mutation in Japanese quail. J Hered 84 : 145–147.
8. PunnettRC (1933) Genetic study in poultry-IX. The blue egg. Genetics 27 : 465–470.
9. WangCT, WanTC, PanCM, ChenYH (1997) Comparisons of physical-chemical properties and alkalizing process between greenish and whitish eggs of Brown Tsaiya duck. J Chin Agri Chem Soc 35 : 263–272.
10. LangMR, WellsJW (1987) A review of eggshell pigmentation. World Poult Sci J 43 : 238–245.
11. KennedyGY, VeversHG (1973) Eggshell pigments of the Araucano fowl. Comp Biochem Physiol B 44 : 11–25.
12. BrucknerJH, HuttFB (1939) Linkage of Pea Comb and Blue Egg in the Fowl. Science 90 : 88.
13. BitgoodJJ, ShoffnerRN, OtisJS, BrilesWE (1980) Mapping of the genes for pea comb, blue egg, barring, silver, and blood groups A, E, H, and P in the domestic fowl. Poult Sci 59 : 1686–1693.
14. BitgoodJJ, OtisJS, ShoffnerRN (1983) Refined linkage value for comb and blue Egg: lack of effect of pea comb, blue egg, and naked on age at first egg in the domestic fowl. Poult Sci 62 : 235–238.
15. BitgoodJJ, BrilesRW, BrilesWE (2000) Further tests for genetic linkages of three morphological traits, three blood groups, and break points of two chromosome translocations on chromosome one in the chicken. Poult Sci 79 : 293–295.
16. BartletJR, JonesCP, SmithEJ (1996) Linkage analysis of endogenous viral element 1, blue eggshell, and pea comb loci in chickens. J Hered 87 : 67–70.
17. WrightD, BoijeH, MeadowsJR, Bed'homB, GourichonD, et al. (2009) Copy number variation in intron 1 of SOX5 causes the Pea-comb phenotype in chickens. PLoS Genet 5: e1000512.
18. ZhaoR, LiuZZ, XuGY, YangN (2007) Analysis of SNP markers for chicken blue-shelled gene using PCR-SSCP. Chin J Agric Biotech 4 : 53–56.
19. WangXT, BaiJR, ZhaoCJ, ZhangH, BaoHG, et al. (2010) Localization of the genomic sequence interval for the blue eggshell gene using an F2 resource population of Dongxiang chickens. Brit Poult Sci 51 : 507–509.
20. WangZP, WangXT, LiuRF, LiJY, DengXM (2010) Refined localization of the O gene for blue egg phenotype on chicken chromosome 1. J Anim Vet Adv 9 : 2947–2950.
21. WraggD, MwacharoJM, AlcaldeJA, HockingPM, HanotteO (2012) Analysis of genome-wide structure, diversity and fine mapping of Mendelian traits in traditional and village chickens. Heredity 109 : 6–18.
22. SaccoMA, FlanneryDM, HowesK, VenugopalK (2000) Venugopal, Avian endogenous retrovirus EAV-HP shares regions of identity with avian leukosis virus subgroup J and the avian retrotransposon ART-CH. J Virol 74 : 1296–1306.
23. ReynoldsSJ, MartinGR, CasseyP (2009) Is sexual selection blurring the functional significance of eggshell coloration hypotheses? Anim Behav 78 : 209–215.
24. PopovicM, ZajaR, SmitalT (2010) Organic anion transporting polypeptides (OATP) in zebrafish (Danio rerio): Phylogenetic analysis and tissue distribution. Comp Biochem Physiol A 155 : 327–335.
25. HagenbuchB, GuiC (2008) Xenobiotic transporters of the human organic anion transporting polypeptides (OATP) family. Xenobiotica 38 : 778–801.
26. SannaS, BusoneroF, MaschioA, McArdlePF, UsalaG, et al. (2009) Common variants in the SLCO1B3 locus are associated with bilirubin levels and unconjugated hyperbilirubinemia. Hum Mol Genet 18 : 2711–2718.
27. GunnarssonU, KerjeS, Bed'homB, SahlqvistAS, EkwallO, et al. (2010) The Dark brown plumage color in chickens is caused by an 8.3-kb deletion upstream of SOX10. Pigment Cell Melanoma Res 24 : 268–274.
28. PailhouxE, VigierB, ChaffauxS, ServelN, TaouritS, et al. (2001) A 11.7-kb deletion triggers intersexuality and polledness in goats. Nat Genet 29 : 453–458.
29. GiuffraE, TörnstenA, MarklundS, Bongcam-RudloffE, ChardonP, et al. (2002) A large duplication associated with dominant white color in pigs originated by homologous recombination between LINE elements flanking KIT. Mamm Genome 13 : 569–577.
30. Van LaereAS, NguyenM, BraunschweigM, NezerC, ColletteC, et al. (2003) A regulatory mutation in IGF2 causes a major QTL effect on muscle growth in the pig. Nature 425 : 832–836.
31. ClopA, MarcqF, TakedaH, PirottinD, TordoirX, et al. (2006) A mutation creating a potential illegitimate microRNA target site in the myostatin gene affects muscularity in sheep. Nat Genet 38 : 813–818.
32. KarlssonEK, BaranowskaI, WadeCM, Salmon HillbertzNH, ZodyMC, et al. (2007) Efficient mapping of mendelian traits in dogs through genome-wide association. Nat Genet 39 : 1321–1328.
33. ErikssonJ, LarsonG, GunnarssonU, Bed'homB, Tixier-BoichardM, et al. (2008) Identification of the yellow skin gene reveals a hybrid origin of the domestic chicken. PLoS Genet 4: e1000010.
34. Rosengren PielbergG, GolovkoA, SundströmE, CurikI, LennartssonJ, et al. (2008) A cis-acting regulatory mutation causes premature hair graying and susceptibility to melanoma in the horse. Nat Genet 40 : 1004–1009.
35. PayneLN (1998) Retrovirus-induced disease in poultry. Poult Sci 77 : 1204–1212.
36. JenkinsNA, CopelandNG, TaylorBA, LeeBK (1981) Dilute (d) coat colour mutation of DBA/2J mice is associated with the site of integration of an ecotropic MuLV genome. Nature 293 : 370–374.
37. StoyeJP, FennerS, GreenoakGE, MoranC, CoffinJM (1988) Role of endogenous retroviruses as mutagens: the hairless mutation of mice. Cell 54 : 383–391.
38. ChangCM, CovilleJL, CoquerelleG, GourichonD, OulmoudenA, et al. (2006) Complete association between a retroviral insertion in the tyrosinase gene and the recessive white mutation in chickens. BMC Genomics 7 : 19.
39. MatsumineH, HerbstMA, OuSH, WilsonJD, McPhaulMJ (1991) Aromatase mRNA in the extragonadal tissues of chickens with the henny-feathering trait is derived from a distinctive promoter structure that contains a segment of a retroviral long terminal repeat. Functional organization of the Sebright, Leghorn, and Campine aromatase genes. J Biol Chem 266 : 19900–19907.
40. BaconLD, SmithE, CrittendenLB, HavensteinGB (1988) Association of the slow feathering (K) and an endogenous viral (ev21) gene on the Z chromosome of chickens. Poult Sci 67 : 191–197.
41. YuB, WangXT, LiHW, ZhaoCJ, WuCX, et al. (2011) Structural analysis of a 4414-bp element in Drosophila melanogaster. Genet Mol Res 10 : 717–730.
42. LaiYZ, ZhangSZ (1991) Genetic analysis of duck eggshell color. Hereditas 13 : 4–5.
43. GongoraJ, RawlenceNJ, MobegiVA, JianlinH, AlcaldeJA, et al. (2008) Indo-European and Asian origins for Chilean and Pacific chickens revealed by mtDNA. Proc Natl Acad Sci U S A 105 : 10308–10313.
44. Green P, Falls K, Crooks S (1990) Documentation for CRIMAP, version 2.4. Washington University School of Medicine, St Louis, MO.
45. LivakKJ, SchmittgenTD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 25 : 402–408.
Štítky
Genetika Reprodukčná medicína
Článek Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across PathogensČlánek TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association StudiesČlánek Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization inČlánek Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA ExpressionČlánek The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of GenesČlánek The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
Článok vyšiel v časopisePLOS Genetics
Najčítanejšie tento týždeň
2013 Číslo 1- Gynekologové a odborníci na reprodukční medicínu se sejdou na prvním virtuálním summitu
- Je „freeze-all“ pro všechny? Odborníci na fertilitu diskutovali na virtuálním summitu
-
Všetky články tohto čísla
- A Model of High Sugar Diet-Induced Cardiomyopathy
- Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across Pathogens
- Emerging Function of Fat Mass and Obesity-Associated Protein (Fto)
- Positional Cloning Reveals Strain-Dependent Expression of to Alter Susceptibility to Bleomycin-Induced Pulmonary Fibrosis in Mice
- Genetics of Ribosomal Proteins: “Curiouser and Curiouser”
- Transposable Elements Re-Wire and Fine-Tune the Transcriptome
- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- MAML1 Enhances the Transcriptional Activity of Runx2 and Plays a Role in Bone Development
- Predicting Mendelian Disease-Causing Non-Synonymous Single Nucleotide Variants in Exome Sequencing Studies
- A Systematic Mapping Approach of 16q12.2/ and BMI in More Than 20,000 African Americans Narrows in on the Underlying Functional Variation: Results from the Population Architecture using Genomics and Epidemiology (PAGE) Study
- Transcription of the Major microRNA–Like Small RNAs Relies on RNA Polymerase III
- Histone H3K56 Acetylation, Rad52, and Non-DNA Repair Factors Control Double-Strand Break Repair Choice with the Sister Chromatid
- Genome-Wide Association Study Identifies a Novel Susceptibility Locus at 12q23.1 for Lung Squamous Cell Carcinoma in Han Chinese
- Genetic Disruption of the Copulatory Plug in Mice Leads to Severely Reduced Fertility
- The [] Prion Exists as a Dynamic Cloud of Variants
- Adult Onset Global Loss of the Gene Alters Body Composition and Metabolism in the Mouse
- Fis Protein Insulates the Gene from Uncontrolled Transcription
- The Meiotic Nuclear Lamina Regulates Chromosome Dynamics and Promotes Efficient Homologous Recombination in the Mouse
- Genome-Wide Haplotype Analysis of Expression Quantitative Trait Loci in Monocytes
- TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association Studies
- Structural Basis of a Histone H3 Lysine 4 Demethylase Required for Stem Elongation in Rice
- The Ecm11-Gmc2 Complex Promotes Synaptonemal Complex Formation through Assembly of Transverse Filaments in Budding Yeast
- MCM8 Is Required for a Pathway of Meiotic Double-Strand Break Repair Independent of DMC1 in
- Comparative Genomic Analysis of the Endosymbionts of Herbivorous Insects Reveals Eco-Environmental Adaptations: Biotechnology Applications
- Integration of Nodal and BMP Signals in the Heart Requires FoxH1 to Create Left–Right Differences in Cell Migration Rates That Direct Cardiac Asymmetry
- Pharmacodynamics, Population Dynamics, and the Evolution of Persistence in
- A Hybrid Likelihood Model for Sequence-Based Disease Association Studies
- Aberration in DNA Methylation in B-Cell Lymphomas Has a Complex Origin and Increases with Disease Severity
- Multiple Opposing Constraints Govern Chromosome Interactions during Meiosis
- Transcriptional Dynamics Elicited by a Short Pulse of Notch Activation Involves Feed-Forward Regulation by Genes
- Dynamic Large-Scale Chromosomal Rearrangements Fuel Rapid Adaptation in Yeast Populations
- Heterologous Gln/Asn-Rich Proteins Impede the Propagation of Yeast Prions by Altering Chaperone Availability
- Gene Copy-Number Polymorphism Caused by Retrotransposition in Humans
- An Incompatibility between a Mitochondrial tRNA and Its Nuclear-Encoded tRNA Synthetase Compromises Development and Fitness in
- Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization in
- Single-Stranded Annealing Induced by Re-Initiation of Replication Origins Provides a Novel and Efficient Mechanism for Generating Copy Number Expansion via Non-Allelic Homologous Recombination
- Tbx2 Controls Lung Growth by Direct Repression of the Cell Cycle Inhibitor Genes and
- Suv4-20h Histone Methyltransferases Promote Neuroectodermal Differentiation by Silencing the Pluripotency-Associated Oct-25 Gene
- A Conserved Helicase Processivity Factor Is Needed for Conjugation and Replication of an Integrative and Conjugative Element
- Telomerase-Null Survivor Screening Identifies Novel Telomere Recombination Regulators
- Genome-Wide Analysis Reveals Selection for Important Traits in Domestic Horse Breeds
- Coordinated Degradation of Replisome Components Ensures Genome Stability upon Replication Stress in the Absence of the Replication Fork Protection Complex
- Nkx6.1 Controls a Gene Regulatory Network Required for Establishing and Maintaining Pancreatic Beta Cell Identity
- HIF- and Non-HIF-Regulated Hypoxic Responses Require the Estrogen-Related Receptor in
- Delineating a Conserved Genetic Cassette Promoting Outgrowth of Body Appendages
- The Telomere Capping Complex CST Has an Unusual Stoichiometry, Makes Multipartite Interaction with G-Tails, and Unfolds Higher-Order G-Tail Structures
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Loci Associated with -Glycosylation of Human Immunoglobulin G Show Pleiotropy with Autoimmune Diseases and Haematological Cancers
- Switchgrass Genomic Diversity, Ploidy, and Evolution: Novel Insights from a Network-Based SNP Discovery Protocol
- Centromere-Like Regions in the Budding Yeast Genome
- Sequencing of Loci from the Elephant Shark Reveals a Family of Genes in Vertebrate Genomes, Forged by Ancient Duplications and Divergences
- Mendelian and Non-Mendelian Regulation of Gene Expression in Maize
- Mutational Spectrum Drives the Rise of Mutator Bacteria
- Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA Expression
- The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of Genes
- Sex-Specific Signaling in the Blood–Brain Barrier Is Required for Male Courtship in
- A Newly Uncovered Group of Distantly Related Lysine Methyltransferases Preferentially Interact with Molecular Chaperones to Regulate Their Activity
- Is Required for Leptin-Mediated Depolarization of POMC Neurons in the Hypothalamic Arcuate Nucleus in Mice
- Unlocking the Bottleneck in Forward Genetics Using Whole-Genome Sequencing and Identity by Descent to Isolate Causative Mutations
- The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
- MTERF3 Regulates Mitochondrial Ribosome Biogenesis in Invertebrates and Mammals
- Downregulation and Altered Splicing by in a Mouse Model of Facioscapulohumeral Muscular Dystrophy (FSHD)
- NBR1-Mediated Selective Autophagy Targets Insoluble Ubiquitinated Protein Aggregates in Plant Stress Responses
- Retroactive Maintains Cuticle Integrity by Promoting the Trafficking of Knickkopf into the Procuticle of
- Phenome-Wide Association Study (PheWAS) for Detection of Pleiotropy within the Population Architecture using Genomics and Epidemiology (PAGE) Network
- Genetic and Functional Modularity of Activities in the Specification of Limb-Innervating Motor Neurons
- A Population Genetic Model for the Maintenance of R2 Retrotransposons in rRNA Gene Loci
- A Quartet of PIF bHLH Factors Provides a Transcriptionally Centered Signaling Hub That Regulates Seedling Morphogenesis through Differential Expression-Patterning of Shared Target Genes in
- A Genome-Wide Integrative Genomic Study Localizes Genetic Factors Influencing Antibodies against Epstein-Barr Virus Nuclear Antigen 1 (EBNA-1)
- Mutation of the Diamond-Blackfan Anemia Gene in Mouse Results in Morphological and Neuroanatomical Phenotypes
- Life, the Universe, and Everything: An Interview with David Haussler
- Alternative Oxidase Expression in the Mouse Enables Bypassing Cytochrome Oxidase Blockade and Limits Mitochondrial ROS Overproduction
- An Evolutionarily Conserved Synthetic Lethal Interaction Network Identifies FEN1 as a Broad-Spectrum Target for Anticancer Therapeutic Development
- The Flowering Repressor Underlies a Novel QTL Interacting with the Genetic Background
- Telomerase Is Required for Zebrafish Lifespan
- and Diversified Expression of the Gene Family Bolster the Floral Stem Cell Network
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Increased Maternal Genome Dosage Bypasses the Requirement of the FIS Polycomb Repressive Complex 2 in Arabidopsis Seed Development
- WNK1/HSN2 Mutation in Human Peripheral Neuropathy Deregulates Expression and Posterior Lateral Line Development in Zebrafish ()
- Synergistic Interaction of Rnf8 and p53 in the Protection against Genomic Instability and Tumorigenesis
- Dot1-Dependent Histone H3K79 Methylation Promotes Activation of the Mek1 Meiotic Checkpoint Effector Kinase by Regulating the Hop1 Adaptor
- A Heterogeneous Mixture of F-Series Prostaglandins Promotes Sperm Guidance in the Reproductive Tract
- Starvation, Together with the SOS Response, Mediates High Biofilm-Specific Tolerance to the Fluoroquinolone Ofloxacin
- Directed Evolution of a Model Primordial Enzyme Provides Insights into the Development of the Genetic Code
- Genome-Wide Screens for Tinman Binding Sites Identify Cardiac Enhancers with Diverse Functional Architectures
- PLOS Genetics
- Archív čísel
- Aktuálne číslo
- Informácie o časopise
Najčítanejšie v tomto čísle- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
Prihlásenie#ADS_BOTTOM_SCRIPTS#Zabudnuté hesloZadajte e-mailovú adresu, s ktorou ste vytvárali účet. Budú Vám na ňu zasielané informácie k nastaveniu nového hesla.
- Časopisy



